Volume 32, Number 4—April 2026
Research
Border Region Surveillance of Malaria Drug Resistance, Northern Burundi, 2023–2024
Cite This Article
Citation for Media
Abstract
To evaluated artemisinin partial resistance (ART-R) in malaria in Burundi, during December 2023–June 2024, we studied 423 children <5 years of age with uncomplicated Plasmodium falciparum malaria in 8 health facilities in the northern part of the country. After artemether/lumefantrine treatment with only the first dose directly observed, 4.5% remained parasitemic on day 3. No pfkelch13 mutations, validated or candidate markers of ART-R, were detected. However, markers of antifolate and 4-aminoquinoline resistance were widespread: the dhfr triple mutant N51I/C59R/S108N was nearly fixed (92%), dhps double and triple mutants were common (41% and 47%), and pfcrt CVIET, associated with chloroquine and amodiaquine resistance, predominated (84%). Geographic differences occurred in day-3 positivity and haplotype frequencies. Although ART-R markers were absent, delayed parasite clearance and near fixation of multidrug-resistant haplotypes serve as a warning. Strengthened efficacy monitoring and regional molecular surveillance are urgently needed to prevent drug-resistant P. falciparum from becoming established in Burundi.
Malaria remains a leading cause of illness and death in low- and middle-income countries, and sub-Saharan Africa bears the greatest burden. In 2024, an estimated 280 million malaria cases and 610,000 deaths occurred globally, of which 94% of cases and 95% of deaths were in sub-Saharan Africa. Children <5 years of age and pregnant women are the most affected groups (1,2).
To accelerate control, the World Health Organization (WHO) launched the Global Technical Strategy for Malaria 2016–2030, targeting a 90% reduction in incidence and mortality and elimination in 35 countries by 2030 (3). Burundi has scaled up core interventions, including artemisinin-based combination therapies (ACTs) since 2003, long-lasting insecticidal nets, indoor residual spraying, and intermittent preventive treatment in pregnancy (IPTp) (4–7). First-line treatment shifted from artesunate/amodiaquine to artemether/lumefantrine in 2019 (4). More recently, perennial malaria chemoprevention (PMC) was introduced in 5 districts, and RTS,S/AS01 vaccine (GSK, https://www.glaxosmithkline.com) was integrated into routine immunization in March 2025, with initial deployment in 25 of the country’s 47 health districts (5,6). Despite those efforts, malaria transmission remains intense; 4.9 million cases and 809 deaths were reported by the national surveillance system in 2023–2024, including 2.2 million cases among children <5 years of age (5,6).
The emergence and spread of drug resistance is a looming threat. Artemisinin partial resistance (ART-R), defined by day 3 parasitemia and validated pfkelch13 mutations, has been confirmed in Rwanda and Tanzania (8–14). Given porous borders and high population mobility, Burundi faces increased risk for resistant Plasmodium falciparum introduction. We conducted a prospective study in northern Burundi using a simplified day 3 follow-up design after artemether/lumefantrine treatment to assess delayed parasite clearance and the key molecular resistance markers, pfkelch13, pfcrt, pfmdr1, dhfr, and dhps.
Study Design and Population
Study Design and Setting
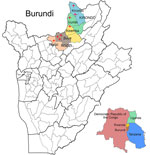
We conducted the study during December 2023–June 2024 in 8 health facilities across 5 districts of northern Burundi, a high P. falciparum transmission zone bordering Rwanda (Figure). Sites were located in Kirundo Province (2 facilities in Kirundo, 1 in Vumbi) and Ngozi Province (3 facilities in Buye, 1 in Ngozi, and 1 in Kiremba). This region experiences intense perennial transmission with seasonal peaks during March–June and October–December. According to national surveillance data, parasitemia prevalence in children <5 years of age reaches 79% in Kirundo and 54% in Ngozi (15), and clinical incidence approaches 400 cases/1,000 population annually (5,6,15).
Study Population and Eligibility Criteria
We screened children <5 years of age who had fever (≥37.5°C axillary or history of fever within the previous 24 h). Inclusion required microscopically confirmed P. falciparum monoinfection. We applied no minimum parasite density or hemoglobin thresholds for enrollment. Exclusion criteria were signs of severe malaria, mixed infections, or known hypersensitivity to artemisinin derivatives.
Treatment, Follow-up Procedures, and Outcomes
Participants received a standard 3-day course of artemether/lumefantrine (Lumartem DT, https://www.cipla.com; WHO-prequalified dispersible tablets, 20 mg artemether/120 mg lumefantrine) according to Burundi national guidelines (4). Dosing followed WHO weight-based recommendations: children 5 to <15 kg received 1 dispersible tablet per dose, and children 15 to <25 kg received 2 tablets per dose. Each treatment course consisted of 6 doses administered at 0, 8, 24, 36, 48, and 60 hours. The first dose was directly observed and was administered with fatty food to enhance lumefantrine absorption. Caregivers were instructed to administer subsequent doses with food at home with reinforced instructions. On day 0 and day 3, we collected capillary blood samples for Giemsa-stained smears and dried blood spots for molecular analysis. On day 3, all children underwent clinical assessment including temperature measurement and symptom evaluation. We referred children with persistent fever or clinical deterioration for management, according to national guidelines. We did not re-treat those with persistent parasitemia but no danger signs because the study assessed day 3 positivity for surveillance purposes rather than to guide individual treatment.
