Volume 4, Number 1—March 1998
Perspective
Calicivirus Emergence from Ocean Reservoirs: Zoonotic and Interspecies Movements
Cite This Article
Citation for Media
Abstract
Caliciviral infections in humans, among the most common causes of viral-induced vomiting and diarrhea, are caused by the Norwalk group of small round structured viruses, the Sapporo caliciviruses, and the hepatitis E agent. Human caliciviruses have been resistant to in vitro cultivation, and direct study of their origins and reservoirs outside infected humans or water and foods (such as shellfish contaminated with human sewage) has been difficult. Modes of transmission, other than direct fecal-oral routes, are not well understood. In contrast, animal viruses found in ocean reservoirs, which make up a second calicivirus group, can be cultivated in vitro. These viruses can emerge and infect terrestrial hosts, including humans. This article reviews the history of animal caliciviruses, their eventual recognition as zoonotic agents, and their potential usefulness as a predictive model for noncultivatable human and other animal caliciviruses (e.g., those seen in association with rabbit hemorrhagic disease).
In vitro cultivation of caliciviruses indicates that these pathogens have been emerging periodically from ocean sources for 65 years (1). The best-documented example of ocean caliciviruses causing disease in terrestrial species is the animal disease vesicular exanthema of swine (VES) (1). Feline calicivirus (the only member of the group with a seemingly ubiquitous and continuous terrestrial presence) also appears to have ocean reservoirs (2). The source of caliciviruses causing gastroenteritis in humans is frequently shellfish, which do not always come from beds contaminated with human waste (3,4). The origins of hepatitis E are often obscure, but water is one suspected source (5). The most recent emerging calicivirus is associated with rabbit hemorrhagic disease (RHD), and although an ocean association has not been reported, the agent readily moves between continents and crosses ocean channels (6). Finally, the only reported in vitro isolation and sequential propagation of a calicivirus pathogenic for humans is a virus residing in the sea (7).
The Caliciviridae are divided into five groups, tentatively designated distinct genera, on the basis of sequence relatedness and genomic organization (8). Four are known human pathogens—Sapporo, Norwalk-like small round structured viruses, hepatitis E, and the marine (animal) caliciviruses—while the fifth group, which includes RHD virus, is not yet proven to be a human pathogen. The human Sapporo viruses are more closely related to the marine caliciviruses than to the other human group causing gastroenteritis, the Norwalk-like viruses. On the basis of homology and genomic organization, RHD virus falls between these two groups. In addition, the genomic organization of hepatitis E is most closely related to that of the only other hepatotropic calicivirus currently described, RHD virus (8).
Many marine calicivirus strains in the tentative genus of VES virus-like caliciviruses have been passaged in vitro; their characterization has facilitated understanding of calicivirus geographic distribution and host versatility (1). Dozens of serotypes were described on the basis of serum neutralization tests; this antigenic complexity complicated serodiagnosis and hampered studies of effects on host species.
The illnesses associated with two recently discovered viruses classified as Caliciviridae, hepatitis E virus and RHD virus, have altered the notion that caliciviruses produce only transient clinical disease but not death (1,9,10). Hepatitis E virus is fatal for 25% of the pregnant women in developing countries who contract hepatitis E (11); RHD can kill 95% of infected rabbits within 24 to 48 hours of exposure (6).
The oceans are reservoirs in which caliciviruses are exposed in a water substrate to life forms from zooplankton to whales and in which they, like other RNA viruses, can amplify to very high numbers with variants occurring in every replicative cycle (12). Such a varied replicative setting has served this parasite well. With the right tools, evidence of previous infection with caliciviruses can often be shown in fish, avian, and many mammalian species, including humans (1). It is not known why some caliciviruses have become potential hemorrhagic agents associated with purpura hemorrhagica in aborted piglets (13), neonatal hemorrhagic syndrome in pinnipeds (A.W. Smith, D.E. Skilling, unpub. data), hemorrhagic disease in fatal hepatitis E in humans (5), and RHD (6). However, it is known that caliciviral diseases can be difficult or impossible to contain and eradicate. Pathogenic caliciviruses can be expected to continue emerging from the sea in unexpected forms at unexpected times in unexpected places. Studying those that have emerged and are compliant to in vitro propagation can provide insights into those that cannot be cell-culture adapted and those yet to be discovered.
