Volume 27, Number 7—July 2021
Dispatch
Emergence of SARS-CoV-2 B.1.1.7 Lineage at Outpatient Testing Site, Berlin, Germany, January–March 2021
Cite This Article
Citation for Media
Abstract
Within 5 weeks in 2021, B.1.1.7 became the dominant severe acute respiratory syndrome coronavirus 2 lineage at an outpatient testing site in Berlin, Germany. Compared with outpatients with wild-type virus infection, patients with B.1.1.7 had similar cycle threshold values, more frequent sore throat and travel history, and less frequent anosmia/ageusia.
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) B.1.1.7 lineage (variant of concern [VOC] 202012/01 or 20I/501Y.V1) likely emerged during autumn 2020 in the United Kingdom and quickly became dominant there (E. Volz et al., unpub. data, https://www.medrxiv.org/content/10.1101/2020.12.30.20249034v2). B.1.1.7 carries multiple mutations and deletions, including 501Y and deletion ΔH69/ΔV70 (del69–70) in the spike protein. B.1.1.7 reportedly exhibits greater transmissibility and fatality in the community than non-VOC lineages (hereafter referred to as wild-type virus) (1; E. Volz et al., unpub. data). However, increased deaths were not seen in hospitalized patients (2).
The first patient infected with B.1.1.7 at the outpatient SARS-CoV-2 testing site of Charité–Universitätsmedizin Berlin was identified on January 18, 2021. We describe lineage prevalence over time and demographic and clinical characteristics in outpatients with B.1.1.7 or wild-type virus who sought care during January–March 2021. Ethics approval was obtained from Charité’s Institutional Review Board (EA4/083/20).
Details of the testing site have been described (3). Physicians interviewed patients about demographics, medical history, and symptoms. If indicated, a combined oro-nasopharyngeal swab specimen was collected. Specimens were tested by using the cobas 6800/8800 assay (Roche Diagnostics, https://diagnostics.roche.com), targeting open reading frame 1ab and the envelope gene (4). All positive samples were typed for the N501Y and del69–70 polymorphisms by melting curve analysis. Variants including both polymorphisms were considered B.1.1.7.
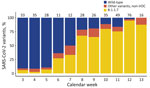
During January 18–March 29, 2021, a total of 349 SARS-CoV-2–positive patients were seen at the testing site, and the proportion of B.1.1.7 increased from 2% to >90% (Figure 1). In total, 35.8% (125/349) of samples belonged to the wild-type lineage, 57.0% (199/349) were B.1.1.7, and 7.2% (25/349) were other non–wild-type variants (non-VOCs).
Six patients previously had received >1 SARS-CoV-2 vaccinations; all were infected with B.1.1.7 but were excluded from analysis because vaccination could interfere with viral dynamics and clinical manifestation (Appendix Table). We excluded patients carrying lineages other than wild-type or B.1.1.7. Half of the patients were female (49%); mean age was 36 (SD +15) years. Almost all (97%) reported symptoms. Median symptom duration until testing was 3 (interquartile range 2–4) days. Symptoms were fatigue (72%), headache (69%), and muscle ache (60%). Fifteen percent reported travel outside Berlin in the previous 14 days, and half (49%) reported contact with a SARS-CoV-2–positive person (Table).
Most assessed characteristics did not substantially differ between patients with wild-type or B.1.1.7, including age, sex, leading symptoms, symptom duration, contact with a SARS-CoV-2–positive person, and time passed since contact (Table). However, B.1.1.7 patients had traveled more frequently than those with the wild-type strain (19% vs. 10%). Patients with B.1.1.7 more often reported sore throat than did patients with wild-type virus (54% vs. 42%) but less frequently reported anosmia or ageusia (24% vs. 38%) (Table).
We observed no difference in cycle threshold (Ct) values for the envelope gene target between B.1.1.7 and wild-type samples (median 20.2 vs. 20.1) (Table). In patients reporting a symptom duration of >7 days, Ct values appeared to be lower for those with B.1.1.7 (26.2 vs. 30.1 for wild-type), but the difference was not significant (p = 0.7 by Mann–Whitney U test) (Table; Figure 2).
Finally, we explored which combination of variables in our dataset best described B.1.1.7 in a logistic regression applying a backward stepwise selection on the basis of the Akaike information criterion (Appendix). This work identified the best set of associated factors as the absence of anosmia or ageusia (p = 0.01), longer symptom duration (p = 0.02), sore throat (p = 0.05), lower Ct value (p = 0.07), travel in the previous 14 days (p = 0.08), lower age (p = 0.09), and absence of rhinorrhea (p = 0.12). We then used the bootstrap technique to repeat the variable selection in 1,000 replicated datasets and evaluated how often these variables were selected with the backward selection. This analysis resulted in anosmia or ageusia, 89%; symptom duration, 78%; travel, 73%; sore throat, 68%; Ct value, 66%; age, 56%; and rhinorrhea, 51%. Absence of anosmia or ageusia, longer symptom duration, and travel were selected most often, indicating their association with B.1.1.7 infection. We performed all analyses in R version 3.6.3 (https://cran.r-project.org).
