Stenotrophomonas maltophilia Bloodstream Infection Outbreak in Acute Care Hospital, California, USA, 2022–20231
Sana M. Khan, Axel A. Vazquez Deida, Steven Langerman, Jennifer C. Hunter, Rebeca Elliott, Alison Laufer Halpin, Alyssa G. Kent, Paige Gable, Heather A. Moulton-Meissner, Frances C. Knight, Thomas Ewing, Kristen Clancy, Amit Chitnis, Eileen F. Dunne, Dustin Heaton, Barbara Allen, Hillary Metcalf, Munira Shemsu, Kathleen Nava, Suada Abdic, Kiran M. Perkins, Elsa Villarino, Jeffrey Silvers, and Kavita K. Trivedi

Author affiliation: Centers for Disease Control and Prevention, Atlanta, Georgia, USA (S.M. Khan, A.A. Vazquez Deida, S. Langerman, J.C. Hunter, A. Laufer Halpin, A.G. Kent, P. Gable, H.A. Moulton-Meissner, F.C. Knight, T. Ewing, K. Clancy, K.M. Perkins); Alameda County Public Health Department, San Leandro, California, USA (S.M. Khan, A. Chitnis, E.F. Dunne, D. Heaton, M. Shemsu, K.K. Trivedi); California Department of Public Health, Richmond, California, USA (R. Elliott, B. Allen, H. Metcalf, E. Villarino); Sutter Health, Sacramento, California, USA (K. Nava, S. Abdic, J. Silvers)
Main Article
Figure 2
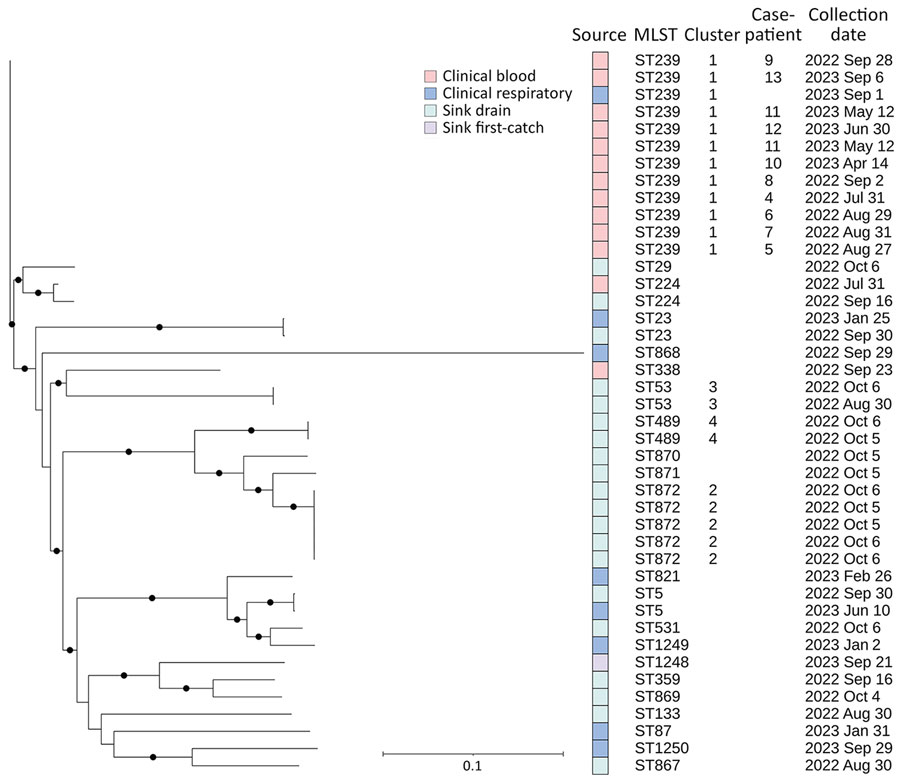
Figure 2. Phylogeny of Stenotrophomonas maltophilia detected in bloodstream infection outbreak in acute care hospital, California, United States, 2022–2023. Case-patient numbering starts at 4 because isolates for the first 3 cases were unavailable for sequencing. Full circles on the phylogenetic tree indicate evolutionary branching in the tree with a high value (n = 1) or strong support for the hypothesis that the branching is true. Cluster 1 (n = 12) differed by 0–4 pairwise high-quality single-nucleotide variants (hqSNVs), across 98.54% of the reference isolate core-genome. Cluster 2 (n = 5) differed by 1–21 hqSNVs, across 97.56% of the core-genome. Cluster 3 (n = 2) differed by 12 hqSNVs, across 98.68% of the core-genome. Cluster 4 (n = 2) differed by 3 hqSNVs, across 99.01% of the core-genome. Scale bar indicates nucleotide substitutions per site.
Main Article
Page created: February 06, 2026
Page updated: March 20, 2026
Page reviewed: March 20, 2026
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.