The 66-year history of the caliciviruses with ocean reservoirs can be divided into three periods: 1932 to 1972, the species-specific era (14); 1972 to 1982, the new era of virology, during which oceans were first found to be reservoirs of viral disease infecting domestic animals (7,9); and 1976 to the present.
The first evidence of infection with caliciviruses of marine origin can be traced to 1932. A large herd of swine in Orange County, California, was being fed raw garbage collected from restaurants and institutions in the Los Angeles area. When some animals became sick with vesicular lesions on the feet and nose, regulatory veterinarians were notified because vesicular diseases of livestock were reportable. The farm with sick swine and adjacent farms were quarantined for foot-and-mouth disease, and more than 19,000 head of exposed cattle and swine were destroyed and buried in quicklime (14). The outbreak was contained. One year later and 100 miles to the south in San Diego, California, the second known outbreak occurred and was contained (14). This time the disease was found not to be foot-and-mouth disease, because the virus would not infect cattle, but instead was described as a new disease of swine and was called VES (14). In 1934, a third outbreak occurred in San Francisco, California, and VES was again contained (14). In 1935, the events repeated themselves, but from 1936 through mid-December 1939, the disease disappeared and then abruptly reappeared, at times involving 40% of California swine herds. All of these outbreaks in the 1930s and 1940s were shown by cross-infectivity studies to be caused by many distinct but related VES virus strains.
The embargoes placed on raw California pork were successful in containing VES within California until 1952. That year, a passenger train between San Francisco and Chicago served California pork and discarded the raw pork trimmings into the garbage in Cheyenne, Wyoming; the garbage was fed to swine subsequently redistributed by auction sale yard. Within 14 months, all major swine-growing areas in the United States (41 states) had reported VES. For the first time, the federal government, rather than just the California state government, activated eradication and quarantine measures against VES, including enforcement of federal laws requiring garbage to be cooked before it was fed to swine. By 1956, the last reported outbreak of VES had been contained, and the disease was said to have been eradicated. In 1959, VES was declared a foreign animal disease, even though it had never been reported outside the United States (14).
Forty years later, the natural history of this calicivirus still contained few details. Its origins were not known but were said to have been de novo or from some unknown wild animal reservoir, which was extensively sought but not found (14,15). Swine were the only naturally infected host species; no evidence of human infection had been observed (14). Control of VES had been a notable success story for regulatory veterinary medicine in the United States; within 24 years of its discovery as an entirely new disease, it was said to have been eradicated (14). Internationally accepted animal disease diagnostic tests using swine, horse, and bovine infectivity profiles for vesicular stomatitis virus, foot-and-mouth virus, and VES virus were used routinely; VES virus did not infect cattle, whereas the other viruses did (14,15).
The discovery of a new paradigm in "viral traffic" (16) began the second period of calicivirus history. The movement of a member of the Caliciviridae from ocean reservoirs to terrestrial hosts changed the understanding of the natural history of a virus thought to be host specific and eradicated (1,15,17).
The first virus isolate from a pinniped occurred in 1972. The agent, named San Miguel sea lion virus type 1 (SMSV-1), was a calicivirus that caused classic VES in swine (17). Thus began a series of isolating and characterizing viruses in ocean species that were officially designated as "viruses indistinguishable from VES virus" because they were additional VES virus types. They could not be called VES virus (18) since VES had been officially eradicated. Should the VES virus reappear, its status as a foreign animal disease would mandate immediate implementation of eradication measures; eradication was viewed as an impossibility because of the wide range of reservoirs for VES virus, both migratory and ocean species. The 13 marine caliciviruses serotypes (100 TCID50 vs. 20 units of neutralizing antibody) isolated from swine before 1956 were called VES viruses; those isolated after 1972 have been designated SMSV-1 through 17 or given more proper nomenclature, e.g., the pygmy chimpanzee isolate (primate calicivirus Pan paniscus type 1) (1,19).
