Disclaimer: Early release articles are not considered as final versions. Any changes will be reflected in the online version in the month the article is officially released.
Volume 32, Number 5—May 2026
Dispatch
Replication Efficiency of Contemporary Highly Pathogenic Avian Influenza A(H5N1) Virus Isolates in Human Nasal Epithelium Model
Figure 3
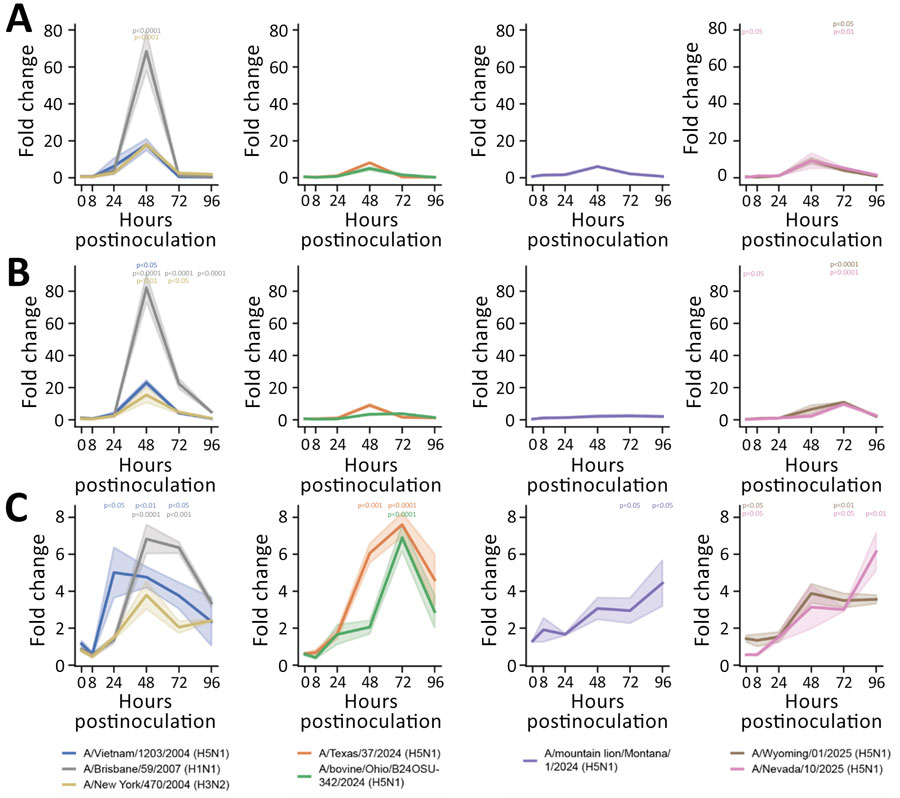
Figure 3. Induction of proinflammatory cytokines in human nasal epithelium infected with seasonal influenza A and highly pathogenic avian influenza (HPAI) A(H5N1) viruses in study of replication efficiency of contemporary HPAI H5N1 virus isolates in human nasal epithelium model. We inoculated Mattek EpiNasal cultures (https://www.mattek.com) with a multiplicity of infection of 0.1 50% tissue culture infectious dose per cell. RNA was extracted from cells at 0, 8, 24, 48, 72, and 96 hours postinoculation. We ran quantitative reverse transcription PCR using primers (Integrated DNA Technologies, https://www.idtdna.com) to detect interleukin 6 (A), tumor necrosis factor α (B), and interleukin 1β (C) for historical and clade 2.3.4.4b HPAI H5N1 genotype B3.13, B3.6, and D1.1 strains. Lines indicate medians; shading indicates 95% CIs. We normalized data to internal controls (ACTB and GAPDH) and calculated fold change relative to timepoint-matched mock-infected controls. Fold change is reported for 3 biologic replicates. We performed statistical analysis using 2-way analysis of variance with Dunnett posttest. p values are shown by asterisks in colors matching isolates: *p<0.05; **p<0.01; ***p<0.001; ****p<0.0001.
1These authors contributed equally to this article.
2Current affiliation: LSU Health, Shreveport, Louisiana, USA.