The first B.1.1.7 case in Germany was recorded in late December 2020 (5). At our testing site, B.1.1.7 was observed 3 weeks later and replaced wild-type virus as the dominant strain within just 5 weeks. The rapid emergence and dominance of this lineage likely results from its increased transmissibility (E. Volz et al., unpub. data), which is potentially caused by spike protein polymorphisms (including 501Y) conferring enhanced mucosal binding (6); 681H, near a region vital for transmission (T.P. Peacock et al., unpub. data, https://doi.org/10.1101/2020.09.30.318311); and deletion 69–70, linked to immune escape (7). Viral replication in vitro does not differ from earlier strains (J.C. Brown et al., unpub. data, https://doi.org/10.1101/2021.02.24.432576).
Compared to the wild-type virus, B.1.1.7 is reportedly associated with more deaths in nonhospitalized patients but not in inpatients (1,2). In our young outpatient study population, we did not observe major, lineage-dependent differences in leading symptoms. Nevertheless, anosmia and ageusia, among the most specific coronavirus disease symptoms (3), were less common in patients with B.1.1.7., whereas sore throat was more common. A survey in the United Kingdom revealed patients with B.1.1.7 experienced anosmia and ageusia less frequently but more frequently experienced sore throat, cough, fatigue, myalgia, and fever (8). In contrast, no associations between SARS-CoV-2 B.1.1.7 and self-reported symptoms, disease duration, or hospital admissions were seen in another UK study (9). The main factors for B.1.1.7 infection prediction in our study appeared to be lack of anosmia or ageusia and longer symptom duration, in addition to recent travel. The association with recent travel at the time of B.1.1.7 spread is probably no longer relevant because B.1.1.7 is now the most common variant in Berlin and Germany.
With regard to Ct values, one study observed similar figures in patients with B.1.1.7 and wild-type lineages; however, a longer duration of PCR–positivity with B.1.1.7 was suggestive by repeated sampling over time (S.M. Kissler et al., unpub. data, https://doi.org/10.1101/2021.02.16.21251535). Likewise, longer persistence has been observed for B.1.1.7 (10), but lower Ct values (indicating higher viral load) have been observed compared to wild-type samples. Lower Ct values were also seen in other studies on population (11) and inpatient levels (2). In our cross-sectional assessment of recently ill outpatients, we did not observe such differences. Still, increased transmissibility may result from the variant’s prolonged excretion (10; S.M. Kissler et al., unpub. data). Test timing appears crucial for interpretation of Ct values. Outpatients are commonly tested earlier than inpatients. The combination of prolonged viral shedding with different test timing might explain increased viral load in B.1.1.7 samples in inpatients but not in recently ill outpatients. In outpatients with ≥7 days symptom duration, Ct values in B.1.1.7 samples were suggestively reduced. However, comparison groups were small. Our data enabled detection of a maximum effect size on overall Ct values, which corresponds to B.1.1.7 Ct values being 1.5 units below or 1.0 above those for wild-type virus.
The main limitation of our study was its limited subgroup sizes, which reduced the likelihood of detecting differences between rare characteristics. Other limitations are the 1-time assessment, subjective symptom duration, and the variable manifestation of SARS-CoV-2 infection (3). Strengths include standardized procedures conducted by trained medical staff and its prospective nature evaluating patient groups during the same period, reducing the likelihood of biases because of temporal effects.
In summary, SARS-CoV-2 VOC B.1.1.7 is now the dominant lineage in Berlin. In outpatients, no major difference in clinical manifestations has been observed.
Ms. Van Loon is a PhD student at the Research Group Malaria and Infectious Diseases Epidemiology, Institute of Tropical Medicine and International Health, Charité–Universitätsmedizin Berlin, Germany. Her primary research interests include the epidemiology of infectious diseases.
This article was preprinted at https://www.medrxiv.org/content/10.1101/2021.04.15.21255389v1.
Acknowledgment
We thank the Corona Untersuchungsstelle staff at the Charité–Universitätsmedizin Berlin for their dedication and hard work during the coronavirus disease pandemic.
