Bat Influenza A(HL18NL11) Virus in Fruit Bats, Brazil
Angélica Cristine Almeida Campos, Luiz Gustavo Bentim Góes, Andres Moreira-Soto, Cristiano de Carvalho, Guilherme Ambar, Anna-Lena Sander, Carlo Fischer, Adriana Ruckert da Rosa, Debora Cardoso de Oliveira, Ana Paula G. Kataoka, Wagner André Pedro, Luzia Fátima A. Martorelli, Luzia Helena Queiroz, Ariovaldo P. Cruz-Neto, Edison Luiz Durigon
1, and Jan Felix Drexler
1
Author affiliations: Charité-Universitätsmedizin Berlin, corporate member of Freie Universität Berlin, Humboldt-Universität zu Berlin, and Berlin Institute of Health, Institute of Virology, Berlin, Germany (A.C.A. Campos, L.G.B. Góes, A. Moreira-Soto, A.-L. Sander, C. Fischer, J.F. Drexler); Universidade de São Paulo-USP, Instituto de Ciências Biomédicas-ICB, São Paulo, Brazil (A.C.A. Campos, L.G.B. Góes, E.L. Durigon); Universidade Estadual Paulista Faculdade de Medicina Veterinária de Araçatuba, Araçatuba, Brazil (C. de Carvalho, W.A. Pedro, L.H. Queiroz); Universidade Estadual Paulista, Instituto de Biociências, Rio Claro, Brazil (G. Ambar, A.P. Cruz-Neto); Centro de Controle de Zoonoses, São Paulo (A.R. da Rosa, D.C. de Oliveira, L.F.A. Martorelli, A.P.G. Kataoka); German Centre for Infection Research, Germany (J.F. Drexler); Martsinovsky Institute of Medical Parasitology, Tropical and Vector-Borne Diseases, Sechenov University, Moscow, Russia (J.F. Drexler)
Main Article
Figure 2
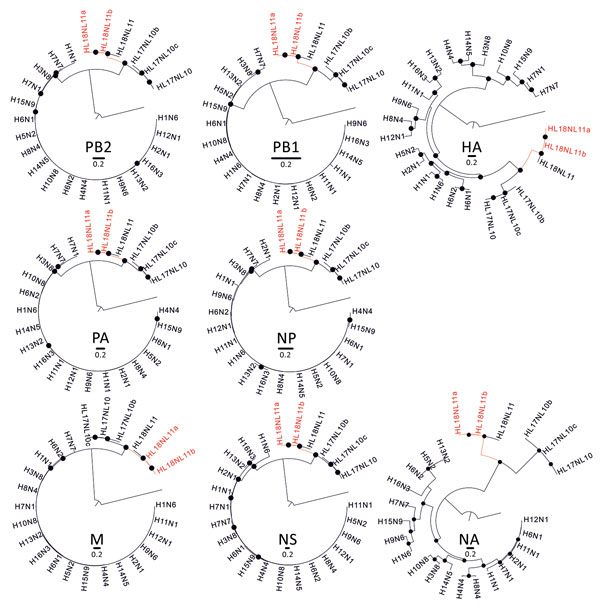
Figure 2. Phylogenetic relationships between bat influenza A viruses from Brazil and reference viruses. Phylogenetic trees show comparison of the 8 segments of representative influenza A virus genomes (PB2, PB1, PA, HA/HL, NP, NA/NL, M, NS) with A/great fruit-eating bat/Brazil/2301/2012 (HL18NL11a; GenBank accession nos. MH682200–7) and A/great fruit-eating bat/Brazil/2344/2012 (HL18NL11b; GenBank accession nos. MH682208–15), shown in red. Maximum-likelihood trees were inferred using a general time-reversible substitution model with a gamma distribution and invariant sites. Black dots represent bootstrap values >75% (1,000 replicates). Trees were generally rooted using influenza B/Lee/1940 (GenBank accession nos. DQ792894–901) (data not shown). Trees were constructed by using MEGA 6.0 (http://www.megasoftware.net). HA, hemagglutinin; M, matrix; NA, neuraminidase; NS1, nonstructural protein 1; NP, nucleoprotein; PA, polymerase acidic; PB, polymerase basic. Scale bars indicate nucleotide substitutions per site.
Main Article
Page created: January 18, 2019
Page updated: January 18, 2019
Page reviewed: January 18, 2019
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.
