Volume 27, Number 2—February 2021
Research
Outbreak of Severe Vomiting in Dogs Associated with a Canine Enteric Coronavirus, United Kingdom
Cite This Article
Citation for Media
Abstract
The lack of population health surveillance for companion animal populations leaves them vulnerable to the effects of novel diseases without means of early detection. We present evidence on the effectiveness of a system that enabled early detection and rapid response to a canine gastroenteritis outbreak in the United Kingdom. In January 2020, prolific vomiting among dogs was sporadically reported in the United Kingdom. Electronic health records from a nationwide sentinel network of veterinary practices confirmed a significant increase in dogs with signs of gastroenteric disease. Male dogs and dogs living with other vomiting dogs were more likely to be affected. Diet and vaccination status were not associated with the disease; however, a canine enteric coronavirus was significantly associated with illness. The system we describe potentially fills a gap in surveillance in neglected populations and could provide a blueprint for other countries.
Population health data is lacking for companion animals such as dogs, cats, and rabbits, leaving a surveillance gap for endemic diseases and delayed detection of incursions of disease, such as equine influenza virus (H3N8) (1), avian influenza (H3N2) (2,3), and parvoviruses (3). In the absence of legislated programs of population surveillance, several attempts have been made to fill this gap using secondary data, particularly from pet insurance providers (4). More recently, researchers have exploited the rapid digitization of electronic health records (EHRs) for passive surveillance. Data can be collected at great scale and analyzed in near–real time. EHR data are now routinely used in human heath efforts (5–8), in which their timeliness, simplicity, and breadth of coverage complements surveillance based on diagnostic data (9,10). Such approaches are beginning to find healthcare value in veterinary species, especially among companion animals (4,11–13), a high proportion of which visit veterinarians (14).
In January 2020, one of the authors of this article (D.G.), a primary care veterinarian in northwest England, contacted the other authors about seeing an unusually high number of cases (≈40) of severe vomiting in dogs; responses to a social media post suggested other veterinarians might have been experiencing similar events. Vomiting is a common complaint among dogs whose owners seek treatment for them (15,16). However, documented outbreaks are rare because established vaccines are available for most common known pathogens (17). In the absence of robust populationwide data, such sporadic reports frequently do not raise awareness of outbreaks.
For the response we describe, we obtained data from syndromic surveillance and text mining of EHRs collected from sentinel veterinary practices and diagnostic laboratories, which we then linked with data from field epidemiology and enhanced genomic testing. In 8 weeks, using this approach, we described the temporal and spatial epidemiology, identified a possible causative agent, and provided targeted advice to control the outbreak. Ethics approval was given by Liverpool University Research Ethics Committees (Liverpool, UK; VREC922/RETH000964).
Data Sources
Veterinary Practices
During March 17, 2014–February 29, 2020, we collected data from 7,094,397 consultation records (4,685,732 from dogs and 1,846,493 from cats) from EHRs from the Small Animal Veterinary Surveillance Network (SAVSNET), a volunteer network of 301 veterinary practices (663 sites) in the United Kingdom, recruited based on convenience (11). In brief, EHRs included data collected during individual consultations on species, breed, sex, neuter status, age, owners’ postcodes, and vaccination status. Each EHR is also compulsorily annotated by the veterinary clinician with a main presenting complaint (MPC) at time of visit, using a questionnaire window embedded in the practice management system. Options for reasons for visit included gastroenteric, respiratory, pruritus, tumor, kidney disease, other unwell, post-op check, vaccination, or other healthy.
Given that severe vomiting was a key outbreak feature, we undertook 2 complementary analyses. First, we used regular expressions to identify clinical narratives describing frequent vomiting, but excluded common false positive search results (Appendix Table 1). Second, we used data on prescriptions to describe the frequency of all veterinary-authorized products containing the antiemetic maropitant (18). We calculated trend lines using Bayesian binomial generalized linear modeling trained on weekly prevalence during 2014–2019 (19), which allowed us to identify extreme (>99% credible interval [CrI]) or moderate (>95% CrI) observations.
Laboratories
SAVSNET also collects EHRs from participating diagnostic laboratories on samples submitted from more than half of UK veterinary practices. Canine diagnostic test results from January 2017 through February 2020 were queried from 6 laboratories for 6 gastroenteric pathogens. Test numbers, percentage of positive results, and associated 95% CIs were summarized (Table 1). The number of sites was surmised from the submitting practices’ postcodes.