References
- Davies NG, Jarvis CI, Edmunds WJ, Jewell NP, Diaz-Ordaz K, Keogh RH; CMMID COVID-19 Working Group. Increased mortality in community-tested cases of SARS-CoV-2 lineage B.1.1.7. Nature. 2021;593:270–4. DOIPubMedGoogle Scholar
- Frampton D, Rampling T, Cross A, Bailey H, Heaney J, Byott M, et al. Genomic characteristics and clinical effect of the emergent SARS-CoV-2 B.1.1.7 lineage in London, UK: a whole-genome sequencing and hospital-based cohort study. Lancet Infect Dis. 2021;
S1473-3099(21)00170-5 ; [Epub ahead of print].PubMedGoogle Scholar - Maechler F, Gertler M, Hermes J, van Loon W, Schwab F, Piening B, et al. Epidemiological and clinical characteristics of SARS-CoV-2 infections at a testing site in Berlin, Germany, March and April 2020-a cross-sectional study. Clin Microbiol Infect. 2020;26:1685.e7–12. DOIPubMedGoogle Scholar
- Poljak M, Korva M, Knap Gašper N, Fujs Komloš K, Sagadin M, Uršič T, et al. Clinical evaluation of the cobas SARS-CoV-2 test and a diagnostic platform switch during 48 hours in the midst of the COVID-19 pandemic. J Clin Microbiol. 2020;58:e00599–20. DOIPubMedGoogle Scholar
- Robert Koch Institute. Coronavirus disease 2019 (COVID-19) daily situation report of the Robert Koch Institute. 2020 Dec 26 [cited 2021 Apr 3]. https://www.rki.de/DE/Content/InfAZ/N/Neuartiges_Coronavirus/Situationsberichte/Dez_2020/2020-12-26-en.pdf?__blob=publicationFile
- Chan KK, Tan TJC, Narayanan KK, Procko E. An engineered decoy receptor for SARS-CoV-2 broadly binds protein S sequence variants. Sci Adv. 2021;7:eabf1738.
- Kemp SA, Collier DA, Datir R, Ferreira IATM, Gayed S, Jahun A, et al. Neutralising antibodies in Spike mediated SARS-CoV-2 adaptation. Nature. 2021;592:277–82. DOIPubMedGoogle Scholar
- Office for National Statistics. Coronavirus (COVID-19) infection survey: characteristics of people testing positive for COVID-19 in England, 27 January 2021 [cited 2021 Apr 3]. https://www.ons.gov.uk/peoplepopulationandcommunity/healthandsocialcare/conditionsanddiseases/articles/coronaviruscovid19infectionsinthecommunityinengland/characteristicsofpeopletestingpositiveforcovid19inengland27january2021
- Graham MS, Sudre CH, May A, Antonelli M, Murray B, Varsavsky T, et al.; COVID-19 Genomics UK (COG-UK) Consortium. Changes in symptomatology, reinfection, and transmissibility associated with the SARS-CoV-2 variant B.1.1.7: an ecological study. Lancet Public Health. 2021;6:e335–45. DOIPubMedGoogle Scholar
- Calistri P, Amato L, Puglia I, Cito F, Di Giuseppe A, Danzetta ML, et al. Infection sustained by lineage B.1.1.7 of SARS-CoV-2 is characterised by longer persistence and higher viral RNA loads in nasopharyngeal swabs. Int J Infect Dis. 2021;105:753–5. DOIPubMedGoogle Scholar
- Kidd M, Richter A, Best A, Cumley N, Mirza J, Percival B, et al. S-variant SARS-CoV-2 lineage B1.1.7 is associated with significantly higher viral loads in samples tested by ThermoFisher TaqPath RT-qPCR. J Infect Dis. 2021;
jiab082 ; Epub ahead of print. DOIPubMedGoogle Scholar
Figures
Table
Cite This ArticleOriginal Publication Date: May 26, 2021
Table of Contents – Volume 27, Number 7—July 2021
| EID Search Options |
|---|
|
|
|
|
|
|
![Comparison of median Ct values in severe acute respiratory syndrome coronavirus 2 wild-type and B.1.1.7 lineage by symptom duration, Berlin, Germany, January–March 2021. A) Symptom duration <7 days. B) Symptom duration >7 days. The boxplots indicate medians (center) and 25th (top) and 75th (bottom) percentiles (i.e., quartile [Q] 1 and Q3). The upper whiskers reach the largest value with a maximum Q3 +1.5 interquartile range. The lower whiskers reach the smallest value with a minimum Q1 –1.5 interquartile range. The numbers on top of the boxplots indicate the total number of observations included in the comparison. Ct, cycle threshold.](/eid/images/21-0845-F2-tn.jpg)
Please use the form below to submit correspondence to the authors or contact them at the following address:
Welmoed van Loon, Institute of Tropical Medicine and International Health, Charité–Universitätsmedizin Berlin, Campus Virchow-Klinikum, Augustenburger Platz 1, 13353 Berlin, Germany
Top