Volume 23, Number 8—August 2017
Research
Human Infection with Highly Pathogenic Avian Influenza A(H7N9) Virus, China
Figure 4
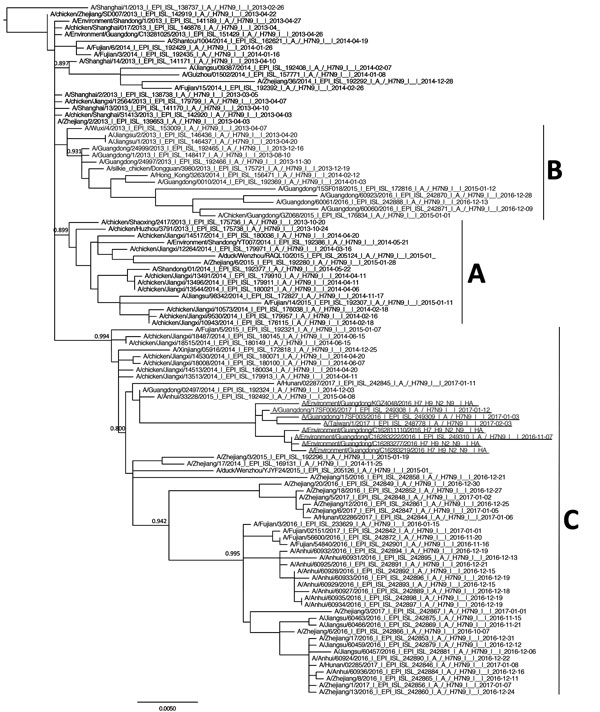
Figure 4. Phylogenetic analysis of the hemagglutinin gene of highly pathogenic avian influenza A(H7N9) viruses in Guangdong Province, China, and reference viruses. Maximum likelihood trees were constructed with PhyML by using the general time reversible plus gamma distribution plus proportion of invariable sites model. Node support was estimated by the SH-like aLRT method, and values >0.8 are shown. Virus clades A, B, and C—previously defined as W2-A, W2-B, and W2-C (5)—are labeled. A/Guangdong/17SF006/2017 (from a 56-year-old man), A/Guangdong/17SF003/2016, A/Taiwan/1/2017, and environmental isolates are underlined. Scale bar indicates nucleotide substitutions per site.
References
- World Heath Organization. Human infection with avian influenza A(H7N9) virus—China. Disease outbreak news 2017 Feb 20 [cited 2017 May 1]. http://www.who.int/csr/don/20-february-2017-ah7n9-china/en/
- World Heath Organization. Human infection with avian influenza A(H7N9) virus—China. Disease outbreak news 2017 Feb 22 [cited 2017 May 1]. http://www.who.int/csr/don/22-february-2017-ah7n9-china/en/
- Qin Y, Horby PW, Tsang TK, Chen E, Gao L, Ou J, et al. Differences in the epidemiology of human cases of avian influenza A(H7N9) and A(H5N1) viruses infection. Clin Infect Dis. 2015;61:563–71. DOIPubMedGoogle Scholar
- Yu H, Cowling BJ, Feng L, Lau EH, Liao Q, Tsang TK, et al. Human infection with avian influenza A H7N9 virus: an assessment of clinical severity. Lancet. 2013;382:138–45. DOIPubMedGoogle Scholar
- Chen Z, Liu H, Lu J, Luo L, Li K, Liu Y, et al. Asymptomatic, mild, and severe influenza A(H7N9) virus infection in humans, Guangzhou, China. Emerg Infect Dis. 2014;20:1535–40. DOIPubMedGoogle Scholar
- Lin YP, Yang ZF, Liang Y, Li ZT, Bond HS, Chua H, et al. Population seroprevalence of antibody to influenza A(H7N9) virus, Guangzhou, China. BMC Infect Dis. 2016;16:632. DOIPubMedGoogle Scholar
- Lam TT, Zhou B, Wang J, Chai Y, Shen Y, Chen X, et al. Dissemination, divergence and establishment of H7N9 influenza viruses in China. Nature. 2015;522:102–5. DOIPubMedGoogle Scholar
- Gao R, Cao B, Hu Y, Feng Z, Wang D, Hu W, et al. Human infection with a novel avian-origin influenza A (H7N9) virus. N Engl J Med. 2013;368:1888–97. DOIPubMedGoogle Scholar
- Chan MC, Chan RW, Chan LL, Mok CK, Hui KP, Fong JH, et al. Tropism and innate host responses of a novel avian influenza A H7N9 virus: an analysis of ex-vivo and in-vitro cultures of the human respiratory tract. Lancet Respir Med. 2013;1:534–42. DOIPubMedGoogle Scholar
- Zhu H, Wang D, Kelvin DJ, Li L, Zheng Z, Yoon SW, et al. Infectivity, transmission, and pathology of human-isolated H7N9 influenza virus in ferrets and pigs. Science. 2013;341:183–6. DOIPubMedGoogle Scholar
- Luczo JM, Stambas J, Durr PA, Michalski WP, Bingham J. Molecular pathogenesis of H5 highly pathogenic avian influenza: the role of the haemagglutinin cleavage site motif. Rev Med Virol. 2015;25:406–30. DOIPubMedGoogle Scholar