Microscopy
We prepared thick and thin smears and stained with 10% Giemsa (pH 7.2) for 15 minutes. We used thick smears for parasite detection and density quantification and thin smears for species identification on the basis of parasite morphology when mixed infections were suspected or when species confirmation was required. Two certified microscopists independently read slides at 1,000× magnification; discrepancies >25% in parasite density were resolved by a third reader, and we calculated the final parasite density as the mean of the 2 closest readings. We calculated parasite density per 200 leukocytes (8,000/µL) and recorded gametocyte presence.
Molecular Analysis
We collected dried blood spots on Whatman 903 filter paper, air-dried, stored with desiccant, and shipped to the University of Strasbourg (Strasbourg, France) for analysis. We extracted DNA from dried blood spots by using the QIAamp DNA Blood Mini Kit (QIAGEN, https://www.qiagen.com). We confirmed Plasmodium species by real-time PCR targeting cytb and by nested species-specific PCR (16).
We analyzed drug-resistance markers by targeted amplicon sequencing of pfkelch13 (codons 430–720), pfcrt (codons 72–76, 93, 97, 145, 218, 343, 350, 353, and 356), pfmdr1 (codons 86, 184, 1034, 1042, and 1246), dhfr (codons 16, 50, 51, 59, 108, and 164), and dhps (codons 431, 436, 437, 540, 581, and 613) (17). We multiplexed and barcoded amplicons, then pooled, purified, and sequenced on an Illumina MiSeq platform (300-cycle v2 kit; Illumina, https://www.illumina.com). We quality-trimmed reads (Phred score >30; https://www.phrap.com/phred) and aligned reads to the P. falciparum 3D7 reference genome (v45) by using CLC Genomics Workbench version 22 (QIAGEN). We called variants that were supported by >20 reads and allele frequency >5%. Reference strains Dd2, 7G8, HB3, and 3D7 served as positive controls (17).
Sample Size Considerations
This surveillance was designed to detect circulating pfkelch13 mutant parasites, particularly the R561H allele, validated in neighboring Rwanda and Tanzania. We targeted ≈50 children per site (400 total) to ensure broad geographic coverage of the border region while maintaining logistical feasibility. This sample size provides 95% confidence to detect mutant alleles circulating at >0.75% prevalence if none are observed (rule of three: 3/n). The study was not powered for formal interdistrict comparisons; district-level differences in baseline characteristics and molecular markers are reported as exploratory observations to generate hypotheses for future targeted investigations.
Study Outcomes
The primary outcome was the percentage of children with persistent parasitemia on day 3 (day 3 positivity), defined as microscopically detectable asexual P. falciparum parasites on day 3 after treatment initiation. That indicator serves as a clinical proxy for delayed parasite clearance associated with ART-R. Secondary outcomes included the prevalence of molecular markers associated with malaria drug resistance: pfkelch13 mutations (ART-R), pfcrt and pfmdr1 haplotypes (chloroquine, amodiaquine, and piperaquine resistance), and dhfr/dhps mutations (sulfadoxine/pyrimethamine resistance).
Statistical Analysis
We summarized demographic and clinical variables by using descriptive statistics and compared continuous variables by using the Kruskal-Wallis test and categorical variables by using χ2 or Fisher exact tests. We considered p<0.05 significant.
We assessed day 3 positivity after treatment with only the first dose directly observed, estimated prevalence with exact binomial 95% CIs and compared across provinces (Kirundo and Ngozi) to assess geographic heterogeneity. We calculated allele frequencies for pfkelch13, pfcrt, pfmdr1, dhfr, and dhps and stratified by district. We classified mixed infections as mutant for allele-frequency estimates. We reconstructed inferred dhfr-dhps haplotypes from unphased single-nucleotide polymorphism data and compared them between districts; we interpreted results as inferred patterns rather than observed genotypes. We performed analyses with MedCalc version 20.218 (MedCalc Software Ltd, https://www.medcalc.org).
Ethical Considerations
The study followed the Declaration of Helsinki and WHO guidelines for human research. Approval was obtained from the Burundi National Ethics Committee for the Protection of Human Subjects (approval no. CNE/25/2023). Written informed consent was obtained from parents or legal guardians. Participant confidentiality and data security were strictly maintained.
We screened 873 febrile children <5 years of age across the 8 health facilities. Of those, 423 met the inclusion criteria and completed clinical follow-up through day 3, corresponding to 1 day after completion of artemether/lumefantrine treatment. Of the 423 samples collected, 365 (86.3%) day 0 samples were successfully confirmed as P. falciparum infections and subsequently analyzed by molecular sequencing to address the secondary objective: detection of pfkelch13 mutations, notably R561H, a validated marker of ART-R. Sequencing success varied by district: 100% in Kiremba and Ngozi, 91% in Kirundo, 75% in Buye, and 76% in Vumbi. We excluded the other 58 samples (13.7%) because of insufficient DNA quality.