Questionnaires
Online questionnaires to enable case reporting were made available to both veterinarians and owners beginning January 29, 2020. The required case definition of >5 vomiting episodes in a 12-hour period was based on clinical observations of early cases. Veterinarians were also asked to complete control questionnaires. Initially, we requested only controls matched to veterinary practices contributing case data; however, to increase recruitment, a nonmatched control questionnaire open to any veterinarian was deployed on February 5. The questionnaires (Appendix) requested a range of information including owner postcode, animal signalment, vaccination status, clinical signs, treatment and diagnostic testing, animal contacts, diet, and recovery status.
We performed all statistical analyses using R version 3.6.1 (https://cran.r-project.org). Case details were described for both veterinarian- and owner-reported data. We calculated proportions and 95% CIs for categorical variables and median and range for continuous variables. We constructed univariable and multivariable mixed-effects logistic regression models using data submitted by veterinarians using R package lme4. Explanatory variables from univariable logistic regression were considered in multivariable models for likelihood ratios of p≤0.20, which underwent manual stepwise backward elimination to reduce Akaike’s and Bayesian information criteria. Practice was included as a random effect. We assessed confounding by the effect on model fit with sequential removal of variables and assessed 2-way interaction terms for improved model fit. We defined final statistical significance as p <0.05.
Spatiotemporal Analysis of Cases
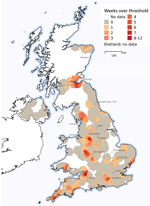
We obtained records of consults weekly during November 4, 2019–March 21, 2020; cases were geolocated by pet owners’ postcodes. We considered records of gastroenteric MPC as a binary outcome (i.e., 1 for gastroenteric consult, 0 for nongastroenteric consult). We used a logistic geostatistical model to investigate spatial clustering of cases for each week. We defined a spatial hotspot as a location having 95% posterior probability of prevalence exceeding the national mean prevalence over any 1-week period. With no discernible epidemic wave apparent over successive weeks, we aggregated weekly measures across the study period to show the number of weeks each location was a hotspot (Appendix).
Sample Collection, PCR, and Phylogenetic Analyses
Veterinarians submitting questionnaires were also asked to submit samples for microbiological testing including mouth swabs, fecal samples, and for gastrointestinal cases, vomit. In brief, we extracted nucleic acids using a QIAGEN QIAamp viral RNA kit (https://www.qiagen.com), reverse transcribed samples using ThermoFisher Superscript III (https://www.thermofisher.com), and tested for canine enteric coronavirus (CeCoV) by M-gene PCR (20). To expedite results and reduce contamination risks, the PCR was run as a single-stage PCR rather than as the published nested reaction. We purified positive samples using QIAquick (QIAGEN) and sequenced them bidirectionally (Sanger sequencing; Source Biosciences, https://www.sourcebioscience.com) to produce consensus sequences (ChromasPro 2.1.8, http://technelysium.com.au).
To rapidly explore the potential involvement of other viruses, we extracted nucleic acid from 19 random cases and 5 controls for deep sequencing. RNA was amplified by sequence-independent, single-primer-amplification (21), multiplexed libraries were prepared using 30 ng of cDNA with an Oxford Nanopore SQK-LSK109 ligation sequencing kit (Oxford Nanopore, https://nanoporetech.com) and sequenced using an Oxford Nanopore MinION Mk1B device for 48 hours. To perform real-time fast basecalling, we used the Oxford Nanopore MinKNOW Guppy toolkit and FASTQ files uploaded to an Oxford Nanopore EPI2ME data analysis platform for identification.
For deeper sequencing coverage, we also processed 10 samples (6 CeCoV-positive cases, 3 negative cases, 1 control) for Illumina sequencing at the University of Liverpool Centre for Genomic Research (https://www.liverpool.ac.uk/genomic-research). We treated nucleic acids with RNase and prepared fragment libraries using a NEBNext Ultra II kit (https://www.neb.com) before performing paired-end, 2 × 150–bp sequencing on an Illumina HiSeq 4000 system (https://www.illumina.com). Adaptor sequences were trimmed using cutadapt (https://cutadapt.readthedocs.io) and sickle (https://github.com), with a minimum quality score of 20. Reads >19 bp matching the dog genome (CanFam3.1, http://genome.ucsc.edu) using Bowtie2 sequence alignment tool (http://bowtie-bio.sourceforge.net) were removed. Remaining reads were assembled using the SPAdes toolkit (https://github.com) and contigs >700 nt blasted against the NCBI RefSeq nonredundant proteins database (https://www.ncbi.nlm.nih.gov/refseq). Sequences matching CeCoV were aligned using the ClustalW multiple sequence alignment program (https://www.genome.jp) and phylogenies reconstructed using bootstrap analyses and neighbor-joining in MEGA6 software (https://www.megasoftware.net). Each sequence was assigned a local laboratory number based on the order in which the sequences were analyzed.
