Volume 28, Number 8—August 2022
Dispatch
Zoonotic Threat of G4 Genotype Eurasian Avian-Like Swine Influenza A(H1N1) Viruses, China, 2020
Cite This Article
Citation for Media
Abstract
We investigated genetic and biologic characteristics of 2 Eurasian avian-like H1N1 swine influenza viruses from pigs in China that belong to the predominant G4 genotype. One swine isolate exhibited strikingly great homology to contemporaneous human Eurasian avian-like H1N1 isolates, preferential binding to the human-type receptor, and vigorous replication in mice without adaptation.
Pigs have long been considered a crucial genetic mixing vessel for influenza A viruses (IAVs) of different hosts (1) because of the dual expression of human (SAα-2,6Gal) and avian (SAα-2,3Gal) viral receptors on their respiratory epithelium. Swine IAVs such as H1N1 and H3N2 subtypes sporadically infect humans and are prone to cause bidirectional interspecies transmission at the swine–human interface (2–5). So far, Eurasian avian-like (EA) H1N1 has dominated prevalence in pig herds in China and caused >10 human infections (6–9). In particular, the dominant genotype 4 (G4) EA H1N1 containing 2009 pandemic influenza A(H1N1) polymerase basic (PB) 1 and 2, polymerase acid (PA), nucleoprotein (NP), and matrix (M) genes, plus the triple-reassortant (TR) nonstructural (NS) gene, is thought to be a candidate virus of potential pandemic (10,11). Indeed, a case of human infection with G4 EA H1N1 was reported in Yunan Province, China, in 2021 (8). It is imperative to conduct surveillance on swine IAVs and evaluate their risk to public health.
During monthly surveillance of swine IAVs in China during October–December 2020, we collected a total of 376 nasal swab samples from apparently healthy pigs in a slaughterhouse accommodating swine from neighboring regions (Jiangsu, Shandong, and Anhui Provinces in eastern China). We detected H1 subtype swine influenza virus in 9 of those by real-time reverse transcription quantitative PCR; 2 were confirmed as hemagglutinin (HA) positive after inoculating into MDCK cells (12). We further evaluated these 2 swine IAV isolates, A/swine/Jiangsu/HD11/2020 (H1N1) [HD11] and A/swine/Anhui/HD21/2020 (H1N1) [HD21], for their genetic and biologic characteristics.
The genome sequences of HD11 and HD21 deposited in the GenBank database (accession no. OL744678–93) shared 95.4%–99.0% nucleotide identities across the coding regions of 8 genes. We performed searches of those sequences on BLAST (https://blast.ncbi.nlm.nih.gov/Blast.cgi) and the GISAID database (http://platform.gisaid.org) to present a more comprehensive scene of the homologous reference influenza viruses. As shown by the closest BLAST hits (Table 1), HD11 and HD21 were not only highly related to swine origin IAVs collected during 2012–2018 but also remarkably similar to contemporaneous human H1N1 isolates from 2020 and 2021.
We constructed a phylogenetic gene tree analysis with H1N1 reference strains to confirm the intimate genetic relationship between these 2 swine IAVs and human viruses (Appendix Figure 1). In each tree, HD11 consistently clustered with 3 human H1N1 viruses, A/Tianjin/00030/2020(H1N1), A/Shandong/00204/2021(H1N1), and A/Sichuan/01208/2021(H1N1). As for HD21, the virus aggregated closely with the HD11-involved subbranch in PB2, HA, NP, NA, and M gene trees but gathered more intimately with another 3 human H1N1 viruses containing A/Hubei-Wujiagang/1324/2020(H1N1), A/Gansu-Xifeng/1143/2021(H1N1) and A/Gansu-Xifeng/1194/2021(H1N1) in PB1, PA, and NS gene trees. Taken together, HD11 and HD21 were both closest to contemporaneous human H1N1 strains, and they uniformly possessed the EA H1N1-like HA and NA genes, pandemice influenza–like RNP (PB2, PB1, PA, and NP) and M genes, and TR-like NS gene that made the G4 type gene constellation. We observed that 2 additional swine reference viruses of A/swine/Shandong/LY142/2017(H1N1) and A/swine/China/Qingdao/2018(H1N1) assembled tightly with the HD11/HD21 cluster, further supporting the possibility of IAV interspecies transmission from swine to human.
