Volume 29, Number 2—February 2023
Dispatch
Molecular Detection of Candidatus Orientia chuto in Wildlife, Saudi Arabia
Cite This Article
Citation for Media
Abstract
Scrub typhus is a zoonosis caused by 3 species of Orientia bacteria, including Candidatus Orientia chuto. This species is known only from a human case in Dubai and infections in wildlife in Kenya. We report molecular detection of Candidatus O. chuto in 2 wild rodent species from Saudi Arabia.
Scrub typhus is a zoonotic bacterial disease caused by 3 intracellular species of bacteria in the genus Orientia (Rickettsiales: Rickettsiaceae). The disease is widespread in the Asia-Pacific Region and is associated with fever, as well as pneumonitis, encephalitis, and myocarditis if not promptly treated. The median case fatality rate is ≈6% (1), and one third of infections during pregnancy result in adverse outcomes (2).
Orientia spp. are transmitted to humans through the bite of infected trombiculid mite larva (chiggers), which feed primarily on small mammals or birds and only incidentally attack humans. Orientia spp. are maintained by vertical transmission in trombiculid mites, but wild vertebrate hosts can become infected. However, whether host species are epidemiologically meaningful Orientia reservoirs is controversial because horizontal transmission of Orientia spp. to chiggers during feeding rarely translates into successful transstadial transfer and transovarial transmission into the next generation (3). At minimum, wild hosts contribute to scrub typhus risk by amplifying trombiculid populations; individual hosts can potentially be infested with thousands of chiggers simultaneously (4).
Until the early 21st Century, only 1 species of Orientia was known: Orientia tsutsugamushi, which is restricted to the tsutsugamushi triangle across the Asia-Pacific region. The precise western limit of O. tsutsugamushi endemicity is unclear, but reports beyond the Hindu Kush region, where Afghanistan, Tajikistan, and Pakistan converge, are very rare. However, in 2006, a woman contracted scrub typhus while visiting Dubai (5). The pathogen was isolated in culture, and molecular characterization established that it was sufficiently distinct from O. tsutsugamushi to be classified as a new species, Candidatus Orientia chuto. Since that report, Candidatus O. chuto–like sequences rarely have been detected, but 1 report describes detection from chiggers infesting a Natal multimammate mouse (Mastomys natalensis) in Baringo County, Kenya (6). A third species of scrub typhus pathogen, Candidatus O. chiloensis, was recently described from patients in Chile (7). We investigated whether Orientia spp. are circulating in small mammals and chiggers in Saudi Arabia.
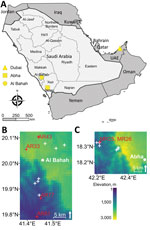
We trapped rodents in southwestern Saudi Arabia, as previously described (8) (Figure 1). We humanely euthanized rodents by inhaled anesthetic overdose and preserved any attached chiggers in 70% ethanol. The fieldwork was approved by the Saudi Wildlife Authority (approval no. 288/33/A) and the University of Liverpool’s Animal Welfare and Ethics Review Board.
We dissected rodents and fixed the internal organs (lungs, spleen, liver, and kidney) in 70% ethanol. To identify chiggers, we cleared a 10% subsample from each rodent in Berlese fluid and mounted these for microscopic examination (8). We pooled the remaining chiggers by species (10–30 specimens per pool) from each rodent. We crushed chigger pools with a pellet-pestle, then extracted DNA by using the DNeasy Blood & Tissue Kit (QIAGEN, https://www.qiagen.com) according to the manufacturer’s instructions and eluted DNA in 30 μL ultrapure water.
We purified genomic DNA from individual mammal host organs (10 mg spleen or 25 mg for other organs) by using the DNeasy Blood & Tissue Kit (QIAGEN) and eluting DNA in 50 μL ultrapure water. We identified rodents by amplifying a cytochrome B gene fragment and performing BLAST analysis (https://blast.ncbi.nlm.nih.gov).
To detect Orientia DNA, we screened rodent and chigger extracts by using a quantitative PCR targeting the multicopy traD gene (9). We subjected positive samples to nested PCRs designed to amplify the 47 kDa high-temperature requirement A (htrA) gene, also known as TSA47, from O. tsutsugamushi or Candidatus O. chuto (6). We sent amplicons from the second round of the nested PCR to Eurofins Genomics (https://www.eurofins.com) for Sanger sequencing in both directions, then trimmed results to 698 bp, and aligned with reference sequences from GenBank. We used MrBayes (10) to construct a phylogenetic tree by using MUSCLE alignment in the Phylogeny.fr web service (11). We estimated the best-fit model of nucleotide substitution by using Akaike information criterion in jModelTest 2.1.7 (12) and selected the general time-reversible plus gamma distribution plus invariable site model as the best fit. We calculated pairwise distances between sequences by using MEGA 11 (https://www.megasoftware.net).