Syndromic Surveillance
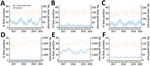
On the basis of MPCs identified in the EHRs, we found a specific and significant increase in the number of dogs recorded as exhibiting gastroenteric signs; the final 10 weeks, during December 2019–March 2020, were outside the 99% CrI (extreme outliers; Figure 1, panel A). A similar trend was observed in maropitant therapy for dogs (Figure 1, panel B). Both measures, peaked in the week ending February 2, 2020, at approximately double the preceding baseline. We observed no similar trends for respiratory disease in dogs, for gastroenteric MPCs, for maropitant treatment in cats (Figure 1, panels C–E), or for antibiotic use in dogs (data not shown), together suggesting the signal was specific to canine gastroenteric disease, a finding supported by similar increases in the regular expression identifying vomiting dogs (Figure 1, panel F).
Spatiotemporal mapping of weekly cases of gastroenteric MPC showed prevalence was spatially clustered (Figure 2). In particular, locations in northwest and southwest England and in Edinburgh, Scotland, had strong evidence of many weeks of prevalence higher than the national mean.
Diagnostic Tests
The patterns of test results for different PCR tests, generally carried out concurrently, were broadly similar (Figure 3, panels A–C). The same was true for results based on cultured samples (Figure 3, panels D, E). Of particular interest, CeCoV showed strong seasonality, positive tests peaking during the winter months (Figure 3, panel A). However, similar peaks seen in previous years suggested the observed peak in February 2020 could not itself explain this outbreak.
Questionnaire
By March 1, 2020, a total of 1,258 case questionnaires had been received. After excluding 59 questionnaires missing key data, we used data from 165 veterinary-reported cases, 1,034 owner-reported cases (Table 2), and 60 veterinary-reported controls (Appendix Table 2) for analyses.
Most cases were from households in England (Table 2). Median case age at examination was 4.0 years (range 0.3–15.0 years) based on veterinary reports and 4.8 years (range 0.2–15.5 years) based on owner reports. Most animals had been vaccinated against core pathogens (17) and leptospirosis within the preceding 3 years and dewormed within the previous 3 months. A range of breeds (data not presented) were observed, broadly corresponding to previous studies (6). Most cases were fed dog food, with ≈20%–37% of dogs scavenged food when walked. Of those from multidog households, just over half reported the presence of another dog recently vomiting within the same household. Around 30% of dogs had recently traveled, most commonly visiting a daycare facility.
Date of onset of clinical signs ranged from November 16, 2019, through February 28, 2020, for veterinary-reported cases, and September 4, 2019, through March 1, 2020, for owner-reported cases. Most cases involved inappetence (75.6%–86.1%) and vomiting without blood (88.7%–91.5%) (Table 3). Approximately half of cases reported diarrhea, most without blood. Diagnostic testing was performed in 32.1% of veterinary-reported cases, most (78.9%) using hematology or biochemistry assays, or both.
Dogs in >90% of veterinary-reported cases were treated, compared with in 61.7% of owner-reported cases. In both, antiemetics were most often prescribed: in 89.1% (CrI 84.3%–93.9%) of veterinary-reported cases and in 48.1% (CrI 45.0%–51.1%) of owner-reported cases. The most common recovery time was 3–7 days; the dogs died in 0.6% of veterinary-reported and 1.0% of owner-reported cases.
Descriptive data about the control population, submitted by veterinarians, and univariable findings from analyses of the veterinary case controls are presented in Appendix Tables 2 and 3; multivariable findings are shown in Table 4. Both neutered and nonneutered male dogs were at significantly increased odds of contracting the illness, compared with neutered females, as were dogs living in the same household as another dog that had also been vomiting compared to those in households where other dogs were healthy. However, dogs living in a single-dog household were at increased odds of contracting the illness compared with dogs living in the same household as another dog that had not recently vomited. Dogs that had been in recent contact with another animal species (including humans) that had recently vomited were at reduced odds of vomiting, compared with those who had not. Other potential causes considered early in the outbreak, including foodborne etiologies, vaccine preventable diseases, or the possibility of interspecies transmission, were not significantly associated (Appendix Table 3).
Sampling and Molecular Testing
During January 30–March 12, 2020, we collected a total of 95 samples from 71 animals (50 cases, 21 controls): 22 from feces, 60 from oral swabs, and 13 from vomit. Dogs with prolific vomiting were significantly more likely to test positive for CeCoV in >1 sample (17/50, 34%) compared with controls (0/21) (p = 0.002 by Fisher exact test). Positive test results were most likely in samples from feces (10/16 [62.5%] cases, 0/6 controls; p = 0.01) and vomit (6/13 [46%] cases, 0 controls). Samples from oral swabs were least likely to test positive (7/43 [16%] cases, 0/17 controls; p = 0.17). Of 17 CeCoV-positive cases, 12 met the case definition, 2 did not (<5 episodes of vomiting in 12 hours), and 3 lacked questionnaire data.