The median age of enrolled children was 30 (IQR 17–45) months, and the sex ratio was balanced (1.0). Baseline characteristics showed variation across districts (Table). Median axillary temperatures ranged from 37.8°C (Buye) to 39.0°C (Kirundo). Pretreatment asexual P. falciparum parasite density varied widely; median values ranged from 10,543/µL (Vumbi) to 42,400/µL (Kirundo). Gametocyte carriage was detected in 5.4% of children overall, with district-specific prevalence ranging from 1.2% (Kirundo) to 22.0% (Ngozi). Mixed-species infections were uncommon (1.2% P. malariae in Ngozi Province), confirming P. falciparum as the predominant species.
Among the 423 children enrolled, 19 (4.5%; 95% CI 2.7%–6.9%) remained parasitemic on day 3 after artemether/lumefantrine treatment. That overall prevalence was below the WHO 10% threshold, but we observed marked heterogeneity across districts. Vumbi (13.6%) exceeded the WHO 10% threshold and neighboring districts, identifying a localized hotspot of delayed clearance. Those differences were statistically significant within Kirundo Province districts (p = 0.0002), but we observed no significant variation between provinces (p = 0.06).
The median asexual parasite density on day 3 was 260 parasites/µL (IQR 141–1,659 parasites/µL), with no significant variation across districts (p = 0.96). Nevertheless, district-specific values showed marked variation, from very low medians in Buye and Kiremba (≈100 parasites/µL) to >1,800 parasites/µL in Ngozi district. Gametocytes were detected microscopically in 11 (2.6%) children on day 3. Their prevalence varied geographically: uncommon in Kirundo Province (1.2%) but more frequent in Ngozi Province (4.8%) and peaked at 10.0% in Kiremba district (Table).
For day 0 isolates, of the 423 children enrolled, 365 (86.3%) samples yielded high-quality sequences and were included in the molecular analysis (Appendix Tables 1, 2). Most (97.5%, 356/365), isolates were wild-type at pfkelch13. We detected no validated or candidate WHO-associated mutations linked to ART-R. We did observe nonsynonymous variants in 9 (2.5%) isolates. The most frequent variant was A578S (4 cases, 1.1%), whereas R513S, A626S, H644R, and N672H were each detected in 1 or 2 isolates. None of those polymorphisms are currently associated with delayed parasite clearance, but their detection might indicate ongoing low-level diversification of the pfkelch13 locus in this setting.
Mutant pfcrt haplotypes were nearly fixed. The CVIET triple mutant, defined by mutations at pfcrt codons 72–76 (C72/V73/I74/E75/T76), predominated (84.4%, 308/365), whereas the CVIET+356T haplotype was detected in 9.3% (34/365) of isolates. The CVMNK wild-type was uncommon (4.9%, 18/365). Minor variants included SVMNT (72S/76T, 1.1%) and 356T alone (0.3%). We observed no significant difference in overall pfcrt haplotype distribution between provinces (p = 0.25). However, intraprovincial heterogeneity was evident, particularly for codon 356 in Ngozi Province (p = 0.01), suggesting microgeographic clustering of resistant variants. The predominance of CVIET confirms that chloroquine resistance remains highly fixed in northern Burundi, with additional mutations such as 356T emerging at low frequency. Of note, we detected no pfcrt mutations associated with piperaquine resistance. All isolates were wild-type at positions H97, F145, I218, M343, C350, and G353. Although the 356T variant occurred in 9.6% (35/365) of isolates, that mutation alone does not confer piperaquine resistance.
Mutant haplotypes were common at the pfmdr1 locus. We identified the wild-type NYSND haplotype, defined by pfmdr1 codons 86-184-1034-1042-1246 (N86/Y184/S1034/N1042/D1246), in 31.2% (114/365) of isolates. The most frequent mutant was the single 184F allele, detected in 57.3% (209/365) of isolates, followed by 86Y (7.9%). Additional combinations involving 86Y, 1034C, 1042D, or 1246Y remained uncommon (<1.6% each). We noted significant differences in haplotype distribution between provinces (p = 0.03), suggesting that selective pressures on pfmdr1 might vary geographically, potentially reflecting differences in drug use or drug pressure intensity.
At the dhfr locus, resistance alleles were nearly fixed. The triple mutant 51I/59R/108N predominated (92.1%, 336/365), confirming widespread pyrimethamine resistance. Other haplotypes were uncommon, including the double mutant 51I/108N (1.1%) and the single 108N variant (0.5%), and we detected the quadruple mutant 51I/59R/108N/164L in 12 (3.3%) isolates. Although infrequent, the quadruple mutant genotype is clinically relevant because the addition of 164L confers high-grade pyrimethamine resistance and has been associated with sulfadoxine/pyrimethamine treatment failure in East Africa. We detected the wild-type haplotype in only 12 (3.3%) isolates. Variation between provinces was not statistically significant (p = 0.2).