We trapped 27 rodents in Asir Province, all of which we identified as Eastern spiny mice (Acomys dimidiatus). We trapped 55 rodents of 3 different species in Al-Bahah Province (Table 1). Using htrA primers for Candidatus O. chuto, we identified 7 sequence-confirmed positive organs from 6 individual animals, 2 (7.4%) from Asir Province and 4 (7.3%) from Al-Bahah Province (Table 2). We found infected A. dimidiatus in both provinces, and a single infected Wagner’s gerbil (Dipodillus dasyurus) in Al-Bahah. Positive rodents were widely distributed both in terms of habitat type and elevation (388–2,477 meters above sea level) at the trap location (Table 2; Figure 1). One A. dimidiatus from Asir had 2 organs that tested positive for Candidatus O. chuto, but all other rodents had a single PCR-positive organ (Table 2). Only 2 rodents were infested with chiggers, and we identified a total of 5 chigger species. None of the chigger samples were PCR-positive for Orientia spp. DNA.
Phylogenetic analysis of publicly available htrA gene sequences revealed 3 well-supported clades representing O. tsutsugamushi, Candidatus O. chiloensis, and Candidatus O. chuto (Figure 2). The Candidatus O. chuto lineage was split into 2 main clades comprising isolates from the Arabian Peninsula, including isolates from this study and the clinical isolate from Dubai in 1 clade and the isolate from Kenya in the other clade. The Candidatus O. chuto sequences from this study were distinct from the Dubai isolate (pairwise distance of 0.7%–1.3%) and comprised 2 genotypes exhibiting a pairwise distance of 0.9% (Appendix Table), which was linked to geographic origin from Al-Bahah or Asir Province (Figure 2).
Only 2 published records of Candidatus O. chuto have been available worldwide, 1 from Dubai (5) and 1 from Baringo, Kenya (6), and only the Dubai isolate has been cultured in vitro and fully sequenced. We identified 2 Orientia genotypes from Saudi Arabia, neither of which was identical to previously described isolates. Separation between Orientia spp. at the htrA locus is ≈11%–14%, whereas within O. tsutsugamushi, htrA attains a maximum separation of 1.5%. The maximum separation between the sequences from the Arabian Peninsula and Kenya was 2.5% (Appendix Table), indicating that whole-genome sequencing studies are needed to resolve relationships between Candidatus O. chuto–like organisms.
The potential range of Candidatus O. chuto is vast, and the bacterium is suspected of causing scrub typhus on the continent of Africa, which is indicated by serologic and clinical evidence, albeit without confirmation by sequencing or culture to date. The Orientia-infected rodents detected in this study were found in locations >2,000 km northeast of Baringo County, Kenya, and >1,500 km southwest of Dubai. Thus, the endemic zone of Candidatus O. chuto clearly includes northeastern Africa and the Arabian Peninsula, at least, but also could potentially extend eastwards to the western extremity of the tsutsugamushi triangle. Of note, Orientia spp. detected by 16S amplicon sequencing in house mice (Mus musculus domesticus) in West Africa resembled Candidatus O. chuto, although sequences obtained from rodents in France formed a clade that was distinct from known Orientia spp. (13).
Scrub typhus has not been reported from Saudi Arabia, but our findings highlight the need for vigilance. Moreover, further investigations of the diverse chigger fauna of Saudi Arabia are required to identify the local vectors of Orientia spp. Because other agents of febrile illness are endemic in the southwest of the country, including Plasmodium vivax (14) and dengue virus (15), and the region is popular with tourists because of its relatively cool climate, we recommend scrub typhus be included in the differential diagnosis for fever in Saudi Arabia.
Mrs. Alkathiry is a biologist and PhD student of parasitology in the Institute of Infection, Veterinary & Ecological Sciences at the University of Liverpool, Liverpool, England, UK. Her primary research interests include rodents, vector-borne diseases, and microbiomes.
H.A.A., S.Q.A. and B.L.M. designed the study; H.A.A., S.Q.A. and A.N.A. selected field sites and trapped rodents; H.A.A. identified rodents and mites; H.A.A. and H.E.J.M. performed DNA extractions, PCR assays and sequence analysis; M.E.N. produced Figure 1; J.J.K. conducted the phylogenetics; B.L.M. supervised the study and wrote the first manuscript draft. All authors read and approved the manuscript. The authors declare no conflict of interest.
Acknowledgments
We are grateful to Al-Bahah University for access to research facilities during fieldwork.