We gathered useable M-gene sequences from 21 samples (16 dogs). When we sequenced 2 samples from the same animal, the sequences were identical and subsequently represented only once in analyses (Figure 4). All sequences clustered with previously reported type II CeCoVs (22) in 1 of 3 lineages. Sequences from 14 of 16 dogs were identical, suggesting a single outbreak strain geographically distributed across England. Sequences from dogs 15 and 16 were phylogenetically distinct.
Results of MinION sequencing rapidly confirmed an alphacoronavirus as the predominant virus (24,190 out of 33,826,933 reads) and failed to identify any other prevalent candidates (next highest, betabaculovirus: 4,541 reads). Although bacterial reads were present in high numbers, none showed consistently high results across most samples.
Complete CeCoV genomes were assembled from 6 PCR-positive cases by Illumina sequencing. We identified no coronavirus sequences in 3 cases and 1 control that tested negative for CeCoV by PCR. The only other mammalian virus sequence detected matched a canine rotavirus (1 case, 1 control; data not presented). Consistent with M-gene sequencing, 5 of the CeCoV genomes clustered together (>99% similarity), distinct from the genome from dog 15 (Figure 4). The outbreak strain was most similar to a virus from Taiwan isolated in 2008 from a young dog with diarrhea (94.5% similarity; L. Chueh, pers. comm. [email] Apr. 27, 2020) and did not show any obvious sequence differences to published strains that might explain the unusual pattern of disease observed in the outbreak. Based on spike gene analyses, the outbreak strain clustered with IIb, having a TGEV-like N-terminal spike domain (23). Sequences were submitted to GenBank (accession nos. MT877072, MT906864, and MT906865).
Using EHRs annotated with syndromic information by veterinarians, we rapidly identified an outbreak of canine gastroenteric disease that had started in November 2019. This finding was corroborated by parallel increases in relevant prescriptions and records of frequent vomiting. These data were augmented by data from responses to a questionnaire, diagnostic laboratories, and enhanced microbiological analyses. This system enabled us to determine case definitions and outcomes and to identify risk factors as well as a potential viral cause, within a 3-month period; findings were rapidly disseminated to veterinarians (24,25) and owners. This combined approach represents an efficient system that can fill a previously neglected national population health surveillance need for companion animals.
The first indication of an outbreak came from time-series analyses of syndromic data. Such syndromic surveillance is increasingly being used to monitor the impact of national events like natural disasters and bioterrorism on human population health, as well as changes in gastroenteric and influenza-like illness (6–9). Such data can be simple to collect, provide real-time wide geographic coverage, and be flexibly applied to different conditions (10,11). Although in some cases these data can identify outbreaks earlier than more active surveillance, their predictive value can sometimes be low, particularly where there is a low signal to noise complaint ratio. In our case, the outbreak was large compared with background levels, associated with near doubling of the gastroenteric syndrome, and had many weeks in which the syndrome statistically exceeded the baseline.
The richness of data within EHRs enabled us to validate this outbreak using numbers of antiemetic prescriptions and text mining. Prescription data have been used to understand, for example, human health inequalities (26), and the use of critical antimicrobials in both humans (27) and animals (28,29). We used these data to identify and track an outbreak, benefitting from a clear link between the syndrome (vomiting) and its therapy (antiemetic). It will be useful to identify other disease-therapy associations that could be used for similar surveillance.
We used text mining to identify records of frequent vomiting in clinical narratives. Such approaches can circumvent the need for practitioner-derived annotation and be flexibly and rapidly adapted to emerging syndromes as soon as case-definitions are determined. Similar approaches have been described in human health for conditions such as fever (30–32) but can suffer low sensitivity (31). Indeed, the outbreak peak based on text mining was ≈20% of that based on MPC analysis. However, it is also likely the outbreak as defined by the MPC included a considerable number of animals with milder signs that would not be detected by data mining using the regular expression developed here. Although data from text mining are unlikely to give an accurate estimate of the true prevalence of a given condition, they can still be used to track outbreaks.
To compliment syndromic surveillance, we implemented a rapid case-control study, collecting >1,200 responses from veterinarians and owners in 4.5 weeks. There was no evidence for similar disease in people or other species. The timing of the outbreak as shown by case data was in broad agreement with our syndromic surveillance. Questionnaires from owners and veterinarians were in broad agreement on date of onset, geographic density, clinical signs, and recovery. These data informed targeted health messages posted online and on social media on February 28, 2020, 4 weeks after we first became aware of the outbreak.
