Volume 26, Number 10—October 2020
Dispatch
Drug Resistance Spread in 6 Metropolitan Regions, Germany, 2001–20181
Cite This Article
Citation for Media
Abstract
We analyzed 1,397 HIV-1 pol sequences of antiretroviral therapy–naive patients in a total of 7 university hospitals in Bonn, Cologne, Frankfurt, Hamburg, Hannover, and Munich, Germany. Phylogenetic and network analysis elucidated numerous cases of shared drug resistance mutations among genetically linked patients; K103N was the most frequently shared mutation.
The use of antiretroviral therapy (ART) has shown markedly decreased sickness and death rates in persons living with HIV (PLWH) (1–3). Meanwhile, the emergence of antimicrobial drug resistance in HIV-1 is raising public health concerns (4,5). Nationwide estimates of the prevalence of drug resistance mutations (DRMs) are not available in Germany (6); the reported prevalence of transmitted HIV-1 DRMs differ across regions and risk groups from 10.4%–17.2%, as described in 2 cohort studies from the German ClinSurv-HIV cohort and the Cologne-Bonn cohort (6,7).
Information about the dynamics and patterns of HIV transmission within defined areas and communities remains incomplete. Thus, we combined phylogenetic analysis with clinical and sociodemographic data, to determine the spread and dynamics of HIV-1 DRMs in 6 metropolitan regions in Germany, including the cities with the highest rates of new HIV-1 infections in 2017: Munich (17.3/100,000 population), Cologne (13.3/100,000 population), and Frankfurt (12.3/100,000 population), (8). We conducted this retrospective study in a cooperative effort of partner sites of the Translational Platform HIV (TP-HIV) (Cologne, Germany) and the University of California, San Diego (San Diego, CA, USA). The study was approved by the local ethics committees of the university hospitals of Bonn, Cologne, Munich, Hannover, Frankfurt, and Hamburg. All study participants gave written informed consent.
We analyzed HIV-1 partial pol sequences (HXB2 position 2550–3356), obtained as part of clinical routine care, and sociodemographic data of PLWH who received HIV care at the university hospitals of Bonn, Cologne, Frankfurt, Hamburg, and Hannover and at 2 hospitals in Munich during 2001–2018. Patients could participate in the study if they had recently received their diagnosis of HIV-1 and were ART naive; this conservative approach excluded participants for whom the exact start date of ART or history of prior ART was not accurately documented.
Blood samples were collected before ART initiation. We sequenced partial HIV-1 pol region as previously described (7,9). We set the mixed mutation calling threshold at >10%, consistent with Sanger sequencing sensitivity (10). We identified major DRMs by using the Stanford University Genotypic Resistance Interpretation HVdb version 8.9, (https://hivdb.stanford.edu). We inferred the genetic transmission network as previously described (7,9,11); we inferred putative linkage for genetic distances <1.5% (12) (Appendix).
We performed statistical analyses by using Stata version 14 (StataCorp LP, https://www.stata.com). We applied Fisher exact or χ2 test and univariable and multivariable logistic regression models, as appropriate, to determine characteristics that are associated with shared DRM and clustering. A shared DRM was defined as any DRM present in >2 genetically linked persons.
Overall, 1,397 HIV-1 infected participants were included. Most were male (82.9%; 1,158/1,397), originated from Germany (69.6%; 972/1,397), and infected with HIV-1 subtype B (72.8%; 1,017/1,397). The most commonly reported transmission risk was men who have sex with men (MSM) (56.7%; 792/1,397) (Table 1).
We identified an overall prevalence of any DRM at the time of diagnosis, excluding polymorphic mutations, of 17.8% (95% CI 15.7%–19.8%), 248/1,397 participants. The proportion varied significantly (p<0.001) by region, ranging from 9.1% (95% CI 5%–13%; 21/231) in Munich, up to 31.4% (95% CI 24%–38%; 53/169) in Hannover. Resistance mutations associated with nucleoside reverse transcriptase inhibitors (NRTIs) (172/1,397; 12.3%) were most frequent, followed by nonnucleoside reverse transcriptase inhibitors (NNRTIs) (124/1,397; 8.9%). The most common single mutations related to NNRTIs were K103N (31/124; 25.0%), and G190A (8/124; 6.5%). Of the NRTI resistance mutations, M41L (25/172; 14.5%), and T215S (18/172; 10.5%) were most frequently observed (Table 2).
