Reassortment of Influenza A Viruses in Wild Birds in Alaska before H5 Clade 2.3.4.4 Outbreaks
Nichola J. Hill, Islam T.M. Hussein, Kimberly R. Davis, Eric J. Ma, Timothy J. Spivey, Andrew M. Ramey, Wendy Blay Puryear, Suman R. Das, Rebecca A. Halpin, Xudong Lin, Nadia B. Fedorova, David L. Suarez, Walter M. Boyce, and Jonathan A. Runstadler

Author affiliations: Massachusetts Institute of Technology, Cambridge, Massachusetts, USA (N.J. Hill, I.T.M. Hussein, K.R. Davis, E.J. Ma, W.B. Puryear, J.A. Runstadler); University of Alaska Fairbanks, Alaska, USA (T.J. Spivey); US Geological Survey, Anchorage, Alaska, USA (T.J. Spivey, A.M. Ramey); Vanderbilt University Medical Center, Nashville, Tennessee, USA (S. Das); J. Craig Venter Institute, Rockville, Maryland, USA (R.A. Halpin, X. Lin, N.B. Fedorova); Department of Agriculture, Athens, Georgia, USA (D.L. Suarez); University of California, Davis, California, USA (W.M. Boyce)
Main Article
Figure 1
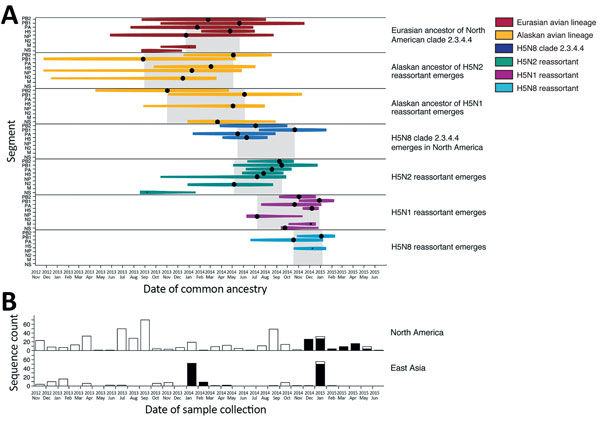
Figure 1. Molecular dating of the emergence of H5 clade 2.3.4.4 influenza A virus in Eurasia and North America and concurrent trends in surveillance effort. A) Events contributing to the evolution of H5 clade 2.3.4.4 estimated using multiple influenza segments. Time of most recent common ancestry (indicated by a black circle) is size-scaled by the posterior probability (0.0–1.0), and the 95% highest posterior density is color-coded by lineage. Gray shading indicates time of most recent common ancestry of multiple segments with a posterior probability >0.85. B) Surveillance effort estimated by the number of hemagglutinin sequences (high and low pathogenicity) available in the Influenza Research Database (https://www.fludb.org). Black bars indicate surveillance effort for H5 clade 2.3.4.4; white bars indicate surveillance effort for other clades. M, matrix gene; N2, neuraminidase 2 gene; NP, nucleoprotein gene; PA, polymerase acidic, PB1, basic polymerase protein 1 gene; PB2, basic polymerase protein 2 gene; tMRCA, time of most recent common ancestry.
Main Article
Page created: March 16, 2017
Page updated: March 16, 2017
Page reviewed: March 16, 2017
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.