Clearly, evidence of transmission driving the outbreak was vital to providing disease control advice. Dogs in multidog households were more likely to vomit if other dogs in the household were also affected, suggesting either transmission between dogs or a common environmental source; these observations informed advice to the public around isolating affected dogs. Of note, dogs in single-dog households were also at increased odds of being affected compared to multidog households where only a single dog was vomiting. Some authors have shown that dogs from single-dog households are walked more and therefore could be at greater risk for infection (33). Factors affecting dog walking are clearly likely to be important for control of infectious disease transmission and should be explored further.
In addition to collecting epidemiologic data, we collected microbiological samples from cases and controls. Based on its known (34) and observed seasonality (Figure 3, panel A), we tested all samples for CeCoV. Cases were significantly more likely to show positive results both when all samples (oral swabs, feces and vomit) were considered or when just fecal samples were considered, suggesting a possible role for CeCoV in the outbreak. However, many case samples tested negative: 33 of 50 overall, 6 of 16 dogs for which feces samples were submitted, and 7 of 13 dogs for which vomit samples were submitted. There are several potential reasons for these negative findings, including the sensitivity of the PCR, the high numbers of oral swabs (although simpler to collect, oral swabs were more likely to test negative), the timing of samples in relation to viral shedding, and the storage and transport of samples. In addition, it is important to note that our case definition, based as it was on a syndrome and lacking more specific confirmatory testing, is likely to include some animals that were not part of the outbreak. Indeed, at its peak, the outbreak only doubled the background level of gastroenteric disease seen at other times of the year; therefore, we might expect only half of our cases to be truly associated with the outbreak.
Sequencing results identified a predominant CeCoV strain in outbreak cases across the United Kingdom, in contrast with earlier studies showing that CeCoV strains tend to cluster in households, veterinary practices, or local areas (35). This finding lends further support to the role of this strain in the observed outbreak. In Sweden, a single strain was also implicated in several small wintertime canine vomiting outbreaks (36); genetically, however, the virus strain we identified was distinct from the strain from Sweden (data not shown). Ultimately, it will be necessary to perform a challenge study to confirm or refute the role of this CeCoV strain as the cause of this outbreak, as well as to explore the range of clinical signs associated with infection.
If this strain is proven to be the cause of the outbreak, several features mark the observed pattern of disease as unusual, including the outbreak scale, its geographic distribution, the severity of signs in some animals, a lack of notable viral co-infections, and the involvement of adult dogs. CeCoV is generally associated with mild gastroenteritis (37). Although sporadic outbreaks of more severe hemorrhagic diseases with high mortality (38–40), as well as systemic diseases (41,42), have been reported, these typically affect individual households, and are often associated with mixed infections (43). Such observations suggest that the genetic variability of CeCoVs may affect virulence and are supported by experimental infections recreating more severe disease (38). The genetic mechanism underlying such shifts in virulence in CeCoV have not been defined. However, mutations impacting virulence are described in closely related alphacoronaviruses (44–47).
In conclusion, this multidisciplinary approach enabled a rapid response to a newly described outbreak of canine gastroenteritis and identified a CeCoV as a potential cause. Previous CeCoV seasonality suggests further outbreaks may occur. Having such an efficient surveillance system provides the ideal platform to inform and target population health messaging. Several challenges remain for addressing the lack of national population health structures for companion animals: to systematically capture discussions of disease in social and mainstream media; to sustainably fund these activities, which currently are largely resourced by research grants; to understand and broaden the representativeness of such sentinel networks; and to link surveillance information with agencies empowered to act (12).
Dr. Radford is a professor of veterinary health informatics at the University of Liverpool. His primary research interests are the molecular epidemiology of viral pathogens, particularly those of veterinary importance, and combining this subject with electronic health data to study animal diseases at a population level and their impact on people.
Acknowledgments
We thank data providers both in veterinary practice (VetSolutions, Teleos, CVS Group, and independent practitioners) and participating veterinary diagnostic laboratories (Axiom Veterinary Laboratories, Batt Laboratories, BioBest, Idexx, NationWide Laboratories Microbiology Diagnostics Laboratory at the University of Liverpool, the Department of Pathology and Infectious Diseases at the University of Surrey, and the Veterinary Pathology Group), without whose support and participation this research would not have been possible. We are especially grateful for the help and support provided by SAVSNET team members Susan Bolan and Steven Smyth.
This work was funded in part by the Dogs Trust as part of SAVSNET-Agile, and by the Biotechnology and Biological Sciences Research Council, and previously by the British Small Animal Veterinary Association.