Transmission network analyses revealed that 20.7% (289/1,397) of participants had a putative linkage forming 102 transmission clusters. The largest cluster included 12 participants, mostly MSM from Bonn, Cologne, Munich, and Frankfurt (Figure 1, panels A, B). Participants <25 years and 25–45 years of age were significantly more likely to cluster compared with participants >45 years (<25 years, adjusted OR [aOR] 4.38, 95% CI 2.55–7.52, p<0.001; 25–45 years, aOR 1.91, 95% CI 1.36–2.678; p<0.001). Participants infected with HIV-1 subtype B were more likely to cluster than those with non-B subtype (aOR 4.05, 95% CI 2.37–6.90; p<0.001). Geospatial distribution differed; participants from Bonn were linked significantly more often than those from Cologne (aOR 1.63; 95% CI 1.06–2.49; p = 0.025), even though the cities are geographically close (Appendix Table).
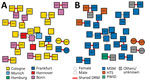
The prevalence of transmitted DRM was comparable in clustering (47/289, 16.3%) and nonclustering (201/1,108; 18.1%) participants (p = 0.46) (Appendix Table). Of the 47 sequences harboring DRM, 19 (40.4%) were preferentially shared by participants living predominantly in Cologne (14/19, 73.7%) and Bonn (3/19, 15.8%) (Figure 2, panel A) and by participants reporting MSM as main risk factor (15/19; 78.9%) (Figure 2, panel B). Younger age (<25 years) was associated with a higher proportion of shared DRM (3/11; 3.5%) compared with older age (24–45 years, 13/856 [1.5%]; >45 years, 2/430 [0.5%]) (Table 1).
The most frequently observed putatively shared DRM was K103N, detected in 9/19 (47.4%) participants forming 4 distinct clusters, predominantly originating from Cologne (7/9, 77.8%). The second most common shared DRM was D67N, found in 6/19 (31.6%) participants from Cologne and Bonn.
The increasing prevalence of DRMs in PLWH has become a serious matter of concern for clinicians and public health entities (4). In our study, we observed a 17.8% prevalence of DRMs, higher than in previous studies (6,7). The proportion of NNRTI resistance mutations was 8.9%, which is potentially associated with the common use of NNRTI in first-line ART regimens. K103N represented one quarter of NNRTI resistance mutations, reducing susceptibility to the first-generation drugs nevirapine and efavirenz (13). Transmission network analyses revealed that K103N was the most frequently shared DRM.
K65R, K70RT, and M184IV were the most common of the NRTI resistance mutations we observed, particularly among the risk group of MSM living in Cologne and Hannover, indicating potential resistance to preexposure prophylaxis (PrEP) with tenofovir/emtricitabine. Such resistance might be an upcoming challenge as PrEP use increases. Monitoring for HIV infections with these mutations is of utmost importance for preventing an epidemic among high-risk PrEP users; one mitigation is to consider alternative PrEP regimens in regions with high resistance.
Our study had several limitations. First, our sample population could have been biased because participants were not randomly selected; our dataset was limited to ART-naive patients who received an HIV diagnosis at 7 university hospitals during 2001–2018. Although we know no reason why a university hospital setting would not be representative of the region, it is possible that populations treated outside these centers may have different transmission networks and risks; results are not generalizable to the entire regions or nationwide. Second, mixing of heterosexual patients and MSM in clusters may be due to missing single or multiple risk factors. Thus, their role could not be represented in the transmission networks. Third, we have not tested clinical correlates and drug resistance; the clinical relevance was inferred from the Stanford database.
In summary, we found that the overall rate of DRM was high in Germany. Network analysis elucidated cases of shared DRMs among genetically linked persons, mainly in MSM-dominated clusters. Our findings highlight regional differences and illustrate the need to test MSM, especially younger men, for HIV regularly and to evaluate local HIV programs and adapt screening and treatment strategies to local epidemics.
Dr. Stecher is an epidemiologist in the Department for Infectious Diseases at the University Hospital of Cologne, Germany. Her primary research interests are infectious diseases, HIV epidemiology, and cohort studies.
Acknowledgments
This work was supported by funds from the German Center for Infection Research (DZIF) (grant no. NCT02149004] and supported by the NIH grants (grant nos. AI036214, MH113477, and AI106039). M.H. received grant funding from Gilead.
The study was approved by the local ethics committees of the university hospitals of Bonn (reference no. 279-14), Cologne (reference no. 13-364), Munich (reference no. 438-14), Hannover (reference no. 279-14), Frankfurt (reference no. 279-14), and Hamburg (reference no. 279-14).
The datasets used and analyzed during the current study are available from the corresponding author on reasonable request.