Mutations at the dhps locus were also widespread. Only 21/365 (5.8%) isolates were wild-type. The double mutant 437G/540E was detected in 148/365 (40.5%) isolates, whereas the triple mutant 437G/540E/581G was even more frequent (172/365 isolates, 47.1%). Minor haplotypes included 436A (3.8%) and 613S (0.3%). Although we observed no significant difference in overall haplotype proportions between provinces (p = 0.6), district-level variation within Kirundo Province was significant for codons 436A (p = 0.0001), 437G (p = 0.001), and 540E (p = 0.02).
Analysis of combined genotypes enabled inference of multilocus dhfr/dhps haplotypes, highlighting the extensive accumulation of antifolate resistance alleles. The fully wild-type dhfr/dhps combination was nearly absent (1/365 isolates, 0.3%). The most prevalent inferred haplotype was the sextuple mutant (dhfr triple 51I/59R/108N + dhps triple 437G/540E/581G), detected in 161/365 (44.1%) isolates. That genotype has been associated with markedly reduced efficacy of IPTp in East Africa. The classic quintuple mutant (dhfr triple 51I/59R/108N + dhps double 437G/540E) was also common, found in 134/365 (36.7%) isolates. Together, those 2 high-resistance genotypes accounted for >80% of infections. Less frequent combinations included quadruple or septuple haplotypes, each detected in <3% of isolates. Although overall haplotype distribution did not differ significantly between provinces (p = 0.2), variation occurred at the district level.
We performed molecular analysis on isolates from the 19 children who remained parasitemic on day 3. All isolates were wild-type at pfkelch13, confirming the absence of validated or candidate ART-R mutations. At pfcrt, the CVIET haplotype predominated (84.2%), and additional variants included CVIET+356T (10.5%) and SVMNT (5.3%). At pfmdr1, the 184F allele occurred in 57.9% of isolates, consistent with findings from day 0. Mutations at the dhfr locus were nearly fixed, the triple mutant 51I/59R/108N was present in 94.7% of isolates. At dhps, the double (437G/540E) and triple (437G/540E/581G) mutants together accounted for >85% of samples. Inferred dhfr/dhps haplotypes showed a high prevalence of quintuple and sextuple genotypes, mirroring the overall population.
Paired analysis of baseline (day 0) and follow-up (day 3) samples revealed that most isolates retained identical molecular profiles across time points. Nonetheless, we observed a few discordances, particularly in pfmdr1 (184F/Y) and dhps alleles (e.g., detection of 581G or 164L in some day 3 samples). Those differences likely reflect within-host selection of minority clones under drug pressure rather than de novo mutations.
ART-R is a growing threat to malaria control in sub-Saharan Africa, and validated pfkelch13 mutations are already established in Rwanda, Uganda, and Tanzania (8,9,13,17–20). Of particular concern is the detection of the pfkelch13 R561H mutation, a validated marker of delayed parasite clearance, in the Huye district of Rwanda and the Kagera region of Tanzania (8–10,12–14,21,22). Those findings raise the alarm over possible spread into neighboring countries, including Burundi, underscoring the urgent need for localized therapeutic and molecular surveillance.
We adopted a simplified surveillance design to detect ART-R, combining day 3 parasitemia assessment with pfkelch13 molecular screening. That approach provides complementary clinical and genetic indicators without replacing traditional therapeutic efficacy studies, which remain essential for evaluating ACT efficacy and informing treatment policy. The limited follow-up period and dried blood spot collection enable rapid geographic mapping of ART-R risk across multiple sites in resource-constrained contexts (23).
In 5 districts across northern Burundi, 4.5% of children treated with artemether/lumefantrine remained parasitemic on day 3, below the 10% WHO threshold for confirmed ART-R (24), but the Vumbi district exceeded the WHO threshold. However, the sample size was calculated for surveillance purposes and was not powered to detect statistically significant differences between districts. Consequently, the geographic comparisons we describe should be interpreted as exploratory findings, rather than definitive evidence of spatial heterogeneity.
None of the day 3 positivity cases harbored validated or candidate pfkelch13 mutations. That discordance suggests either false-positive results driven by imperfect adherence to the 6-dose artemether/lumefantrine regimen (25,26), variability in lumefantrine exposure (food intake, vomiting, drug–drug interactions), age-dependent immunity in young children (27), or genuine delayed clearance mediated by polygenic architecture of artemisinin response (e.g., coronin, ap2µ, or ubp-1) (28,29). Pharmacokinetics/adherence verification and broader genomic profiling are needed to resolve those alternatives.
The overall clinical response to artemether/lumefantrine treatment appeared satisfactory despite 4.5% day 3 positivity, likely because lumefantrine, the partner drug with a longer half-life, remains fully effective at eliminating residual parasites. The pfmdr1 mutations observed (low N86, high 184F) indicate drug pressure but are not validated markers of lumefantrine failure. Those findings suggest lumefantrine efficacy is currently maintaining artemether/lumefantrine effectiveness in Burundi, underscoring the importance of monitoring both artemisinin and partner drug susceptibility.
