Volume 26, Number 11—November 2020
Research Letter
Seroprevalence of SARS-CoV-2 and Infection Fatality Ratio, Orleans and Jefferson Parishes, Louisiana, USA, May 2020
Cite This Article
Citation for Media
Abstract
Using a novel recruitment method and paired molecular and antibody testing for severe acute respiratory syndrome coronavirus 2 infection, we determined seroprevalence in a racially diverse municipality in Louisiana, USA. Infections were highly variable by ZIP code and differed by race/ethnicity. Overall census-weighted seroprevalence was 6.9%, and the calculated infection fatality ratio was 1.61%.
Seroprevalence studies around the world have estimated the spread of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) to range from 1.79% (1) in Boise, Idaho, USA, to 25% in Breves, Brazil (P. Hallal, unpub. data, https://doi.org/10.1101/2020.05.30.20117531). Coronavirus disease (COVID-19) has also been reported to disproportionately affect Black patients, but we do not know the infection fatality ratio (IFR), which requires knowing how many persons are at risk (i.e., infected). We estimated SARS-CoV-2 infections in Orleans and Jefferson Parishes, Louisiana, USA, and determined the COVID-19–related IFR by race.
The protocol was approved by the Ochsner Clinic Foundation Institutional Review Board (New Orleans, LA, USA) and designed to enroll and test up to 3,000 persons at 10 sites during May 9–15, 2020. To recruit a representative sample for this high-throughput method, a novel 2-step system developed by Public Democracy (https://www.publicdemocracy.io) considered >50 characteristics, including social determinants of health and US Census population data, to establish a pool of potential participants reflective of the demographics of the parishes, from which a randomized subset of 150,000 was selected. Of these, >25,000 volunteers were recruited through dynamic, cross-device digital advertisements, supplemented by television advertisements and a call-in number to register (Appendix). This volunteer pool was stratified by the same attributes and then randomly issued a text message inviting them to private testing locations. Invitations were adjusted daily on the basis of response rates to achieve a representative sample. Volunteers checked in with a digital code to discourage unsolicited walk-ins. We did not turn uninvited persons away but excluded them from analysis if they did not fit criteria. Housemates of participants (n = 234) or persons from ineligible ZIP codes (n = 34) were excluded. Six people withdrew consent. All study materials were created in English, Spanish, and Vietnamese. Participants were offered free transportation if needed. Verbal consent was electronically documented, and participants were asked a short list of questions followed by a blood draw and nasopharyngeal swab.
Tests approved by the US Food and Drug Administration’s Emergency Use Authorization were used. Real-time reverse transcription PCR tests of nasopharyngeal swabs were performed on the Abbott m2000 RealTime System (Abbott, https://www.abbott.com) and qualitative IgG blood tests on the ARCHITECT i2000SR (Abbott). The IgG test meets criteria described by the Centers for Disease Control and Prevention as yielding high positive predictive value, which was validated by a laboratory at Ochsner Health and others (1,2). Study participants for whom either or both tests were positive were considered to be infected with SARS-CoV-2.
US Census values, weighted by race and parish of residence, were divided by the total sample for exposure (a PCR-positive test, an IgG-positive test, or both), point prevalence (PCR-positive only), and seroprevalence (IgG-positive tests regardless of PCR test result). The positive-testing population included persons with early-stage infections (PCR-positive only) and persons recovering (PCR-positive and IgG-positive) and recovered (IgG-positive only). Early-stage infections were excluded from IFR estimation because their outcomes would not yet be registered as deaths. Therefore, weighted seroprevalence was used to calculate persons presumed to be recovered (3). IFR was calculated by dividing cumulative deaths by race (4) by the number of persons presumed to be recovered. Methodology and symptoms observed have been described elsewhere (A. Feehan, unpub. data, https://ssrn.com/abstract=3633166).
Among the 2,640 persons in the sample, 63.5% were female and 60.9% were White; average age was 50.6 years, and average household size was 2.55 persons. Among the 183 participants who tested positive, 49% were Black. The unadjusted exposure rate of SARS-CoV-2 in the sample population was 6.9% (7.8%, census-weighted); 0.9% were positive for active viral shedding but had no detectable antibody. By race, seroprevalence was highest (9.8%) in Black participants, followed by multiracial (7.1%), Asian (5.5%), and White (4.5%) participants. Hispanic participants had 5.3% seroprevalence. We multiplied 2018 population estimates by weighted seroprevalence to generate the number of persons presumed to be recovered (Table). Reported deaths (4) were divided by number of persons presumed to be recovered plus deaths to calculate the IFR, which was 1.61% overall. The IFR was statistically similar for White (1.55%), Black (1.69%), and multiracial (1.38%) persons but was significantly lower for Asian persons (0.61%). No COVID-19–related data on Hispanic persons were collected by the Louisiana Department of Public Health during the study period.