By 1982, 11 species of pinnipeds and cetaceans of the North Pacific Ocean and Bering Sea (monk seals, California sea lions, northern sea lions, northern elephant seals, northern fur seals, walrus, gray whales, sei whales, sperm whales, bowhead whales, and Pacific bottlenosed dolphins) were known to be susceptible to calicivirus infection, as was an ocean fish, the opaleye perch (Girella nigricans) (20). Furthermore, in many instances, the virus had crossed the intertidal zone to infect terrestrial species (18). On the basis of these data and the established ocean ranges of known calicivirus host species, the shores of Mexico, the United States, Canada, Russia, Korea, Japan, China, and perhaps others bordering the North Pacific Ocean had been regularly exposed to large numbers of marine caliciviruses with unknown host ranges and tissue trophisms (21). By this time, type-specific neutralizing antibodies to two of four serotypes tested were reported in human patients in the United States (22). Cumulatively, these findings lead to the conclusion that fish and perhaps other ocean products provide a vehicle for transmission of these marine caliciviruses to terrestrial animals.
The magnitude of potential exposure to marine caliciviruses from the sea is substantial. For example, a 35-ton gray whale, shown by electron microscopy to have more than 106 caliciviruses per gram of feces, can eat 5% or more of its body weight per day and eliminate an equivalent quantity of feces containing an estimated 1013 caliciviruses daily. Marine caliciviruses remain viable more than 14 days in 15°C seawater (20).
Although marine mammals were often infected, fish and fish products were more likely to transport the virus from sea to land (23). In contrast to the 1932 to 1936 introductions of VES from raw fish, the rapid and uncontrolled spread of VES virus throughout California after 1939 and then across the United States in 1952 was a pig-to-pig cycle through raw garbage feed. However, new virus serotypes were also introduced through feeding raw fish scraps to swine (1,15). That VES was not a species-specific disease became accepted, but the possibility of human infection, although suspected (22), was largely untested.
During the third historical period (1976 to present), which overlaps with the second, viral traffic across the land/sea interface has been observed repeatedly, as the following examples show. A calicivirus isolated from an opaleye perch and designated SMSV-7 produced fulminating VES in exposed swine and spread from pig to pig by contact transmission (23). A reptilian calicivirus Crotalus-1 was isolated from three species of snakes and one species of amphibian (24) and from three species of marine mammals whose population distributions spanned the North Pacific from Mexico to the Bering Sea (1). Mink fed a diet of ground-up calicivirus-infected seal meat became infected with VES virus (25). Parasitic nematode larvae from California sea lions in San Diego, California, were used to infect opaleye fish with calicivirus (SMSV-5); when the fish were killed and fed to Northern fur seals on the Pribilof Islands in the Bering Sea 30 days later, the seals developed vesicular disease, and the virus was recovered from the lesions (1). Shellfish resident to the tidal zone were exposed to marine caliciviruses and held at less than 10°C in a continuous flow of sterile seawater. The caliciviruses were reisolated 60 days later in mammalian cell lines (26). When tested with a cDNA calicivirus group-specific hybridization probe from a marine calicivirus (SMSV-5), some shellfish beds on U.S. coasts were positive for caliciviruses of unknown type (26). Feline calicivirus was shown to cause disease not only in dogs, but also in seals (on the basis of 17 of 20 adult sea lions having neutralizing antibody to FCV-F9 with titers of 1:15 to 1:220). Only 11 of 20 of these sea lions had neutralizing antibody to a sea lion isolate, SMSV-13 (titer 1:10). This demonstrates a probable feline calicivirus ocean presence in California sea lions (2). In swine, a so-called mystery pig disease (porcine respiratory reproductive syndrome) was reproduced in pregnant sows in 1992 with a three-plaque passage purified cytolytic calicivirus isolate from stillborn piglets with mystery pig disease (13). A second calicivirus serotype isolated from the same piglets was the same as that isolated from walruses in 1976 (1). A white tern (Gygis alba rothschildi), a migratory sea bird sampled in the mid-Pacific (French Frigate Shoals), had a blistering disease caused by a calicivirus (27). In the first fully documented human case of clinical disease caused by a marine calicivirus, SMSV-5 was isolated from blisters on the hands and feet of a patient (7). A second, less well-documented, case involved a field biologist who was handling sea lions and developed severe facial blistering. An untypable calicivirus was isolated in tissue culture (Vero cells) from throat washings (7).