The absence of pfkelch13 mutants in Burundi, despite their presence in neighboring countries, might reflect intense transmission (5,7), where parasite diversity and immunity limit resistant lineage expansion. However, pfkelch13 mutations have been documented in high-transmission areas of Uganda (20), demonstrating that transmission intensity alone does not prevent resistance selection or spread. The absence of validated mutations should not be interpreted as protection. Importation risk remains high, given porous borders and regional connectivity, underscoring the urgency of cross-border surveillance (24).
Beyond ART-R, this study revealed a concerning accumulation of antifolate resistance. The dhfr triple mutant was nearly fixed, and >80% of infections carried highly resistant multilocus haplotypes. At the dhps locus, mutations were also highly prevalent. The 437G/540E/581G triple mutant was present in 47.1% of isolates, whereas only 5.8% of parasites remained wild-type. That pattern indicates widespread sulfadoxine resistance, with direct implications for malaria prevention in Burundi. Sulfadoxine/pyrimethamine is used for IPTp and more recently in PMC. The high prevalence (44%) of the sextuple haplotype raises serious concerns because that genotype is associated with reduced IPTp effectiveness and adverse birth outcomes and likely compromises PMC efficacy. Those findings warrant urgent reassessment of sulfadoxine/pyrimethamine-based prevention strategies in Burundi (30–32).
ACT partner-drug resistance markers were entrenched. The pfcrt CVIET haplotype (74I/75E/76T) predominated (84.4%), with an additional subset carrying 356T (9.3%). Wild-type CVMNK was infrequent (4.9%). Those results confirm that chloroquine resistance remains essentially fixed in the parasite population, consistent with observations across East Africa, despite chloroquine withdrawal (33,34). That pattern contrasts with observations from several other countries in Africa, where chloroquine-sensitive parasites have reexpanded after drug withdrawal (35). The persistence of CVIET in Burundi likely reflects the use of artesunate/amodiaquine as first-line treatment until 2019, which maintained selective pressure favoring pfcrt mutant parasites. Similar persistence of CVIET has been reported in other countries that used artesunate/amodiaquine, such as South Sudan. A decline in CVIET prevalence might require several more years to become apparent (36–39).
We detected no pfcrt mutations associated with piperaquine resistance (e.g., F145I, H97Y, M343L, G353V) in this study (40–42). Those mutations have been identified as key drivers of dihydroartemisinin/piperaquine treatment failures in Southeast Asia, but they appear absent in East Africa parasite populations.
At the pfmdr1 locus, the Y184F mutation was the dominant variant (57.3%), whereas the wild-type NYSND was found in 31.2%. Other variants involving 86Y, 1034C, 1042D, or 1246Y were uncommon (<1.6%). The predominance of 184F is consistent with reports from Zanzibar (Tanzania) and Senegal (43,44). The Y184F mutation is frequently selected by artemether/lumefantrine treatment and can modulate parasite responses in vitro, but it should be regarded as a marker of drug pressure rather than a validated clinical marker of lumefantrine resistance (45).
The limitations of this study include partially supervised drug treatment, unmeasured drug concentrations, day 3 follow-up only, and no assessment of copy number variation in pfmdr1 and plasmepsin 2/3, the established markers associated with reduced lumefantrine (46–48) and piperaquine (40,49,50) susceptibility. Study strengths include multiple sentinel sites in a high-risk border region, combined clinical and molecular surveillance, and an integrated baseline dataset informing national and regional strategies.
WHO recommends deploying multiple first-line ACTs to delay malaria drug resistance emergence. In Burundi, where artemether/lumefantrine is the sole first-line treatment, introducing an alternative such as dihydroartemisinin/piperaquine could reduce selective pressure. The absence of pfcrt mutations associated with piperaquine resistance suggests dihydroartemisinin/piperaquine could retain full efficacy and serve as a second first-line option.
In conclusion, our findings highlight a critical juncture for Burundi. We detected no pfkelch13 mutants, but excess localized day 3 positivity and entrenched sulfadoxine/pyrimethamine resistance represent clear warning signals. High-transmission dynamics may currently protect Burundi against pfkelch13-mediated ART-R, but that protection is fragile. Strengthened therapeutic efficacy monitoring, expanded molecular surveillance, including genomic approaches, and cross-border collaboration are urgently required to prevent resistant P. falciparum establishment and spread in the Great Lakes region of Burundi.
Dr. Niyomwungere is a physician and public health specialist at the Institut National de Santé Publique in Bujumbura, Burundi. His work focuses on malaria control and surveillance of malaria drug resistance to support national elimination efforts.
Acknowledgments
We sincerely thank all the children and their caregivers who participated in this study. We are also grateful to the clinical and laboratory teams across the 8 participating health facilities for their dedication and professionalism. We acknowledge the support of the Burundian Ministry of Health, the National Malaria Control Program, and local health authorities for facilitating field implementation. We also thank the staff of the National Reference Laboratory of Institut National de Santé Publique in Burundi and community health workers for their valuable contributions.
The datasets used and analyzed during this study are available from the corresponding author upon reasonable request. The nucleotide sequences generated in this study have been deposited in GenBank (accession nos. PZ122655–9; BankIt submission no. 3061318).