- Schrauwen EJ, Herfst S, Leijten LM, van Run P, Bestebroer TM, Linster M, et al. The multibasic cleavage site in H5N1 virus is critical for systemic spread along the olfactory and hematogenous routes in ferrets. J Virol. 2012;86:3975–84. DOIPubMedGoogle Scholar
- World Organisation for Animal Health and Food and Agriculture Organization Influenza. Influenza A cleavage sites [cited 2017 Feb 24]. http://www.offlu.net/fileadmin/home/en/resource-centre/pdf/Influenza_A_Cleavage_Sites.pdf
- Yang JR, Liu MT. Human infection caused by an avian influenza A (H7N9) virus with a polybasic cleavage site in Taiwan, 2017. J Formos Med Assoc. 2017;116:210–2. DOIPubMedGoogle Scholar
- Zhang F, Bi Y, Wang J, Wong G, Shi W, Hu F, et al. Human infections with recently-emerging highly pathogenic H7N9 avian influenza virus in China. J Infect. 2017;75:71–5. DOIPubMedGoogle Scholar
- Suguitan AL Jr, Matsuoka Y, Lau YF, Santos CP, Vogel L, Cheng LI, et al. The multibasic cleavage site of the hemagglutinin of highly pathogenic A/Vietnam/1203/2004 (H5N1) avian influenza virus acts as a virulence factor in a host-specific manner in mammals. J Virol. 2012;86:2706–14. DOIPubMedGoogle Scholar
- Zeng H, Pappas C, Belser JA, Houser KV, Zhong W, Wadford DA, et al. Human pulmonary microvascular endothelial cells support productive replication of highly pathogenic avian influenza viruses: possible involvement in the pathogenesis of human H5N1 virus infection. J Virol. 2012;86:667–78. DOIPubMedGoogle Scholar
- Gao HN, Lu HZ, Cao B, Du B, Shang H, Gan JH, et al. Clinical findings in 111 cases of influenza A (H7N9) virus infection. N Engl J Med. 2013;368:2277–85. DOIPubMedGoogle Scholar
- Hu Y, Lu S, Song Z, Wang W, Hao P, Li J, et al. Association between adverse clinical outcome in human disease caused by novel influenza A H7N9 virus and sustained viral shedding and emergence of antiviral resistance. Lancet. 2013;381:2273–9. DOIPubMedGoogle Scholar
- Yen HL, McKimm-Breschkin JL, Choy KT, Wong DD, Cheung PP, Zhou J, et al. Resistance to neuraminidase inhibitors conferred by an R292K mutation in a human influenza virus H7N9 isolate can be masked by a mixed R/K viral population. MBio. 2013;4:e00396-13. DOIPubMedGoogle Scholar
- Mok CK, Lee HH, Lestra M, Nicholls JM, Chan MC, Sia SF, et al. Amino acid substitutions in polymerase basic protein 2 gene contribute to the pathogenicity of the novel A/H7N9 influenza virus in mammalian hosts. J Virol. 2014;88:3568–76. DOIPubMedGoogle Scholar
- Neumann G, Kawaoka Y. Host range restriction and pathogenicity in the context of influenza pandemic. Emerg Infect Dis. 2006;12:881–6. DOIPubMedGoogle Scholar
- Yamayoshi S, Fukuyama S, Yamada S, Zhao D, Murakami S, Uraki R, et al. Amino acids substitutions in the PB2 protein of H7N9 influenza A viruses are important for virulence in mammalian hosts. Sci Rep. 2015;5:8039. DOIPubMedGoogle Scholar
1These authors contributed equally to this article.
2These authors contributed equally to this article.