Volume 24, Number 9—September 2018
Dispatch
Association of Batai Virus Infection and Encephalitis in Harbor Seals, Germany, 2016
Cite This Article
Citation for Media
Abstract
We isolated Batai virus from the brain of a euthanized, 26-year-old, captive harbor seal with meningoencephalomyelitis in Germany. We provide evidence that this orthobunyavirus can naturally infect the central nervous system of a mammal. The full-genome sequence showed differences from a previously reported virus isolate from a mosquito in Germany.
Batai virus (BATV) is a member of the Bunyamwera serogroup of orthobunyaviruses of the family Peribunyaviridae. Orthobunyaviruses are single-stranded, negative-sense RNA viruses with a tripartite genome composed of small, medium, and large segments, which encode nucleocapsid, glycoproteins, and polymerase, respectively (1). These segments can be interchanged between viruses of the same genus, resulting in stable reassortant bunyaviruses. For example, the Ngari virus genome consists of segments from BATV and Bunyamwera virus. Ngari virus is associated with outbreaks of hemorrhagic fever in humans and shows a clinical spectrum different from that of both parent viruses (2).
BATV has been documented to cause mild illness in ruminants and humans (3,4). Other hosts include domestic pigs and wild birds (3). BATV is transmitted mainly by Anopheles and Culex spp. mosquitoes and is widely distributed throughout Europe, Asia, and Africa (3).
In Germany, BATV was first detected in Anopheles maculipennis mosquitoes in 2009 (5). Enzootic transmission cycles involving domestic and wild mammals was reported in a serologic study in which 3 (0.55%) of 548 cattle had BATV-neutralizing serum antibodies (6). We report natural BATV infection of 2 captive harbor seals (Phoca vitulina), in Germany, in which meningoencephalomyelitis developed in 1 of them.
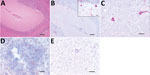
In September 2016, a 26-year-old male harbor seal (Phoca vitulina) in a zoo in northern Germany showed peracute deterioration of its general condition. Because of progression and severity of illness, the seal was euthanized and the carcass sent to the Department of Pathology, University of Veterinary Medicine Hannover (Hannover, Germany), for necroscopic analysis. Macroscopic examination showed signs of distress but no gross lesions were detected. Histologic analysis showed a mild to moderate, multifocal, perivascularly accentuated, lymphohistiocytic meningoencephalomyelitis, which affected the cerebrum, cerebellum (Figure 1, panel A), brain stem, medulla oblongata, and cervical spinal cord. Histologic analysis indicated a virus etiology.
Routine immunohistochemical tests of the seal brain for morbilliviruses, Borna disease virus, and tick-borne encephalitis virus (7,8) and immunofluorescence analysis for rabies virus were performed by the Department of Consumer and Food Safety of Lower Saxony (Hannover, Germany). All tests showed negative results.
We attempted virus isolation from homogenized brain in Vero cells. Cytopathic changes were observed within 3 days and continued to emerge in subsequent passages. To identify the pathogen, we investigated supernatant from the initial Vero cell isolation by using deep sequencing and a modified sequence-independent, single-primer amplification protocol as described (9,10). Analysis of raw reads with Bowtie 2 version 2.2.9 (https://sourceforge.net/projects/bowtie-bio/files/bowtie2/2.2.9/) for DNA mapping and Pauda version 1.0.1 (https://bioconda.github.io/recipes/pauda/README.html) for amino acid mapping identified BATV.
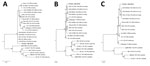
We created a reference assembly for all 3 genome segments (GenBank accession nos. S, MH299972; M, MH299973; and L, MH299974) by using CLC Genomics Workbench version 9.0 (QIAGEN, Hilden, Germany). The isolated virus was closely related to previously identified BATV strains from Europe (Figure 2) but had the highest sequence homology with strains from Russia (nucleotide pairwise identity S, 99%; M, 98.6%; and L, 98.5%).
We tested seal tissues (Table) for BATV by using real-time PCR and fluorescent in situ hybridization (FISH) as described (5,12). We used BATV-specific probe 5′-FAM-aacagtccagttccagacgatggtc-BHQ-1-3′ and primers Fwd-5′-Gctggaaggttactgtatttaatac-3′ and Rv-5′-Caaggaatccactgagtctgtg-3′ specific for the S segment (5). A BATV-specific probe for nucleotides 28–899 of the S segment was designed for FISH experiments (QuantiGene ViewRNA Kits; Affymetrix-Panomics, Santa Clara, CA, USA), which were performed according to the manufacturer’s protocol with minor modifications (12).