This study was conducted as part of the IMPRIMA project and received support from the Global Health EDCTP3 Joint Undertaking (GH EDCTP3 JU; grant agreement no. 101103213). Additional funding was provided by the Fondation pour la Recherche Médicale (FRM) through the ‘Équipes FRM 2024’ program (grant no. EQU202403018026), by the Université de Strasbourg (grant no. R42PHAV), and by the Institut Universitaire de France under the Senior Chair program (2024–2029). The funders had no role in the study design, data collection and analysis, interpretation of results, or writing of the manuscript.
Author contributions: conceptualization: D.N., P.S., D.M., J.N.; methodology: E.C., P.-E.S., L.T., Y.N.G.T., D.N., D.M.; investigation: P.S., D.N., J.N., L. Mugisha, N.N., D.S., M.N., V.N., O.N., J.-C.N., A.W., C.B., J.-L.B., N.N., M.N., F.N., N.N., P.H., P.N., L. Ma, J.R.; data curation: D.N., P.S., D.M.; formal analysis: D.N., P.S., D.M.; data access and verification: D.N., P.S., D.M.; writing, original draft: D.N., D.M.; writing, review and editing: D.N., D.M.; supervision: D.M., J.N.; project administration: D.N., J.N.; funding acquisition: D.M., J.N. All authors read and approved the final version of the manuscript. D.M. and D.N. are joint guarantors of the study.
References
- World Health Organization. World malaria report 2025: addressing the threat of antimalarial drug resistance. Geneva: The Organization; 2025.
- van Eijk AM, Hill J, Noor AM, Snow RW, ter Kuile FO. Prevalence of malaria infection in pregnant women compared with children for tracking malaria transmission in sub-Saharan Africa: a systematic review and meta-analysis. Lancet Glob Health. 2015;3:e617–28. DOIPubMedGoogle Scholar
- World Health Organization. Global technical strategy for malaria 2016–2030. 2021 update. Geneva: The Organization; 2021.
- Programme National Intégré de Lutte contre le Paludisme. Guidelines for malaria case management in Burundi [in French]. Bujumbura (Burundi): Ministère de la Santé Publique; 2019.
- US President’s Malaria Initiative. Burundi malaria profile. Washington: The Initiative; 2023.
- Programme National Intégré de Lutte contre le Paludisme. National strategic plan for the fight against malaria (2021–2027) [in French]. Bujumbura (Burundi): Ministère de la Santé Publique; 2021.
- Sinzinkayo D, Baza D, Gnanguenon V, Koepfli C. The lead-up to epidemic transmission: malaria trends and control interventions in Burundi 2000 to 2019. Malar J. 2021;20:298. DOIPubMedGoogle Scholar
- Uwimana A, Legrand E, Stokes BH, Ndikumana JM, Warsame M, Umulisa N, et al. Emergence and clonal expansion of in vitro artemisinin-resistant Plasmodium falciparum kelch13 R561H mutant parasites in Rwanda. Nat Med. 2020;26:1602–8. DOIPubMedGoogle Scholar
- Uwimana A, Umulisa N, Venkatesan M, Svigel SS, Zhou Z, Munyaneza T, et al. Association of Plasmodium falciparum kelch13 R561H genotypes with delayed parasite clearance in Rwanda: an open-label, single-arm, multicentre, therapeutic efficacy study. Lancet Infect Dis. 2021;21:1120–8. DOIPubMedGoogle Scholar
- Bergmann C, van Loon W, Habarugira F, Tacoli C, Jäger JC, Savelsberg D, et al. Increase in Kelch 13 polymorphisms in Plasmodium falciparum, southern Rwanda. Emerg Infect Dis. 2021;27:294–6. DOIPubMedGoogle Scholar
- Kirby R, Giesbrecht D, Karema C, Watson O, Lewis S, Munyaneza T, et al. Examining the early distribution of the artemisinin-resistant Plasmodium falciparum kelch13 R561H mutation in areas of higher transmission in Rwanda. Open Forum Infect Dis. 2023;10:ofad149. DOIGoogle Scholar
- Assey A, Scialabba S, Zinga MM, Koliopoulos P, van Loon W, Minja CA, et al. Artemisinin and partner drug resistance markers in Plasmodium falciparum from Tanzanian paediatric malaria patients, 2016-2022. Malar J. 2025;24:209. DOIPubMedGoogle Scholar
- Ishengoma DS, Mandara CI, Bakari C, Fola AA, Madebe RA, Seth MD, et al. Evidence of artemisinin partial resistance in northwestern Tanzania: clinical and molecular markers of resistance. Lancet Infect Dis. 2024;24:1225–33. DOIPubMedGoogle Scholar
- Juliano JJ, Giesbrecht DJ, Simkin A, Fola AA, Lyimo BM, Pereus D, et al. Prevalence of mutations associated with artemisinin partial resistance and sulfadoxine-pyrimethamine resistance in 13 regions in Tanzania in 2021: a cross-sectional survey. Lancet Microbe. 2024;5:
100920 . DOIPubMedGoogle Scholar - Institut de Statistiques et d’Études Économiques du Burundi, The DHS Program. Third demographic and health survey in Burundi (EDSB-III), 2016–2017 [in French]. Bujumbura (Burundi): The Institute; 2017.