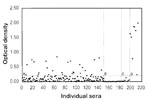
The extent of human disease is not known because test reagents are not readily available and diagnosticians are not alerted to caliciviral causes of human disease, except for diarrhea and occasionally hepatitis. However, evidence of human exposure was shown when 150 serum specimens from normal blood destined for donor use were tested. The samples were antibody-negative for hepatitis B surface and core antigen, HIV-1 and -2, HIV P-24 antigen, human T-cell lymphotropic virus Type 2, and hepatitis C virus. Approximately 19% had antibodies reactive to a polyvalent antigen made up of equal quantities of cesium chloride-banded SMSV-5, 13, and 17. (Figure 1A). To demonstrate that these reactions were not cross-reactions to the human Norwalk calicivirus antibody, serum samples from eight persons with Norwalk virus-induced diarrhea were also tested. Both acute- and convalescent-phase serum specimens were tested by enzyme-linked immunosorbent assay with the same SMSV antigens and Norwalk capsid protein (Figure 1B, C, and D). Although cross-reactivity was not detected, the serum samples may still have been able to cross-react with the calicivirus causing hepatitis E or Sapporo calicivirus, which were not tested. However, sera typed to all 40 marine caliciviruses reacted negatively when tested against Sapporo antigen (data not shown). These results suggest possible human exposure and antigenic response to marine caliciviruses. If that is not the case, such results present a confusing diagnostic picture of calicivirus exposure and diagnosis of human disease.
Organism Characteristics
The history of marine caliciviruses demonstrates that their biologic properties have great plasticity. The VES virus-like caliciviruses can replicate at temperatures of 15°C to 39°C, have diverse tissue trophisms, and travel by land (terrestrial reptiles, amphibians, and mammals), sea (pinnipeds, cetaceans, teleosts, and perhaps filter-feeding mollusks), and air (pelagic birds, e.g., the white tern). They can persist in nonlytic cycles in many reservoir hosts, and they have a wide diversity of successful antigenic types (1) (more than 40 serotypes on the basis of virus neutralization, e.g., no cross-protection between types). Their cup-like surface morphology is characteristic (Figure 2). Finally, they are zoonotic: this is a paradigm shift (7). No other virus has been shown to have its origins and primary reservoirs in the sea yet emerge to cause disease in humans.
To measure calicivirus adaptivity and preclude strong presumptions of host specificity on the basis of calicivirus type or species of origin, the following list of 16 hosts is given for a single virus serotype, SMSV-5: known natural hosts-five genera of seals, cattle, three genera of whales, donkeys, fox, and humans-and susceptible hostso-paleye fish, horses, domestic swine, and primates (1). The lists are still growing. SMSV-5 can also persist for 60 days in shellfish, but infectivity has not been measured (26). Feline calicivirus (FCV-F9) has an apparent ocean presence among California sea lions and is not host-specific (2). All members of the family Felidae are susceptible to infection, not just domestic cats. In addition, cheetahs are susceptible; the agent has naturally infected and caused disease in dogs and experimentally infected coyotes (28,29). Reports of human antibody against the feline virus suggest zoonotic potential for the feline calicivirus (30).
The broad host range and diverse mechanisms of transmission and survival of marine caliciviruses are expected of an RNA virus quasispecies (12). If structural simplicity associated with a capsid made up of a single protein species and replicative strategies conserved across rather broad tissue and phylogenetic distances is a measure, caliciviruses are primitive RNA viruses. Caliciviral RNA replicative mechanisms are thus expected to generate numerous mutants (perhaps as high as one to 10 per template copy (12), which will come in contact with many pelagic and terrestrial biota. Opportunity exists to form clusters of virus adapted across a diversity of life forms. Actual mutation rates have not been demonstrated for the Caliciviridae, but plaque-size reversion studies have found that the mutation rate for this phenomenon is one per 106 replicates (14,15). In addition, the expected versatility from RNA virus replicative infidelity and the resulting successful adaptive mechanisms are manifested in the wide spectrum of calicivirus tissue trophisms.
Disease conditions involving calicivirus tissue trophisms include blistering of the skin (particularly on the appendages and around the mouth), pneumonia, abortion, encephalitis, myocarditis, myositis, hepatitis, diarrhea, and coagulation/hemorrhage (1,3,5,7; Table 1). Caliciviruses have the inherent potential and adaptive mechanisms to successfully parasitize essentially all organ systems of the many animal species that have been examined in detail.
Calicivirus disease manifestations in animals will likely continue but will only become well defined with improved diagnostic means. With cultivatable marine caliciviruses as models, the role of disease-causing caliciviruses can be further defined. Now caliciviruses infecting humans can only be visualized by electron microscopy or histochemistry but cannot be propagated in vitro. Thus, miscarriage and birth defects in human patients, hepatitis other than types A through G, hand-foot-and-mouth–like diseases, viral myocarditis, viral encephalitis of unknown etiology, and joint and muscle disease, for example, should be examined for caliciviruses when other causes of disease are not found.
