Volume 25, Number 9—September 2019
Dispatch
Use of Human Intestinal Enteroids to Detect Human Norovirus Infectivity
Figure 2
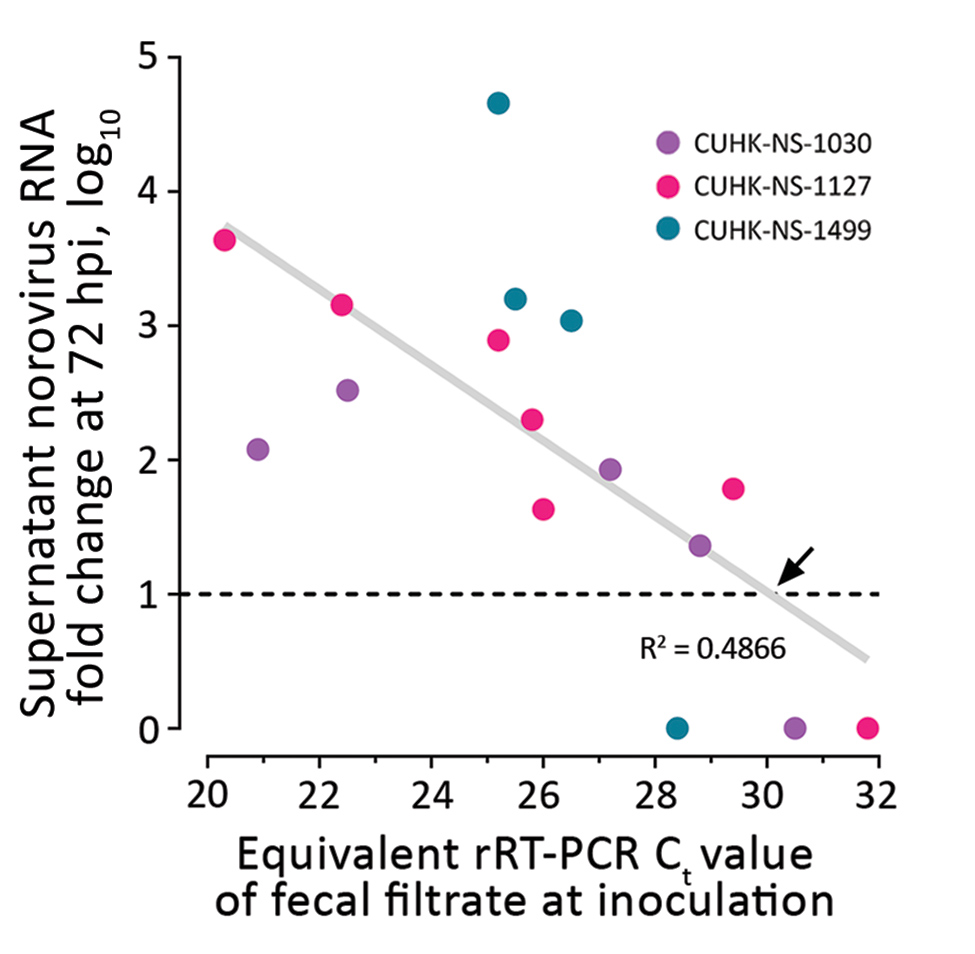
Figure 2. Estimation of a cutoff for rRT-PCR Ct value of inoculating fecal filtrate indicative of the ability to generate productive norovirus replication (i.e., containing infectious norovirus) in a human intestinal enteroid (HIE) line. We tested 3 strains of pandemic human norovirus genogroup II genotype 4 (GII.Pe-GII.4 Sydney) (CUHK-NS-1030, from a 1-year-old boy; CUHK-NS-1127, from a 79-year-old man; CUHK-NS-1499, from a 46-year-old man). We used 3-fold serial dilutions of norovirus-containing fecal filtrates to inoculate a monolayer culture of the adult stem cell–derived HIE line J2 seeded on 96-well cell culture plates. Productive norovirus replication was defined as having a supernatant viral RNA level increase of >10-fold at 72 hpi from baseline (1 hpi) (horizontal dotted line). RNA fold change data were derived from those shown in Figure 1 and were arbitrarily defined as 1 for those without observable viral RNA replication. For each strain, only the first dilution that resulted in undetectable viral RNA is shown and was included in downstream data analysis. The diagonal gray line denotes the best-fit line from linear regression, and the equation is Y = −0.2818X + 9.47, where Y represents RNA fold change and X represents Ct value. The black arrow specifies a Cf cutoff of 30.1 of inoculating fecal filtrate that yields productive norovirus replication as estimated from regression analysis. Ct, cycle threshold; CUHK, Chinese University of Hong Kong; hpi, hours postinfection; rRT-PCR, real-time reverse transcription PCR.