The highest virus load (by real-time PCR) was found in the central nervous system, and lesion-associated Purkinje cells and neurons of the granular cell layer of the cerebellum showed positive FISH results (Figure 1, panel B). A positive cytoplasmic result was also obtained for single spinal cord neurons (Figure 1, panel C). More limited BATV infection was found in peripheral organs, and the lowest cycle threshold was for the intestine. We also found BATV in single cells of the tunica mucosa of the small intestine and in cortical and medullary lymphocytes of the pulmonary lymph node by FISH (Figure 1, panel D). Other organs showed negative results in both assays.
We also performed histopathologic analysis of archived formalin-fixed paraffin-embedded (FFPE) organ samples of a seal that had shared the enclosure with the BATV-infected seal and had died 2 months before the euthanized seal showed the first clinical signs. Glomerular and tubular epithelial kidney cells (Technical Appendix Figure), cells of the tunica mucosa of the small intestine, and cortical and medullary lymphocytes of the pulmonary lymph node showed positive results for BATV by FISH. These FFPE samples did not show positive results by real-time PCR, probably because of low sensitivity of the assay for FFPE samples (13).
Retrospective analysis of FFPE brain samples of seals (n = 7) that had histopathologic changes suggestive of an unknown virus etiology and were isolated from harbor seals in coastal waters of Germany in the past decade all had negative results for BATV by FISH. Therefore, we screened 100 serum samples from harbor seals and 100 serum samples from gray seals (Halichoerus grypus) collected in 2016 and 2017 after admission to a seal rehabilitation center in the Netherlands that covers seal populations partially overlapping those of coastal waters of Germany. We neutralized isolated seal BATV (100 50% tissue culture infective doses) with diluted serum samples before application to reporter cells and examined for cytopathic effects after 3 days. However, no BATV antibodies (titer >1:20) were detected.
We isolated and characterized BATV from the brain of a captive harbor seal in Germany. This seal had lymphohistiocytic meningoencephalomyelitis and evidence of virus replication in Purkinje cells, neurons, enterocytes, and lymphocytes in peripheral tissues. Evidence of BATV infection by FISH was also obtained for a second harbor seal that had died 2 months before in the same enclosure. No additional evidence was found for seals as natural hosts for BATV infection by investigating brains from seals with encephalitis in coastal waters of Germany and by conducting a serosurvey among free-living harbor and gray seals. Results obtained from the 2 BATV infected animals indicated BATV circulation in the area during the mosquito season and that captive seals were possible dead-end hosts. Because seals in their natural environment are most likely less exposed to mosquitoes than seals in captivity, the observed seal BATV infections might be unnatural captivity-associated events. Phylogenetic analysis indicated that BATV isolated from the seal brain differed from BATV isolated from a mosquito in Germany and is more closely related to strains identified in Russia.
This study provides evidence of BATV associated with central nervous system disease in a naturally infected mammal. Other orthobunyaviruses have also been shown to cross the blood–brain barrier and show neurotropic properties (14). For example, a virus from the same serogroup, Bunyamwera virus, was recently associated with neurologic disease and abortion in horses (15). Moreover, possible human BATV infection in disease-endemic regions requires further investigation because BATV infection of mammals, including humans, has been reported in Europe (3) and Sudan (4).
Furthermore, BATV is the donor of the M segment of Ngari virus, which causes hemorrhagic fever in humans (2). The 2 BATV-infected seals could have been exceptionally sensitive to BATV infection because of predisposing factors, such as advanced age, concurrent conditions, genetic predisposition, or immunologic deficiencies. This possibility raises the question whether immunocompromised humans or other mammals might be at increased risk for development of neurologic BATV infection. Collectively, our data indicate the need for increased surveillance of BATV infection in mosquitoes, mammals, and birds in Europe.
Ms. Jo is a doctoral candidate at the University of Veterinary Medicine Hannover Research Center for Emerging Infections and Zoonoses, Hannover, Germany. Her research interests are virus discovery, emerging and reemerging infectious diseases, cross-species transmission of viruses, virus evolution, and host adaptation.
Acknowledgments
We thank M. Tieke, M. Schubert, H. Heidtmann, B. Buck, and P. Grünig for technical assistance; the Department of Consumer and Food Safety of Lower-Saxony, Hannover, for investigative work in searching for the causative agent; the Genomics Laboratory at the Institute of Animal Breeding and Genetics, Hannover, for performing analysis using the MiSeq system; and Joachim Schöne for collaborating in the study.
This study was supported by the Niedersachsen-Research Network on Neuroinfectiology (grant N-RENNT) from the Ministry of Science and Culture of Lower Saxony, Germany; and the European Union Horizon 2020 research and innovation program COMPARE (grant agreement no. 643476).