In the absence of data, extrapolating from cultivatable caliciviruses to predict future effects of poorly characterized caliciviruses should be useful, particularly when there is an urgent need to assess possible human risk. The calicivirus implicated in RHD is a case in point for it might be expected to infect humans.
Additional evidence exists. An anecdotal account mentions a Mexican worker who developed antibodies to RHD while eradicating the disease in Mexico (31). An Australian study designed to assess the risk for illness after RHD escaped from Wardang Island (32) examined a group of 269 persons (153 reporting exposure to rabbits or samples infected with RHD virus and 116 reporting no known RHD virus contact) from two Australian states with the greatest amount of RHD virus activity in rabbits. Exposure was categorized by degree of skin exposure to infected materials. Date of first exposure was noted, but no cumulative exposure index was developed. A "high" exposure category could derive from one exposure, and "low" exposure categories could include multiple exposures, each with low exposure. Symptoms were assessed by recall of illness over the previous 13 months. Because the RHD agent was in high security containment facilities for the first 3 months of the recall period and geographically confined for the following 3 months, that period was considered a low exposure period. Because of the rapid spread of the virus in the two states, the last 6 months of the recall period were considered the high exposure period.
The data (Table 2) show the rate ratios for the occurrence of different illness in the two periods. All rate ratios were considerably greater than 1.00, and the rate ratios for any illness, diarrhea/gastroenteritis, flu/fever, and neurologic illness are significant (p < 0.005). Because each group contained health histories for 3 spring months or 3 autumn months, 1 summer month, and 2 winter months, the data are seasonally adjusted; hence, winter illness does not explain the excess symptoms observed in the high exposure group, and RHD virus exposure remains a plausible explanation for increased disease incidence.
It is difficult to produce pure cultures of noncultivatable caliciviruses to carry out Koch's postulates and establish cause and effect for a single pathogen strain or species. For RHD, both a calicivirus and a parvovirus have been identified in ill rabbits, and a parvovirus has been isolated in vitro and reported to fulfill Koch's postulates (33-35). Yet, caliciviruses have been purified from infected organs to the limits of purity by physical means, and those preparations also cause RHD (35). The caliciviruses purified by physical means cannot be proven to be free of contaminating agents, such as parvovirus (35). If RHD is parvovirus-driven, extrapolation from what is known of other small DNA viruses suggests a rather stable genome and a reduced host range with less likelihood of new host relationships (12). On the other hand, if calicivirus is the primary pathogen, the genomic infidelity that occurs during small RNA virus replication and the documented cross-species transmission of the cultivatable caliciviruses suggest that RHD might also move across species barriers (1,12).
Adequate diagnostic reagents for epidemiologic studies need to be made available; they include antigens, monoclonal antibodies, polymerase chain reaction primer sets, and cDNA probes based on group epitopes. In addition, biotype- or pathotype-specific reagents are needed to differentiate pathogenic from nonpathogenic infections.
The future also holds the confounding problem of vaccines. Although vaccines can be produced, because of calicivirus antigenic diversity, their efficacy would be predictably short-lived and marginal. Other approaches will need to be sought. Conserved traits that render the Caliciviridae viable as a virus with certain predictable genomic expressions must be sought, and if they exist, targeted for immune attack.
Only one of the five known calicivirus groups can be grown in vitro and subjected to the full range of host-parasite tests and conditions necessary to more fully define a virus in nature. Therefore, extrapolations developed from this group, the cultivatable marine caliciviruses, should provide insights as a predictive model to help answer questions for the noncultivatable caliciviruses such as small round structured virus, Sapporo virus, hepatitis E virus, and rabbit caliciviruses. From the replicative strategy of the Caliciviridae (as RNA viruses), one would predict considerable diversity. In vitro cultivation has shown that caliciviruses exhibit survivability and plasticity in nature. Many of the factors regarding host spectrum, zoonotic potential, disease conditions, transport, intermediate hosts, and abrupt appearance or disappearance, which may be unknown in newly emerging calicivirus diseases (e.g., RHD), may be more reliably predicted with an established model such as the cultivatable marine caliciviruses. New and better biologic tools for diagnostic and epidemiologic assessments must be developed. This should be augmented by recognizing the zoonotic potential of the cultivatable caliciviruses of ocean origin and then examining them as possible models to help solve many unanswered questions for pathogenic Caliciviridae.