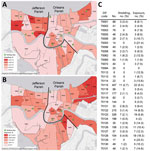
The prevalence of viral shedding (PCR-positive) and overall SARS-CoV-2 exposure (PCR-positive, IgG-positive, or both) were listed and mapped by ZIP code across the 2 parishes (Figure). Prevalence was highly variable across the map and in some areas exceeded 20%.
Prevalence studies help to understand infection spread, especially when testing resources are limited. Our study found the overall SARS-CoV-2 exposure rate in this area to be 7.8% and confirmed a recent report of overrepresentation of Black persons with COVID-19 in the New Orleans area (5). Multiracial, Hispanic, and Asian persons also had higher seroprevalence than White persons. The overall IFR was 1.63%, which is higher than IFRs found in other seroprevalence studies (0.5%–1.2%) (6; M. Emmenegger, unpub. data, https://doi.org/10.1101/2020.05.31.20118554; P. Hallal, unpub. data, https://doi.org/10.1101/2020.05.30.20117531). The similar IFR among most racial groups indicates that viral spread at least partially explains the increased number of deaths among minority populations.
Dr. Feehan is a research scientist at the Ochsner Clinic Foundation’s Infectious Disease Clinical Research Department. Her research focuses on the gut microbiome as a treatment modality for neurologic disease, but more immediately on the SARS-CoV-2 pandemic that has greatly impacted the New Orleans area.
Acknowledgments
The authors would like to especially thank the laboratories at the Ochsner Medical Center Jefferson Highway Campus for testing and keeping track of research samples; Dan Nichols; Sarah Roberts and Gina Mmahat for clinical site management; Samantha Bright, Lyndsey Buckner-Baiamonte, and Ansley Hammons for research site management; Emily Arata for liaising with public leaders; and countless research coordinators, clinical staff, marketing personnel, medical students, and Epic and IT staff for making site testing possible. The Ochsner Health Market Planning and Analysis team designed the maps in Figure. The authors thank Kathleen McFadden for her thorough editing and Mark Roberts for his review and the Ochsner Language Services Department for helping to increase inclusivity. We would also like to acknowledge the New Orleans Mayor’s Office, City of New Orleans Office of Public Health, and the New Orleans City Council, especially council members Helena Moreno and Jason Williams for filming a public service announcement to help recruit participants. We also thank Jefferson Parish President Cynthia Lee Sheng and Parish Council for their support.
We thank George Hutter and ReNOLA for their financial support.
References
- Bryan A, Pepper G, Wener MH, Fink SL, Morishima C, Chaudhary A, et al. Performance characteristics of the Abbott Architect SARS-CoV-2 IgG assay and seroprevalence in Boise, Idaho. J Clin Microbiol. 2020;58:e00941–20. DOIPubMedGoogle Scholar
- Centers for Disease Control and Prevention. Interim guidelines for COVID-19 antibody testing. 2020 [cited 2020 May 15]. https://www.cdc.gov/coronavirus/2019-ncov/lab/resources/antibody-tests-guidelines.html
- Wilson N, Kvalsvig A, Barnard LT, Baker MG. Case-fatality risk estimates for COVID-19 calculated by using a lag time for fatality. Emerg Infect Dis. 2020;26:1339–441. DOIPubMedGoogle Scholar
- Louisiana Department of Public Health. COVID-19. 2020 [cited 2020 May 16]. http://ldh.la.gov/coronavirus
- Price-Haywood EG, Burton J, Fort D, Seoane L. Hospitalization and mortality among Black patients and white patients with Covid-19. N Engl J Med. 2020;382:2534–43. DOIPubMedGoogle Scholar
- Pollán M, Pérez-Gómez B, Pastor-Barriuso R, Oteo J, Hernán MA, Pérez-Olmeda M, et al.; ENE-COVID Study Group. Prevalence of SARS-CoV-2 in Spain (ENE-COVID): a nationwide, population-based seroepidemiological study. [Epub ahead pf print]. Lancet. 2020 Jul 3 Epub ahead of print].
Figure
Table
Cite This ArticleOriginal Publication Date: July 30, 2020
Table of Contents – Volume 26, Number 11—November 2020
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Amy K. Feehan, Ochsner Health, 1st floor AT, Infectious Diseases, 1514 Jefferson Hwy, New Orleans, LA 70121, USA
Top