Volume 30, Supplement—October 2024
SUPPLEMENT ISSUE
Articles
Genomic Epidemiology of Multidrug-Resistant Escherichia coli and Klebsiella pneumoniae in Kenya, Uganda, and Jordan
Figure 1
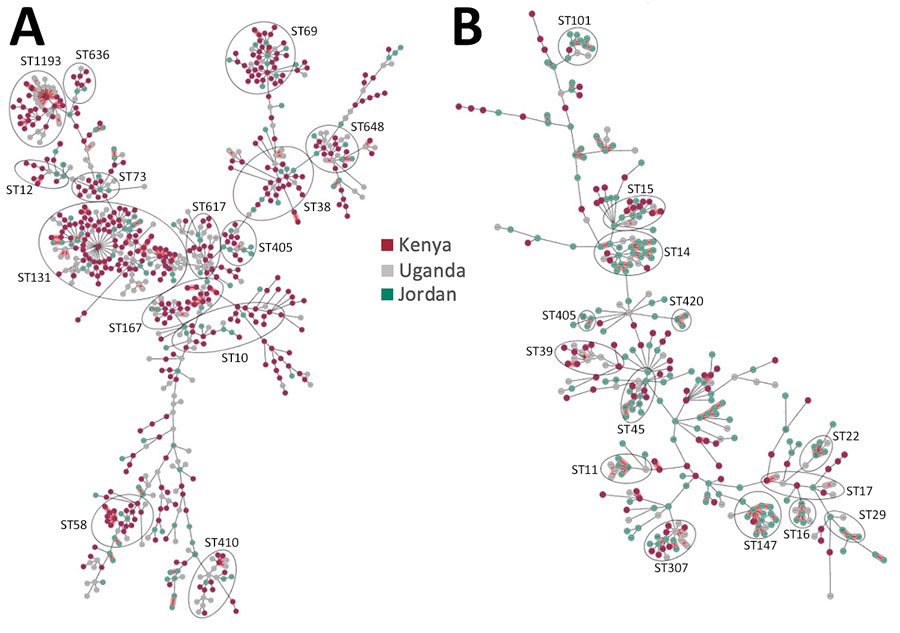
Figure 1. Population structure and diversity of high-risk Escherichia coli and Klebsiella pneumoniae sequence types across Kenya, Uganda, and Jordan. Minimum-spanning trees of E. coli (n = 785) and K. pneumoniae (n = 378) isolates are based on core-genome multilocus sequence typing. Each node represents an isolate; dominant STs are indicated in circled clusters. Branch length between nodes is proportional to the allelic differences between nodes. Purple indicates isolates from Kenya, gray from Uganda, and green from Jordan. ST, sequence type.
Page created: October 30, 2024
Page updated: November 11, 2024
Page reviewed: November 11, 2024
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.