References
- Crawford PC, Dubovi EJ, Castleman WL, Stephenson I, Gibbs EP, Chen L, et al. Transmission of equine influenza virus to dogs. Science. 2005;310:482–5. DOIPubMedGoogle Scholar
- Li G, Wang R, Zhang C, Wang S, He W, Zhang J, et al. Genetic and evolutionary analysis of emerging H3N2 canine influenza virus. Emerg Microbes Infect. 2018;7:73. DOIPubMedGoogle Scholar
- Allison AB, Kohler DJ, Fox KA, Brown JD, Gerhold RW, Shearn-Bochsler VI, et al. Frequent cross-species transmission of parvoviruses among diverse carnivore hosts. J Virol. 2013;87:2342–7. DOIPubMedGoogle Scholar
- O’Neill DG, Church DB, McGreevy PD, Thomson PC, Brodbelt DC. Approaches to canine health surveillance. Canine Genet Epidemiol. 2014;1:2. DOIPubMedGoogle Scholar
- Smith S, Elliot AJ, Mallaghan C, Modha D, Hippisley-Cox J, Large S, et al. Value of syndromic surveillance in monitoring a focal waterborne outbreak due to an unusual Cryptosporidium genotype in Northamptonshire, United Kingdom, June - July 2008. Euro Surveill. 2010;15:19643. DOIPubMedGoogle Scholar
- Fleming DM, Elliot AJ. Lessons from 40 years’ surveillance of influenza in England and Wales. Epidemiol Infect. 2008;136:866–75. DOIPubMedGoogle Scholar
- Hiller KM, Stoneking L, Min A, Rhodes SM. Syndromic surveillance for influenza in the emergency department-A systematic review. PLoS One. 2013;8:
e73832 . DOIPubMedGoogle Scholar - Elliot AJ, Singh N, Loveridge P, Harcourt S, Smith S, Pnaiser R, et al. Syndromic surveillance to assess the potential public health impact of the Icelandic volcanic ash plume across the United Kingdom, April 2010. Euro Surveill. 2010;15:19583.PubMedGoogle Scholar
- Thomas MJ, Yoon PW, Collins JM, Davidson AJ, Mac Kenzie WR. Evaluation of syndromic surveillance systems in 6 US state and local health departments. J Public Health Manag Pract. 2018;24:235–40.PubMedGoogle Scholar
- Smith GE, Elliot AJ, Lake I, Edeghere O, Morbey R, Catchpole M, et al.; Public Health England Real-time Syndromic Surveillance Team. Syndromic surveillance: two decades experience of sustainable systems - its people not just data! Epidemiol Infect. 2019;147:
e101 . DOIPubMedGoogle Scholar - Sánchez-Vizcaíno F, Noble PM, Jones PH, Menacere T, Buchan I, Reynolds S, et al. Demographics of dogs, cats, and rabbits attending veterinary practices in Great Britain as recorded in their electronic health records. BMC Vet Res. 2017;13:218. DOIPubMedGoogle Scholar
- McGreevy P, Thomson P, Dhand NK, Raubenheimer D, Masters S, Mansfield CS, et al. VetCompass Australia: a national big data collection system for veterinary science. Animals (Basel). 2017;7:74. DOIPubMedGoogle Scholar
- O’Neill DG, Church DB, McGreevy PD, Thomson PC, Brodbelt DC. Prevalence of disorders recorded in cats attending primary-care veterinary practices in England. Vet J. 2014;202:286–91. DOIPubMedGoogle Scholar
- Asher L, Buckland EL, Phylactopoulos CI, Whiting MC, Abeyesinghe SM, Wathes CM. Estimation of the number and demographics of companion dogs in the UK. BMC Vet Res. 2011;7:74. DOIPubMedGoogle Scholar
- Elwood C, Devauchelle P, Elliott J, Freiche V, German AJ, Gualtieri M, et al. Emesis in dogs: a review. J Small Anim Pract. 2010;51:4–22. DOIPubMedGoogle Scholar
- O Neill DG, Church DB, McGreevy PD, Thomson PC, Brodbelt DC. Prevalence of disorders recorded in dogs attending primary-care veterinary practices in England. PLoS One. 2014;9:
e90501 . DOIPubMedGoogle Scholar - Day MJ, Horzinek MC, Schultz RD, Squires RA; Vaccination Guidelines Group (VGG) of the World Small Animal Veterinary Association (WSAVA). WSAVA Guidelines for the vaccination of dogs and cats. J Small Anim Pract. 2016;57:E1–45. DOIPubMedGoogle Scholar
- Singleton DA, Sánchez-Vizcaíno F, Arsevska E, Dawson S, Jones PH, Noble PJM, et al. New approaches to pharmacosurveillance for monitoring prescription frequency, diversity, and co-prescription in a large sentinel network of companion animal veterinary practices in the United Kingdom, 2014-2016. Prev Vet Med. 2018;159:153–61. DOIPubMedGoogle Scholar
