Volume 26, Number 7—July 2020
Dispatch
Possible Bat Origin of Severe Acute Respiratory Syndrome Coronavirus 2
Figure 2
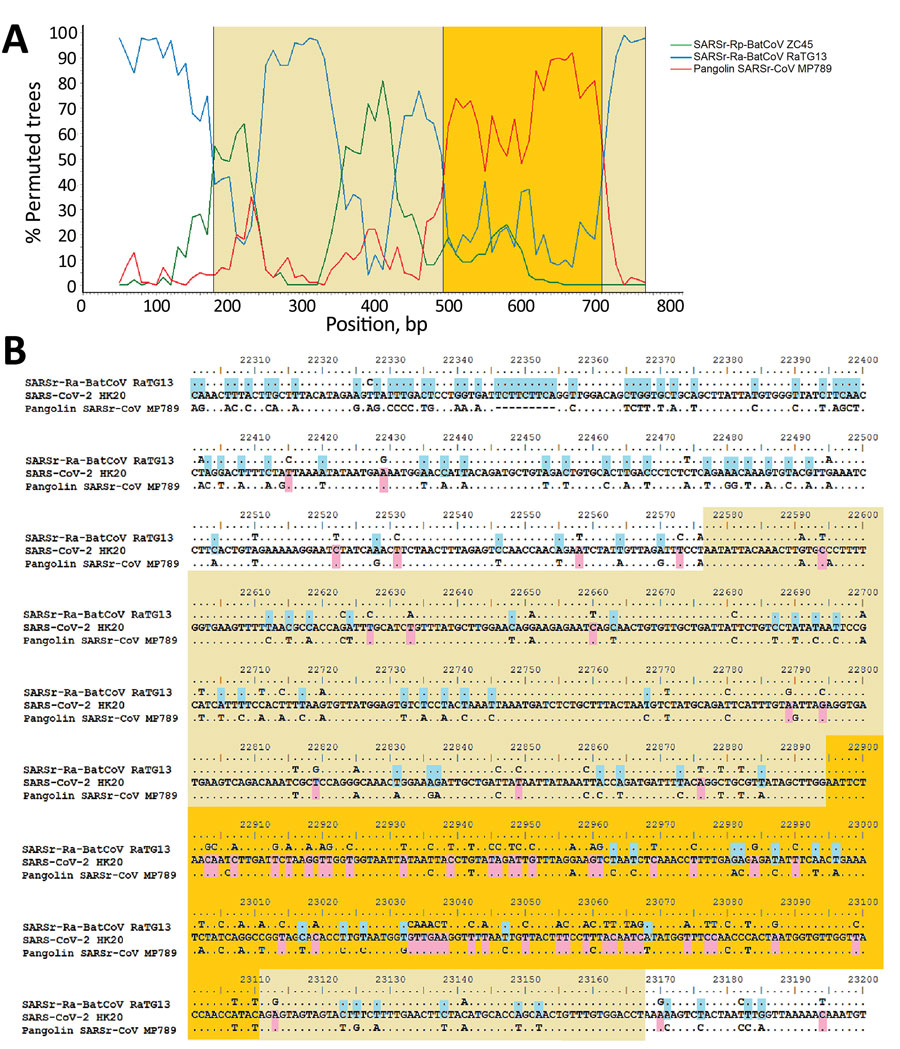
Figure 2. Bootscan analysis and nucleotide sequence alignment for SARS-CoV-2 isolates and closely related viruses. A) Boot scan analysis using the partial spike gene (positions 22397–23167) of SARS-CoV-2 strain HK20 as query sequence. Bootscanning was conducted with Simplot version 3.5.1 (https://sray.med.som) (F84 model; window size, 100 bp; step, 10 bp) on nucleotide alignment, generated with ClustalX (http://www.clustal.org). B) Multiple alignment of nucleotide sequences from genome positions 22300 to 23700. Yellow indicates receptor binding domain; orange indicates receptor binding motif; pink indicates bases conserved between SARS-CoV-2 HK20 and Pangolin-SARSr-CoV/MP789/Guangdong/2019; and blue indicates bases conserved between SARS-CoV-2 HK20 and SARSr-Ra-BatCoVs RaTG13.
1These authors contributed equally to this article.