The 2 G4 genotype EA H1N1 swine isolates both propagated well in specific-pathogen-free chicken embryos with virus titers per 0.1mL >9 log10 50% egg infectious dose (EID50) (Table 2). However, HD11 replicated much better than HD21 in MDCK cells through the titration of the 50% culture infectious dose (TCID50) value and virus growth at 12-hours intervals across 12–60 hours postinfection (hpi). At >24 hpi, HD11 had generated more than 5 log10 TCID50 and reached a peak of 7 log10 TCID50 at 36 hpi, whereas the titer of HD21 virus remained at the relatively lower level <5 log10 TCID50 until the endpoint.
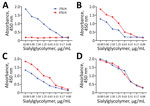
Subsequently, we conducted a solid-phase direct binding ELISA assay with the synthetic glycopolymer-based receptor mimics Neu5Aca2-3Galb1-4GlcNAcb(3ʹSLN)-PAA-biotin and Neu5Aca2-3Galb1-4GlcNAcb(6ʹSLN)-PAA-biotin (GlycoTech, https://www.glycotech.com) to evaluate the viral receptor-binding preference as previously described (13). We used 1 avian H5N1 virus and 1 human seasonal H3N2 virus as controls; the avian virus displayed a complete 3ʹsialyl-N-acetyllactosamine (SLN) affinity, whereas the human virus possessed a dual binding property to both 3ʹSLN and the more advantageous 6ʹSLN (Figure 1). Unlike HD21, which was endowed with comparable avidity between 3ʹSLN and 6ʹSLN, HD11 resembled the binding feature of the human-origin H3N2 virus that preferentially binds the human-type SAα-2,6Gal (Figure 1).
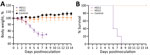
We then investigated the pathogenicity of HD11 and HD21 in mice. We infected 6-week-old BALB/c mice in groups of 5 intranasally with 106.0 EID50 virus dose or mock-inoculated them with phosphate-buffered saline (PBS). We monitored body weight changes and clinical symptoms of the mice daily for 14 days. We humanely euthanized an additional 3 challenged mice per group and analzyed them for virus load in tissues at 3 and 5 days postinfection (dpi). Mice in the control group displayed a steady increase in body weight, the HD21 group experienced a slightly transient weight loss on 3 dpi, and all mice survived during the entire experiment (Figure 2). In contrast, HD11 resulted in a steady decrease in body weights starting at 1 dpi, and all died within 8 days. In addition, we observed that both HD11 and HD21 replicated efficiently in the lungs without prior adaptation and readily disseminated into nasal turbinates and the brain (Table 2). Of note, the virus load in respiratory tissues of HD11-infected mice was significantly higher (p<0.01 in lungs and p<0.001 in turbinates) than that of HD21-infected mice on 3 dpi. On 5 dpi, we observed no significant difference in virus titers in the 3 tissues of the mice infected with these 2 isolates. Moreover, HD11 infections increased the mRNA levels of inflammatory cytokines, including interleukin 6 and 10, interferon β and γ, MX1, and C-X-C motif chemokine ligand 10 11 on 3 dpi, 5 dpi, or both, more dramatically than HD21 virus. Both HD21 and HD11 infections increased tumor necrosis factor α expression at relatively low levels (Appendix Figure 2).
Homology alignment and phylogenetic tree construction analysis suggest that HD11 and HD21, two G4 EA H1N1 swine IAVs isolated in 2020 in China, are strongly related to recent human-origin EA H1N1 viruses. In particular, HD11 had higher affinity for human-type 6ʹSLN at the level that is equivalent to the human seasonal H3N2 virus. Moreover, HD11 replicated much faster in vitro in MDCK cells and in vivo in the lung than di HD21 and was highly pathogenic to BALB/c mice, as evidenced by its lethality, higher viral loads in pulmonary tissues, and higher levels of inflammatory cytokines in the lung. We propose that the HD11-like G4 swine isolates whose genomic sequences share great homology with that of contemporaneous human EA H1N1 viruses may lead to interspecies transmission. Therefore, the public health threat from the zoonotic G4 EA H1N1 viruses should not be underestimated.
Dr. Gu is a faculty member at College of Veterinary Medicine, Yangzhou University. Her research primarily focuses on the epidemiology of zoononic influenza viruses and the mechanisms of virus evolution.
Acknowledgments
We thank Xiulong Xu for revising and editing the manuscript.