- Arsevska E, Singleton DA, Jewell C, Paterson S, Jones PH, Smyth S, et al. Small animal disease surveillance: pruritus and Pseudomonas skin infections. Vet Rec. 2018;183:182–7. DOIPubMedGoogle Scholar
- Pratelli A, Tempesta M, Greco G, Martella V, Buonavoglia C. Development of a nested PCR assay for the detection of canine coronavirus. J Virol Methods. 1999;80:11–5. DOIPubMedGoogle Scholar
- Chrzastek K, Lee DH, Smith D, Sharma P, Suarez DL, Pantin-Jackwood M, et al. Use of Sequence-Independent, Single-Primer-Amplification (SISPA) for rapid detection, identification, and characterization of avian RNA viruses. Virology. 2017;509:159–66. DOIPubMedGoogle Scholar
- Pratelli A, Martella V, Decaro N, Tinelli A, Camero M, Cirone F, et al. Genetic diversity of a canine coronavirus detected in pups with diarrhoea in Italy. J Virol Methods. 2003;110:9–17. DOIPubMedGoogle Scholar
- Decaro N, Mari V, Elia G, Addie DD, Camero M, Lucente MS, et al. Recombinant canine coronaviruses in dogs, Europe. Emerg Infect Dis. 2010;16:41–7. DOIPubMedGoogle Scholar
- Smith SL, Singleton DA, Noble PJ, Radford AD, Brant B, Pinchbeck GL, et al. Possible cause of outbreak of prolific vomiting in dogs. Vet Rec. 2020;186:324. DOIPubMedGoogle Scholar
- Singleton DA, Noble PJ, Radford AD, Brant B, Pinchbeck GL, Greenberg D, et al. Prolific vomiting in dogs. Vet Rec. 2020;186:191. DOIPubMedGoogle Scholar
- Rowlingson B, Lawson E, Taylor B, Diggle PJ. Mapping English GP prescribing data: a tool for monitoring health-service inequalities. BMJ Open. 2013;3:
e001363 . DOIPubMedGoogle Scholar - Zanichelli V, Monnier AA, Gyssens IC, Adriaenssens N, Versporten A, Pulcini C, et al. Variation in antibiotic use among and within different settings: a systematic review. J Antimicrob Chemother. 2018;73(suppl_6):vi17–29. DOIPubMedGoogle Scholar
- Singleton DA, Sánchez-Vizcaíno F, Dawson S, Jones PH, Noble PJM, Pinchbeck GL, et al. Patterns of antimicrobial agent prescription in a sentinel population of canine and feline veterinary practices in the United Kingdom. Vet J. 2017;224:18–24. DOIPubMedGoogle Scholar
- Hur BA, Hardefeldt LY, Verspoor KM, Baldwin T, Gilkerson JR. Describing the antimicrobial usage patterns of companion animal veterinary practices; free text analysis of more than 4.4 million consultation records. PLoS One. 2020;15:
e0230049 . DOIPubMedGoogle Scholar - South BR, Chapman WW, Delisle S, Shen S, Kalp E, Perl T, et al. Optimizing A syndromic surveillance text classifier for influenza-like illness: Does document source matter? AMIA Annu Symp Proc. 2008;2008:692–6.PubMedGoogle Scholar
- Haas SW, Travers D, Waller A, Mahalingam D, Crouch J, Schwartz TA, et al. Emergency Medical Text Classifier: New system improves processing and classification of triage notes. Online J Public Health Inform. 2014;6:
e178 . DOIPubMedGoogle Scholar - Chapman WW, Dowling JN, Wagner MM. Classification of emergency department chief complaints into 7 syndromes: a retrospective analysis of 527,228 patients. Ann Emerg Med. 2005;46:445–55. DOIPubMedGoogle Scholar
- Westgarth C, Christian HE, Christley RM. Factors associated with daily walking of dogs. BMC Vet Res. 2015;11:116. DOIPubMedGoogle Scholar
- Duijvestijn M, Mughini-Gras L, Schuurman N, Schijf W, Wagenaar JA, Egberink H. Enteropathogen infections in canine puppies: (Co-)occurrence, clinical relevance and risk factors. Vet Microbiol. 2016;195:115–22. DOIPubMedGoogle Scholar
- Stavisky J, Pinchbeck GL, German AJ, Dawson S, Gaskell RM, Ryvar R, et al. Prevalence of canine enteric coronavirus in a cross-sectional survey of dogs presenting at veterinary practices. Vet Microbiol. 2010;140:18–24. DOIPubMedGoogle Scholar