References
- Elliott RM. Orthobunyaviruses: recent genetic and structural insights. Nat Rev Microbiol. 2014;12:673–85. DOIPubMedGoogle Scholar
- Yanase T, Kato T, Yamakawa M, Takayoshi K, Nakamura K, Kokuba T, et al. Genetic characterization of Batai virus indicates a genomic reassortment between orthobunyaviruses in nature. Arch Virol. 2006;151:2253–60. DOIPubMedGoogle Scholar
- Hubálek Z. Mosquito-borne viruses in Europe. Parasitol Res. 2008;103(Suppl 1):S29–43. DOIPubMedGoogle Scholar
- Nashed NW, Olson JG, el-Tigani A. Isolation of Batai virus (Bunyaviridae:Bunyavirus) from the blood of suspected malaria patients in Sudan. Am J Trop Med Hyg. 1993;48:676–81. DOIPubMedGoogle Scholar
- Jöst H, Bialonski A, Schmetz C, Günther S, Becker N, Schmidt-Chanasit J. Isolation and phylogenetic analysis of Batai virus, Germany. Am J Trop Med Hyg. 2011;84:241–3. DOIPubMedGoogle Scholar
- Hofmann M, Wiethölter A, Blaha I, Jöst H, Heinemann P, Lehmann M, et al. Surveillance of Batai virus in bovines from Germany. Clin Vaccine Immunol. 2015;22:672–3. DOIPubMedGoogle Scholar
- Lempp C, Jungwirth N, Grilo ML, Reckendorf A, Ulrich A, van Neer A, et al. Pathological findings in the red fox (Vulpes vulpes), stone marten (Martes foina) and raccoon dog (Nyctereutes procyonoides), with special emphasis on infectious and zoonotic agents in Northern Germany. PLoS One. 2017;12:e0175469. DOIPubMedGoogle Scholar
- Uhde AK, Lehmbecker A, Baumgärtner W, Spitzbarth I. Evaluation of a panel of antibodies for the immunohistochemical identification of immune cells in paraffin-embedded lymphoid tissues of new- and old-world camelids. Vet Immunol Immunopathol. 2017;184:42–53. DOIPubMedGoogle Scholar
- Allander T, Emerson SU, Engle RE, Purcell RH, Bukh J. A virus discovery method incorporating DNase treatment and its application to the identification of two bovine parvovirus species. Proc Natl Acad Sci U S A. 2001;98:11609–14. DOIPubMedGoogle Scholar
- Endoh D, Mizutani T, Kirisawa R, Maki Y, Saito H, Kon Y, et al. Species-independent detection of RNA virus by representational difference analysis using non-ribosomal hexanucleotides for reverse transcription. Nucleic Acids Res. 2005;33:e65. DOIPubMedGoogle Scholar
- Huelsenbeck JP, Ronquist F. MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics. 2001;17:754–5. DOIPubMedGoogle Scholar
- Pfankuche VM, Bodewes R, Hahn K, Puff C, Beineke A, Habierski A, et al. Porcine bocavirus infection associated with encephalomyelitis in a Pig, Germany. Emerg Infect Dis. 2016;22:1310–2. DOIPubMedGoogle Scholar
- Bodewes R, Zohari S, Krog JS, Hall MD, Harder TC, Bestebroer TM, et al. Spatiotemporal analysis of the genetic diversity of seal influenza A(H10N7) virus, northwestern Europe. J Virol. 2016;90:4269–77. DOIPubMedGoogle Scholar
- Ludlow M, Kortekaas J, Herden C, Hoffmann B, Tappe D, Trebst C, et al. Neurotropic virus infections as the cause of immediate and delayed neuropathology. Acta Neuropathol. 2016;131:159–84. DOIPubMedGoogle Scholar
- Tauro LB, Rivarola ME, Lucca E, Mariño B, Mazzini R, Cardoso JF, et al. First isolation of Bunyamwera virus (Bunyaviridae family) from horses with neurological disease and an abortion in Argentina. Vet J. 2015;206:111–4. DOIPubMedGoogle Scholar
Figures
Table
Cite This ArticleOriginal Publication Date: July 25, 2018
1These authors contributed equally to this article.
2Current affiliation: Utrecht University, Utrecht, the Netherlands.
Table of Contents – Volume 24, Number 9—September 2018
| EID Search Options |
|---|
|
|
|
|
|
|

Please use the form below to submit correspondence to the authors or contact them at the following address:
Albert Osterhaus, Research Center for Emerging Infections and Zoonoses, University of Veterinary Medicine, Bünteweg 17, Hannover D-30559, Germany
Top