- Canier L, Khim N, Kim S, Sluydts V, Heng S, Dourng D, et al. An innovative tool for moving malaria PCR detection of parasite reservoir into the field. Malar J. 2013;12:405. DOIPubMedGoogle Scholar
- Mihreteab S, Platon L, Berhane A, Stokes BH, Warsame M, Campagne P, et al. Increasing prevalence of artemisinin-resistant HRP2-negative malaria in Eritrea. N Engl J Med. 2023;389:1191–202. DOIPubMedGoogle Scholar
- Balikagala B, Fukuda N, Ikeda M, Katuro OT, Tachibana SI, Yamauchi M, et al. Evidence of artemisinin-resistant malaria in Africa. N Engl J Med. 2021;385:1163–71. DOIPubMedGoogle Scholar
- Fukuda N, Tachibana SI, Ikeda M, Sakurai-Yatsushiro M, Balikagala B, Katuro OT, et al. Ex vivo susceptibility of Plasmodium falciparum to antimalarial drugs in Northern Uganda. Parasitol Int. 2021;81:
102277 . DOIPubMedGoogle Scholar - Rosenthal PJ, Asua V, Conrad MD. Emergence, transmission dynamics and mechanisms of artemisinin partial resistance in malaria parasites in Africa. Nat Rev Microbiol. 2024;22:373–84. DOIPubMedGoogle Scholar
- Kayiba NK, Yobi DM, Tshibangu-Kabamba E, Tuan VP, Yamaoka Y, Devleesschauwer B, et al. Spatial and molecular mapping of Pfkelch13 gene polymorphism in Africa in the era of emerging Plasmodium falciparum resistance to artemisinin: a systematic review. Lancet Infect Dis. 2021;21:e82–92. DOIPubMedGoogle Scholar
- Straimer J, Gandhi P, Renner KC, Schmitt EK. High prevalence of Plasmodium falciparum K13 mutations in Rwanda is associated with slow parasite clearance after treatment with artemether-lumefantrine. J Infect Dis. 2022;225:1411–4. DOIPubMedGoogle Scholar
- World Health Organization. Report on antimalarial drug efficacy, resistance and response: 10 years of surveillance (2010–2019). Geneva: The Organization; 2020.
- World Health Organization. Strategy to respond to antimalarial drug resistance in Africa. Geneva: The Organization; 2022.
- Banek K, Lalani M, Staedke SG, Chandramohan D. Adherence to artemisinin-based combination therapy for the treatment of malaria: a systematic review of the evidence. Malar J. 2014;13:7. DOIPubMedGoogle Scholar
- Challenger JD, Bruxvoort K, Ghani AC, Okell LC. Assessing the impact of imperfect adherence to artemether-lumefantrine on malaria treatment outcomes using within-host modelling. Nat Commun. 2017;8:1373. DOIPubMedGoogle Scholar
- Khoury DS, Cromer D, Akter J, Sebina I, Elliott T, Thomas BS, et al. Host-mediated impairment of parasite maturation during blood-stage Plasmodium infection. Proc Natl Acad Sci U S A. 2017;114:7701–6. DOIPubMedGoogle Scholar
- Mukherjee A, Bopp S, Magistrado P, Wong W, Daniels R, Demas A, et al. Artemisinin resistance without pfkelch13 mutations in Plasmodium falciparum isolates from Cambodia. Malar J. 2017;16:195. DOIPubMedGoogle Scholar
- Demas AR, Sharma AI, Wong W, Early AM, Redmond S, Bopp S, et al. Mutations in Plasmodium falciparum actin-binding protein coronin confer reduced artemisinin susceptibility. Proc Natl Acad Sci U S A. 2018;115:12799–804. DOIPubMedGoogle Scholar
- Hansson H, Minja DTR, Moeller SL, Lusingu JPA, Bygbjerg IC, Yde AM, et al. Reduced birth weight caused by sextuple drug-resistant Plasmodium falciparum infection in early second trimester. J Infect Dis. 2021;224:1605–13. DOIPubMedGoogle Scholar
- Naidoo I, Roper C. Mapping ‘partially resistant’, ‘fully resistant’, and ‘super resistant’ malaria. Trends Parasitol. 2013;29:505–15. DOIPubMedGoogle Scholar
- Plowe CV. Malaria chemoprevention and drug resistance: a review of the literature and policy implications. Malar J. 2022;21:104. DOIPubMedGoogle Scholar
- Hassen J, Alemayehu GS, Dinka H, Golassa L. High prevalence of Pfcrt 76T and Pfmdr1 N86 genotypes in malaria infected patients attending health facilities in East Shewa zone, Oromia Regional State, Ethiopia. Malar J. 2022;21:286. DOIPubMedGoogle Scholar
- Mongella S, Enweji N, Mnong’one N, Minde M, Kamugisha E, Swedberg G. High prevalence of Plasmodium falciparum pfcrt K76T mutation in children with sickle cell disease at a tertiary hospital in north-western Tanzania. Tanzan J Health Res. 2014;16:256–60. DOIPubMedGoogle Scholar