Alvin W. Smith is professor of veterinary virology and head, Laboratory for Calicivirus Studies at the College of Veterinary Medicine, Oregon State University, Corvallis, Oregon. Dr. Smith is interested in mechanisms for the preservation and movement of pathogenic viruses in nature, particularly those contained in ocean reservoirs. His research has focused on the marine caliciviruses.
Acknowledgment
We thank Ms. Christine Robinette for manuscript preparation. This study was funded by the Oregon Agricultural Experiment Station through the College of Veterinary Medicine.
References
- Smith AW, Boyt PM. Caliciviruses of ocean origin: a review. J Zoo Wildl Med. 1990;21:3–23.
- Smith AW, Skilling DE. Comparison of antibody titers for 13 calicivirus serotypes between aborting and full-term adult Zalophus californianus. Special report submitted to the National Marine Mammal Laboratory, National Marine Fisheries Service. Corvallis (OR): Laboratory for Calicivirus Studies, Oregon State University; 1994.
- Gunn RA, Janowski HT, Lieb S, Prather EC, Greenberg HB. Norwalk virus gastroenteritis following raw oyster consumption. Am J Epidemiol. 1982;115:348–51.PubMedGoogle Scholar
- Morse DL, Grabau JJ, Hanrahan R, Stricof M, Shaegani M, Diebal R, Widespread outbreaks of clam- and oyster-associated gastroenteritis: role of Norwalk virus. N Engl J Med. 1986;314:678–81.PubMedGoogle Scholar
- Bradley DW. Hepatitis E virus: a brief review of the biology, molecular virology, and immunology of a novel virus. J Hepatol. 1995;22:140–5.PubMedGoogle Scholar
- Chasey D. Possible origins of rabbit haemorrhagic disease in the United Kingdom. Vet Rec. 1994;135:496–9.PubMedGoogle Scholar
- Smith AW, Berry ES, Skilling DE, Barlough JE, Poet SE, Berke T, In vitro isolation and characterization of a calicivirus causing a vesicular disease of the hands and feet. Clin Infect Dis. 1998. In press.PubMedGoogle Scholar
- Berke T, Golding B, Jiang K, Cubitt DW, Wolfaardt M, Smith AW, A phylogenetic analysis of the caliciviruses. J Med Virol. 1997. In press.PubMedGoogle Scholar
- Barlough JE, Berry ES, Skilling DE, Smith AW. The marine calicivirus story-part I. Compend Contin Educ Pract Vet. 1986;8:F5–14.
- Barlough JE, Berry ES, Skilling DE, Smith AW. The marine calicivirus story-part II. Compend Contin Educ Pract Vet. 1986;8:F75–82.
- Bradley DW. Enterically transmitted non A non B hepatitis. Br Med Bull. 1990;46:442–61.PubMedGoogle Scholar
- Holland J. Replication error, quasispecies populations, and extreme evolution rates of RNA viruses. In: Morse SS, editor. Emerging Viruses. Oxford: Oxford University Press; 1993. p. 203-17.
- Smith AW, Skilling DE, Applegate GL, Trayor TR, Lola TJ, Poet SE. Calicivirus isolation from stillborn piglets in two outbreaks of swine infertility and respiratory syndrome (SIRS). Proceedings of the World Association of Veterinary Microbiologists, Immunologists, and Specialists in Infectious Disease. Davis (CA): University of California, Davis; 1992.
- Smith AW, Madin SH. Vesicular exanthema of swine. In: Leman AD, Straw B, Glock RD, Mengeling WL, Penny RHC, Scholl E, editors. Diseases of swine. 6th edition. Ames (IA): Iowa State Press; 1986. p. 358-68.
- Bankowski RA. Vesicular exanthema. In: Bankowski RA, editor. Advances in veterinary science. Academic Press; 1965. p. 23-64.
- Morse SS. Examining the origins of emerging viruses. In: Morse SS, editor. Emerging viruses. Oxford: Oxford University Press; 1993. p. 10-28.