- Escutenaire S, Isaksson M, Renström LH, Klingeborn B, Buonavoglia C, Berg M, et al. Characterization of divergent and atypical canine coronaviruses from Sweden. Arch Virol. 2007;152:1507–14. DOIPubMedGoogle Scholar
- Decaro N, Buonavoglia C. An update on canine coronaviruses: viral evolution and pathobiology. Vet Microbiol. 2008;132:221–34. DOIPubMedGoogle Scholar
- Buonavoglia C, Decaro N, Martella V, Elia G, Campolo M, Desario C, et al. Canine coronavirus highly pathogenic for dogs. Emerg Infect Dis. 2006;12:492–4. DOIPubMedGoogle Scholar
- Tennant BJ, Gaskell RM, Jones RC, Gaskell CJ. Studies on the epizootiology of canine coronavirus. Vet Rec. 1993;132:7–11. DOIPubMedGoogle Scholar
- Evermann JF, Abbott JR, Han S. Canine coronavirus-associated puppy mortality without evidence of concurrent canine parvovirus infection. J Vet Diagn Invest. 2005;17:610–4. DOIPubMedGoogle Scholar
- Decaro N, Campolo M, Lorusso A, Desario C, Mari V, Colaianni ML, et al. Experimental infection of dogs with a novel strain of canine coronavirus causing systemic disease and lymphopenia. Vet Microbiol. 2008;128:253–60. DOIPubMedGoogle Scholar
- Zicola A, Jolly S, Mathijs E, Ziant D, Decaro N, Mari V, et al. Fatal outbreaks in dogs associated with pantropic canine coronavirus in France and Belgium. J Small Anim Pract. 2012;53:297–300. DOIPubMedGoogle Scholar
- Dowgier G, Lorusso E, Decaro N, Desario C, Mari V, Lucente MS, et al. A molecular survey for selected viral enteropathogens revealed a limited role of Canine circovirus in the development of canine acute gastroenteritis. Vet Microbiol. 2017;204:54–8. DOIPubMedGoogle Scholar
- Porter E, Tasker S, Day MJ, Harley R, Kipar A, Siddell SG, et al. Amino acid changes in the spike protein of feline coronavirus correlate with systemic spread of virus from the intestine and not with feline infectious peritonitis. Vet Res (Faisalabad). 2014;45:49. DOIPubMedGoogle Scholar
- Chang HW, Egberink HF, Halpin R, Spiro DJ, Rottier PJ. Spike protein fusion peptide and feline coronavirus virulence. Emerg Infect Dis. 2012;18:1089–95. DOIPubMedGoogle Scholar
- Licitra BN, Millet JK, Regan AD, Hamilton BS, Rinaldi VD, Duhamel GE, et al. Mutation in spike protein cleavage site and pathogenesis of feline coronavirus. Emerg Infect Dis. 2013;19:1066–73. DOIPubMedGoogle Scholar
- Zhang X, Hasoksuz M, Spiro D, Halpin R, Wang S, Stollar S, et al. Complete genomic sequences, a key residue in the spike protein and deletions in nonstructural protein 3b of US strains of the virulent and attenuated coronaviruses, transmissible gastroenteritis virus and porcine respiratory coronavirus. Virology. 2007;358:424–35. DOIPubMedGoogle Scholar
Figures
Tables
Cite This ArticleOriginal Publication Date: January 13, 2021
Table of Contents – Volume 27, Number 2—February 2021
| EID Search Options |
|---|
|
|
|
|
|
|
![Observed prevalence of main presenting complaint (MPC) and maropitant use in cats and dogs, per 1,000 consultations, in investigation of dogs with vomiting, United Kingdom, January 2017–February 2020. A) Canine records labeled as gastroenteric MPC; B) canine records in which maropitant was prescribed; C) canine records labeled as respiratory MPC; D) feline records in which maropitant was prescribed; E) feline records labeled as gastroenteric MPC; and F) frequent vomiting in dogs based on regular expression searches of the clinical narratives. Red points represent the extreme outliers (outside the 99% credible interval [CrI]), orange points the moderate outliers (outside the 95% CrI, but within the 99% CrI), and green points the average trend (within the 95% CrI).](/eid/images/20-2452-F1-tn.jpg)


Please use the form below to submit correspondence to the authors or contact them at the following address:
Alan Radford, University of Liverpool, Leahurst Campus, Chester High Road, Neston, S. Wirral, CH64 7TE, UK; email;alanrad@liverpool.ac.uk
Top