This work was supported by the National Key Research and Development Program of China (no. 2021YFD1800202), the National Natural Science Foundation of China (no. 32072892), the Jiangsu Provincial Postdoctoral Science Foundation (no. 1501075C), the Priority Academic Program Development of Jiangsu Higher Education Institutions, the Jiangsu Qinglan Project, and the High-end talent support program of Yangzhou University.
References
- Crisci E, Mussá T, Fraile L, Montoya M. Review: influenza virus in pigs. Mol Immunol. 2013;55:200–11. DOIPubMedGoogle Scholar
- Nelson MI, Vincent AL. Reverse zoonosis of influenza to swine: new perspectives on the human-animal interface. Trends Microbiol. 2015;23:142–53. DOIPubMedGoogle Scholar
- Chastagner A, Enouf V, Peroz D, Hervé S, Lucas P, Quéguiner S, et al. Bidirectional human–swine transmission of seasonal influenza A(H1N1)pdm09 virus in pig herd, France, 2018. Emerg Infect Dis. 2019;25:1940–3. DOIPubMedGoogle Scholar
- Deng YM, Wong FYK, Spirason N, Kaye M, Beazley R, Grau MLL, et al. Locally acquired human infection with swine-origin influenza A(H3N2) variant virus, Australia, 2018. Emerg Infect Dis. 2020;26:143–7. DOIPubMedGoogle Scholar
- Anderson TK, Chang J, Arendsee ZW, Venkatesh D, Souza CK, Kimble JB, et al. Swine influenza A viruses and the tangled relationship with humans. Cold Spring Harb Perspect Med. 2021;11:
a038737 . DOIPubMedGoogle Scholar - Feng Z, Zhu W, Yang L, Liu J, Zhou L, Wang D, et al. Epidemiology and genotypic diversity of Eurasian avian-like H1N1 swine influenza viruses in China. Virol Sin. 2021;36:43–51. DOIPubMedGoogle Scholar
- Li X, Guo L, Liu C, Cheng Y, Kong M, Yang L, et al. Human infection with a novel reassortant Eurasian-avian lineage swine H1N1 virus in northern China. Emerg Microbes Infect. 2019;8:1535–45. DOIPubMedGoogle Scholar
- Li Z, Zhao X, Huang W, Yang L, Cheng Y, Tan M, et al. Etiological characteristics of the first human infection with the G4 genotype Eurasian avian⁃like H1N1 swine influenza virus in Yunnan province, China [in Chinese]. Chin J Virol. 2022;38:290–7.
- World Health Organization. Influenza at the human-animal interface summary and assessment, 1 October 2021. 2021 [cited 2021 Dec 14]. https://www.who.int/publications/m/item/influenza-at-the-human-animal-interface-summary-and-assessment-1-october-2021
- Yang H, Chen Y, Qiao C, He X, Zhou H, Sun Y, et al. Prevalence, genetics, and transmissibility in ferrets of Eurasian avian-like H1N1 swine influenza viruses. Proc Natl Acad Sci U S A. 2016;113:392–7. DOIPubMedGoogle Scholar
- Sun H, Xiao Y, Liu J, Wang D, Li F, Wang C, et al. Prevalent Eurasian avian-like H1N1 swine influenza virus with 2009 pandemic viral genes facilitating human infection. Proc Natl Acad Sci U S A. 2020;117:17204–10. DOIPubMedGoogle Scholar
- Chen K, Kong M, Liu J, Jiao J, Zeng Z, Shi L, et al. Rapid differential detection of subtype H1 and H3 swine influenza viruses using a TaqMan-MGB-based duplex one-step real-time RT-PCR assay. Arch Virol. 2021;166:2217–24. DOIPubMedGoogle Scholar
- Yamada S, Suzuki Y, Suzuki T, Le MQ, Nidom CA, Sakai-Tagawa Y, et al. Haemagglutinin mutations responsible for the binding of H5N1 influenza A viruses to human-type receptors. Nature. 2006;444:378–82. DOIPubMedGoogle Scholar
Figures
Tables
Cite This ArticleOriginal Publication Date: July 11, 2022
1These authors contributed equally to this article.
Table of Contents – Volume 28, Number 8—August 2022
| EID Search Options |
|---|
|
|
|
|
|
|

Please use the form below to submit correspondence to the authors or contact them at the following address:
Xiufan Liu, Animal Infectious Diseases Laboratory, College of Veterinary Medicine, Yangzhou University, 48 East Wenhui Road, Yangzhou, Jiangsu 225009, China
Top