Volume 23, Number 2—February 2017
Dispatch
mcr-1−Harboring Salmonella enterica Serovar Typhimurium Sequence Type 34 in Pigs, China
Figure 1
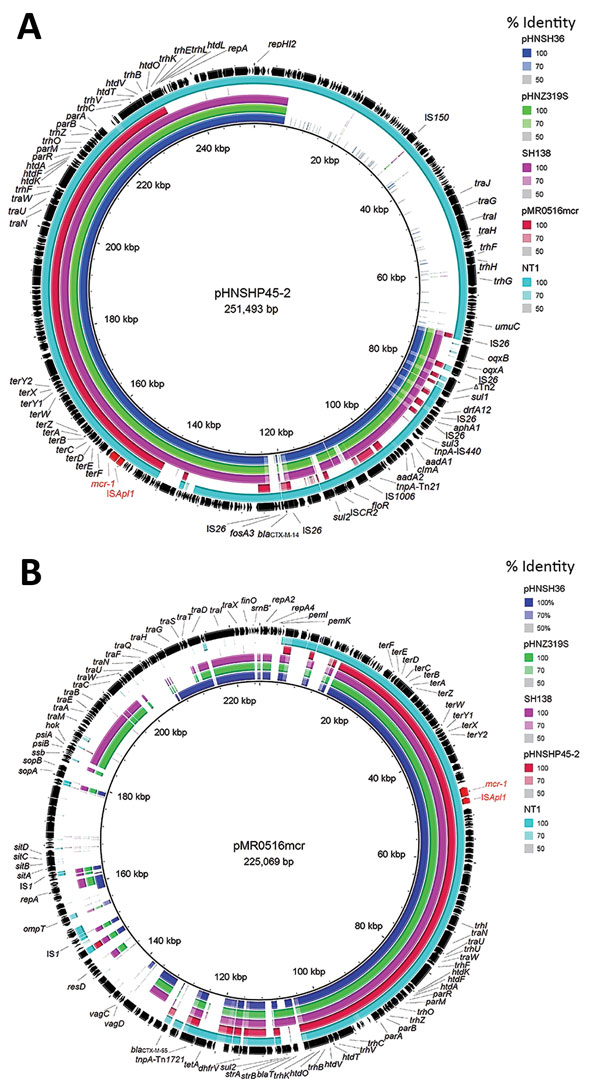
Figure 1. Sequence comparison of scaffolds (portions of genome sequences reconstructed from end-sequenced whole-genome clones) identified in mcr-1–positive plasmids pHNSH36, pHNZ319S, and pHNSH138 with 2 mcr-1–bearing plasmids pHNSHP45-2 (GenBank accession no. KU341381) and pMR0516mcr (GenBank accession no. KX276657), and contigs identified in mcr-1–positive genomes of Escherichia coli strain NT1 in BRIG (11) (GenBank accession LSBW01000090.1) obtained during analysis of mcr-1–positive Salmonella isolates from pigs at slaughter, China, 2013–2014. Arrows indicate positions and direction of transcription of genes. Reference plasmids pHNSHP45–2 (A) and pMR0516mcr (B) are indicated in black in the outer circles.
References
- Liu YY, Wang Y, Walsh TR, Yi LX, Zhang R, Spencer J, et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: a microbiological and molecular biological study. Lancet Infect Dis. 2016;16:161–8. DOIPubMedGoogle Scholar
- Schwarz S, Johnson AP. Transferable resistance to colistin: a new but old threat. J Antimicrob Chemother. 2016;71:2066–70. DOIPubMedGoogle Scholar
- Doumith M, Godbole G, Ashton P, Larkin L, Dallman T, Day M, et al. Detection of the plasmid-mediated mcr-1 gene conferring colistin resistance in human and food isolates of Salmonella enterica and Escherichia coli in England and Wales. J Antimicrob Chemother. 2016;71:2300–5. DOIPubMedGoogle Scholar
- Campos J, Cristino L, Peixe L, Antunes P. MCR-1 in multidrug-resistant and copper-tolerant clinically relevant Salmonella 1,4,[5],12:i:- and S. Rissen clones in Portugal, 2011 to 2015. Euro Surveill. 2016;21:30270. DOIPubMedGoogle Scholar
- Rahn K, De Grandis SA, Clarke RC, McEwen SA, Galán JE, Ginocchio C, et al. Amplification of an invA gene sequence of Salmonella typhimurium by polymerase chain reaction as a specific method of detection of Salmonella. Mol Cell Probes. 1992;6:271–9. DOIPubMedGoogle Scholar
- Sun J, Ke B, Huang Y, He D, Li X, Liang Z, et al. The molecular epidemiological characteristics and genetic diversity of Salmonella Typhimurium in Guangdong, China, 2007-2011. PLoS One. 2014;9:e113145. DOIPubMedGoogle Scholar
- Antunes P, Mourão J, Pestana N, Peixe L. Leakage of emerging clinically relevant multidrug-resistant Salmonella clones from pig farms. J Antimicrob Chemother. 2011;66:2028–32. DOIPubMedGoogle Scholar
- Quesada A, Porrero MC, Téllez S, Palomo G, García M, Domínguez L. Polymorphism of genes encoding PmrAB in colistin-resistant strains of Escherichia coli and Salmonella enterica isolated from poultry and swine. J Antimicrob Chemother. 2015;70:71–4. DOIPubMedGoogle Scholar
- Wong MH, Yan M, Chan EW, Liu LZ, Kan B, Chen S. Expansion of Salmonella Typhimurium ST34 clone carrying multiple resistance determinants in China. Antimicrob Agents Chemother. 2013;57:4599–601. DOIPubMedGoogle Scholar
- Mulvey MR, Finley R, Allen V, Ang L, Bekal S, El Bailey S, et al. Emergence of multidrug-resistant Salmonella enterica serotype 4,[5],12:i:- involving human cases in Canada: results from the Canadian Integrated Program on Antimicrobial Resistance Surveillance (CIPARS), 2003-10. J Antimicrob Chemother. 2013;68:1982–6. DOIPubMedGoogle Scholar
- Alikhan NF, Petty NK, Ben Zakour NL, Beatson SA. BLAST Ring Image Generator (BRIG): simple prokaryote genome comparisons. BMC Genomics. 2011;12:402. DOIPubMedGoogle Scholar
- Aziz RK, Bartels D, Best AA, DeJongh M, Disz T, Edwards RA, et al. The RAST Server: rapid annotations using subsystems technology. BMC Genomics. 2008;9:75. DOIPubMedGoogle Scholar
- Zhi C, Lv L, Yu LF, Doi Y, Liu JH. Dissemination of the mcr-1 colistin resistance gene. Lancet Infect Dis. 2016;16:292–3. DOIPubMedGoogle Scholar
- Brennan E, Martins M, McCusker MP, Wang J, Alves BM, Hurley D, et al. Multidrug-resistant Escherichia coli in bovine animals, Europe. Emerg Infect Dis. 2016;22:1650–2. DOIPubMedGoogle Scholar
- McGann P, Snesrud E, Maybank R, Corey B, Ong AC, Clifford R, et al. Escherichia coli Harboring mcr-1 and blaCTX-M on a Novel IncF Plasmid: First Report of mcr-1 in the United States. Antimicrob Agents Chemother. 2016;60:4420–1. DOIPubMedGoogle Scholar
1These authors contributed equally to this article.