References
- Glass TR, Sterne JA, Schneider MP, De Geest S, Nicca D, Furrer H, et al.; Swiss HIV Cohort Study. Self-reported nonadherence to antiretroviral therapy as a predictor of viral failure and mortality. AIDS. 2015;29:2195–200. DOIPubMedGoogle Scholar
- Saag MS, Benson CA, Gandhi RT, Hoy JF, Landovitz RJ, Mugavero MJ, et al. Antiretroviral drugs for treatment and prevention of HIV infection in adults: 2018 recommendations of the international antiviral society-USA panel. JAMA. 2018;320:379–96. DOIPubMedGoogle Scholar
- Robert Koch-Institut. Epidemiologisches bulletin 47 [in German]. 2018 [cited 2020 Jul 31]. https://www.rki.de/DE/Content/Infekt/EpidBull/Archiv/2018/Ausgaben/47_18.pdf
- Wittkop L, Günthard HF, de Wolf F, Dunn D, Cozzi-Lepri A, de Luca A, et al.; EuroCoord-CHAIN study group. Effect of transmitted drug resistance on virological and immunological response to initial combination antiretroviral therapy for HIV (EuroCoord-CHAIN joint project): a European multicohort study. Lancet Infect Dis. 2011;11:363–71. DOIPubMedGoogle Scholar
- Little SJ, Holte S, Routy JP, Daar ES, Markowitz M, Collier AC, et al. Antiretroviral-drug resistance among patients recently infected with HIV. N Engl J Med. 2002;347:385–94. DOIPubMedGoogle Scholar
- Schmidt D, Kollan C, Fätkenheuer G, Schülter E, Stellbrink HJ, Noah C, et al.; ClinSurv-HIV Drug Resistance Study Group in CHAIN. Estimating trends in the proportion of transmitted and acquired HIV drug resistance in a long term observational cohort in Germany. PLoS One. 2014;9:
e104474 . DOIPubMedGoogle Scholar - Stecher M, Chaillon A, Eis-Hubinger AM, Lehmann C, Fatkenheuer G, Wasmuth JC, et al. Pretreatment human immunodeficiency virus type 1 (HIV-1) drug resistance in transmission clusters of the Cologne-Bonn region, Germany. Clin Microbiol Infect. 2019 Feb;25:253.e1–253.e4.
- Robert Koch-Institut. Epidemiologisches Bulletin 39 [in German]. 2017 [cited 2020 Jul 31]. http://www.rki.de/DE/Content/Infekt/EpidBull/Archiv/2017/Ausgaben/39_17.pdf
- Stecher M, Chaillon A, Eberle J, Behrens GMN, Eis-Hübinger AM, Lehmann C, et al. Molecular epidemiology of the HIV epidemic in three German metropolitan regions—Cologne/Bonn, Munich, and Hannover, 1999–2016. Sci Rep. 2018;8:6799. DOIPubMedGoogle Scholar
- Döring M, Büch J, Friedrich G, Pironti A, Kalaghatgi P, Knops E, et al. geno2pheno[ngs-freq]: a genotypic interpretation system for identifying viral drug resistance using next-generation sequencing data. Nucleic Acids Res. 2018;46(W1):W271–7. DOIPubMedGoogle Scholar
- Kosakovsky Pond SL, Weaver S, Leigh Brown AJ, Wertheim JO. HIV-TRACE (TRAnsmission Cluster Engine): a tool for large scale molecular epidemiology of HIV-1 and other rapidly evolving pathogens. Mol Biol Evol. 2018;35:1812–9. DOIPubMedGoogle Scholar
- Wertheim JO, Leigh Brown AJ, Hepler NL, Mehta SR, Richman DD, Smith DM, et al. The global transmission network of HIV-1. J Infect Dis. 2014;209:304–13. DOIPubMedGoogle Scholar
- European AIDS Clinical Society (EACS). European guidelines for treatment of HIV-positive adults in Europe 9.1. 2018 [cited 2020 Jul 31]. http://www.eacsociety.org/files/2018_guidelines-9.1-english.pdf
Figures
Tables
Cite This ArticleOriginal Publication Date: September 03, 2020
1Preliminary results from this study were presented at the Conference for Retroviruses and Opportunistic Infections (CROI) 2019, March 4–7, 2019, Seattle, Washington, USA.
2These first authors contributed equally to this article.
Table of Contents – Volume 26, Number 10—October 2020
| EID Search Options |
|---|
|
|
|
|
|
|

Please use the form below to submit correspondence to the authors or contact them at the following address:
Martin Hoenigl, Division of Infectious Diseases and Global Public Health, Department of Medicine, University of California, San Diego, 200 West Arbor Dr #8208, San Diego, CA 92103, USA; ; Melanie Stecher, Department I for Internal Medicine, University Hospital of Cologne, Herderstraße 52-54, 50931 Köln, Germany
Top