- Laufer MK, Thesing PC, Eddington ND, Masonga R, Dzinjalamala FK, Takala SL, et al. Return of chloroquine antimalarial efficacy in Malawi. N Engl J Med. 2006;355:1959–66. DOIPubMedGoogle Scholar
- Dagnogo O, Ako AB, Ouattara L, Dago ND, Coulibaly DN, Touré AO, et al. Towards a re-emergence of chloroquine sensitivity in Côte d’Ivoire? Malar J. 2018;17:413. DOIPubMedGoogle Scholar
- Laufer MK, Takala-Harrison S, Dzinjalamala FK, Stine OC, Taylor TE, Plowe CV. Return of chloroquine-susceptible falciparum malaria in Malawi was a reexpansion of diverse susceptible parasites. J Infect Dis. 2010;202:801–8. DOIPubMedGoogle Scholar
- Mwanza S, Joshi S, Nambozi M, Chileshe J, Malunga P, Kabuya JB, et al. The return of chloroquine-susceptible Plasmodium falciparum malaria in Zambia. Malar J. 2016;15:584. DOIPubMedGoogle Scholar
- Ndam NT, Basco LK, Ngane VF, Ayouba A, Ngolle EM, Deloron P, et al. Reemergence of chloroquine-sensitive pfcrt K76 Plasmodium falciparum genotype in southeastern Cameroon. Malar J. 2017;16:130. DOIPubMedGoogle Scholar
- Amato R, Lim P, Miotto O, Amaratunga C, Dek D, Pearson RD, et al. Genetic markers associated with dihydroartemisinin-piperaquine failure in Plasmodium falciparum malaria in Cambodia: a genotype-phenotype association study. Lancet Infect Dis. 2017;17:164–73. DOIPubMedGoogle Scholar
- Dhingra SK, Small-Saunders JL, Ménard D, Fidock DA. Plasmodium falciparum resistance to piperaquine driven by PfCRT. Lancet Infect Dis. 2019;19:1168–9. DOIPubMedGoogle Scholar
- Ross LS, Dhingra SK, Mok S, Yeo T, Wicht KJ, Kümpornsin K, et al. Emerging Southeast Asian PfCRT mutations confer Plasmodium falciparum resistance to the first-line antimalarial piperaquine. Nat Commun. 2018;9:3314. DOIPubMedGoogle Scholar
- Connelly SV, Brazeau NF, Msellem M, Ngasala BE, Aydemir O, Goel V, et al. Strong isolation by distance and evidence of population microstructure reflect ongoing Plasmodium falciparum transmission in Zanzibar. eLife. 2024;12:RP90173. DOIGoogle Scholar
- Ahouidi A, Oliveira R, Lobo L, Diedhiou C, Mboup S, Nogueira F. Prevalence of pfk13 and pfmdr1 polymorphisms in Bounkiling, Southern Senegal. PLoS One. 2021;16:
e0249357 . DOIPubMedGoogle Scholar - Veiga MI, Dhingra SK, Henrich PP, Straimer J, Gnädig N, Uhlemann AC, et al. Globally prevalent PfMDR1 mutations modulate Plasmodium falciparum susceptibility to artemisinin-based combination therapies. Nat Commun. 2016;7:11553. DOIPubMedGoogle Scholar
- Fançony C, Fortes-Gabriel E, Zage F, Alexiou E, Broumou I, Pernaute-Lau L, et al. Artemether-lumefantrine treatment selects Plasmodium falciparum multidrug resistance 1 (pfmdr1) increased copy number among African malaria infections. J Infect Dis. 2025;231:e1119–28. DOIPubMedGoogle Scholar
- Sidhu AB, Uhlemann AC, Valderramos SG, Valderramos JC, Krishna S, Fidock DA. Decreasing pfmdr1 copy number in plasmodium falciparum malaria heightens susceptibility to mefloquine, lumefantrine, halofantrine, quinine, and artemisinin. J Infect Dis. 2006;194:528–35. DOIPubMedGoogle Scholar
- Tumwebaze PK, Conrad MD, Okitwi M, Orena S, Byaruhanga O, Katairo T, et al. Decreased susceptibility of Plasmodium falciparum to both dihydroartemisinin and lumefantrine in northern Uganda. Nat Commun. 2022;13:6353. DOIPubMedGoogle Scholar
- Bopp S, Magistrado P, Wong W, Schaffner SF, Mukherjee A, Lim P, et al. Plasmepsin II-III copy number accounts for bimodal piperaquine resistance among Cambodian Plasmodium falciparum. Nat Commun. 2018;9:1769. DOIPubMedGoogle Scholar
- Witkowski B, Duru V, Khim N, Ross LS, Saintpierre B, Beghain J, et al. A surrogate marker of piperaquine-resistant Plasmodium falciparum malaria: a phenotype-genotype association study. Lancet Infect Dis. 2017;17:174–83. DOIPubMedGoogle Scholar
Figure
Table
Cite This ArticleOriginal Publication Date: March 31, 2026
Table of Contents – Volume 32, Number 4—April 2026
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Denis Niyomwungere, Institut National de Santé Publique, Avenue de l’Hôpital, BP 6807, Bujumbura, Burundi
Top