This work was funded by a doctoral scholarship from Imam Muhammad Ibn Saud Islamic University to H.A.A. The Vice Deanship of Research Chairs, Deanship of Scientific Research of the King Saud University, Kingdom of Saudi Arabia, provided support to A.N.A.
References
- Bonell A, Lubell Y, Newton PN, Crump JA, Paris DH. Estimating the burden of scrub typhus: A systematic review. PLoS Negl Trop Dis. 2017;11:
e0005838 . DOIPubMedGoogle Scholar - McGready R, Prakash JAJ, Benjamin SJ, Watthanaworawit W, Anantatat T, Tanganuchitcharnchai A, et al. Pregnancy outcome in relation to treatment of murine typhus and scrub typhus infection: a fever cohort and a case series analysis. PLoS Negl Trop Dis. 2014;8:
e3327 . DOIPubMedGoogle Scholar - Traub R, Wisseman CL Jr, Jones MR, O’Keefe JJ. The acquisition of Rickettsia tsutsugamushi by chiggers (trombiculid mites) during the feeding process. Ann N Y Acad Sci. 1975;266(1 Pathobiology):91–114.
- Elliott I, Pearson I, Dahal P, Thomas NV, Roberts T, Newton PN. Scrub typhus ecology: a systematic review of Orientia in vectors and hosts. Parasit Vectors. 2019;12:513. DOIPubMedGoogle Scholar
- Izzard L, Fuller A, Blacksell SD, Paris DH, Richards AL, Aukkanit N, et al. Isolation of a novel Orientia species (O. chuto sp. nov.) from a patient infected in Dubai. J Clin Microbiol. 2010;48:4404–9. DOIPubMedGoogle Scholar
- Masakhwe C, Linsuwanon P, Kimita G, Mutai B, Leepitakrat S, Yalwala S, et al. Identification and characterization of Orientia chuto in trombiculid chigger mites collected from wild rodents in Kenya. J Clin Microbiol. 2018;56:e01124–18. DOIPubMedGoogle Scholar
- Abarca K, Martínez-Valdebenito C, Angulo J, Jiang J, Farris CM, Richards AL, et al. Molecular description of a novel Orientia species causing scrub typhus in Chile. Emerg Infect Dis. 2020;26:2148–56. DOIPubMedGoogle Scholar
- Stekolnikov AA, Al-Ghamdi SQ, Alagaili AN, Makepeace BL. First data on chigger mites (Acariformes: Trombiculidae) of Saudi Arabia, with a description of four new species. Syst Appl Acarol. 2019;24:1937–63. DOIGoogle Scholar
- Chao C-C, Belinskaya T, Zhang Z, Jiang L, Ching W-M. Assessment of a sensitive qPCR assay targeting a multiple-copy gene to detect Orientia tsutsugamushi DNA. Trop Med Infect Dis. 2019;4:113. DOIPubMedGoogle Scholar
- Ronquist F, Teslenko M, van der Mark P, Ayres DL, Darling A, Höhna S, et al. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol. 2012;61:539–42. DOIPubMedGoogle Scholar
- Dereeper A, Guignon V, Blanc G, Audic S, Buffet S, Chevenet F, et al. Phylogeny.fr: robust phylogenetic analysis for the non-specialist. Nucleic Acids Res. 2008;36(Suppl 2):
W465-9 . DOIPubMedGoogle Scholar - Posada D. jModelTest: phylogenetic model averaging. Mol Biol Evol. 2008;25:1253–6. DOIPubMedGoogle Scholar
- Cosson JF, Galan M, Bard E, Razzauti M, Bernard M, Morand S, et al. Detection of Orientia sp. DNA in rodents from Asia, West Africa and Europe. Parasit Vectors. 2015;8:172. DOIPubMedGoogle Scholar
- Al-Mekhlafi HM, Madkhali AM, Ghailan KY, Abdulhaq AA, Ghzwani AH, Zain KA, et al. Residual malaria in Jazan region, southwestern Saudi Arabia: the situation, challenges and climatic drivers of autochthonous malaria. Malar J. 2021;20:315. DOIPubMedGoogle Scholar
- Alhaeli A, Bahkali S, Ali A, Househ MS, El-Metwally AA. The epidemiology of Dengue fever in Saudi Arabia: A systematic review. J Infect Public Health. 2016;9:117–24. DOIPubMedGoogle Scholar
Figures
Tables
Cite This ArticleOriginal Publication Date: January 16, 2023
Table of Contents – Volume 29, Number 2—February 2023
| EID Search Options |
|---|
|
|
|
|
|
|

Please use the form below to submit correspondence to the authors or contact them at the following address:
Benjamin Makepeace, Institute of Infection, Veterinary & Ecological Sciences, University of Liverpool, Liverpool Science Park IC2, Liverpool L3 5RF, UK
Top