- Smith AW, Akers TG, Madin SD, Vedros NA. San Miguel sea lion virus isolation, preliminary characterization and relationship to vesicular exanthema of swine virus. Nature. 1973;244:108–10. DOIPubMedGoogle Scholar
- Smith AW, Akers TG. Vesicular exanthema of swine. J Am Vet Med Assoc. 1976;169:700–3.PubMedGoogle Scholar
- Smith AW, Skilling DE, Ensley PK, Benirschke K, Lester TL. Calicivirus isolation and persistence in a pygmy chimpanzee (Pan paniscus). Science. 1983;221:79–81. DOIPubMedGoogle Scholar
- Smith AW, Skilling DE, Prato CM, Bray HL. Calicivirus (SMSV-5) infection in experimentally inoculated opaleye fish (Girella nigricans). Arch Virol. 1981;67:165–8. DOIPubMedGoogle Scholar
- Smith AW, Skilling DE, Barlough JE, Berry ES. Distribution in the North Pacific Ocean, Bering Sea, and Arctic Ocean of animal populations known to carry pathogenic caliciviruses. Dis Aquat Organ. 1986;2:73–80. DOIGoogle Scholar
- Smith AW, Prato CM, Skilling DE. Caliciviruses infecting monkeys and possibly man. Am J Vet Res. 1978;39:287–9.PubMedGoogle Scholar
- Smith AW, Skilling DE, Dardiri AH, Latham AB. Calicivirus pathogenic for swine: a new serotype isolated from opaleye Girella nigricans, an ocean fish. Science. 1980;209:940–1. DOIPubMedGoogle Scholar
- Smith AW, Anderson MP, Skilling DE, Barlough JE, Ensley PK. First isolation of calicivirus from reptiles and amphibians. J Am Vet Med Assoc. 1986;47:1718–21.
- Sawyer JC. Vesicular exanthema of swine and San Miguel sea lion virus. J Am Vet Med Assoc. 1976;169:707–9.PubMedGoogle Scholar
- Smith AW, Reno P, Poet SE, Skilling DE, Stafford C. Retention of ocean-origin caliciviruses in bivalve mollusks maintained under experimental depuration conditions. In: Fenwick B, editor. Proceedings of the 25th Annual Meeting of the International Association of Aquatic Animals; 1994; Vallejo, California. Baltimore: International Association of Aquatic Animal Medicine; 1994.
- Poet SE, Skilling DE, Megyesi JL, Gilmartin WG, Smith AW. Detection of a non-cultivatable calicivirus from the white tern (Gygis alba rothschildi). J Wildl Dis. 1996;32:461–7.PubMedGoogle Scholar
- Evermann JF, Bryan CM, McKiernan AJ. Isolation of a calicivirus from a case of canine glossitis. Canine Pract. 1981;8:36–9.
- Evermann JF, McKiernan AJ, Smith AW, Skilling DE, Ott RL. Isolation of a calicivirus from dogs with enteric infections. Am J Vet Res. 1985;46:218–20.PubMedGoogle Scholar
- Cubitt DW. Proceedings of the European Society of Veterinary Virology. Readings, United Kingdom: Reading University; 1996.
- Bureau of Resource Studies. Rabbit calicivirus disease. Canberra, Australia: Australian Government Printing Office; August 1996. p. 20-56.
- Mead C, Kaldor J, Canton M, Gamer G, Crerar S, Thomas S. Rabbit calicivirus and human health. Canberra, Australia: Department of Primary Industries and Energy, Australian Government (Released under the Official Information Act). Report of the Rabbit Calicivirus Human Health Study Group; 1996.
- Xu WY. Viral haemorrhagic disease of rabbits in the People's Republic of China: epidemiology and virus characterization. Revue Scientifique et Technique. Office International des Epizooties. 1991;10:2393–408.
- Gregg DA, House C, Meyer R, Berninger M. Viral haemorrhagic disease of rabbits in Mexico: epidemiology and viral characterization. Revue Scientifique et Technique. Office International des Epizooties. 1991;10:2435–51.
- Ohlinger VF, Thiel HJ. Identification of the viral haemorrhagic disease in rabbits as a calicivirus. Revue Scientifique et Technique. Office International des Epizooties. 1991;10:2311–23.
Figures
Tables
Cite This ArticleTable of Contents – Volume 4, Number 1—March 1998
| EID Search Options |
|---|
|
|
|
|
|
|