Volume 6, Number 2—April 2000
Synopsis
The bdr Gene Families of the Lyme Disease and Relapsing Fever Spirochetes: Potential Influence on Biology, Pathogenesis, and Evolution
Identification of Borrelia Direct Repear (bdr) Related Genes
Evolutionary Analyses of bdr-Related Sequences: Revised Nomenclature for the Bdr-Related Proteins
Molecular Features and Physical Properties of the Bdr Proteins
Immunologic Analyses of the Bdr Proteins
The Bdr Proteins and Borrelia Biology: An Overview
Cite This Article
Cite This Article
Citation for Media
Abstract
Species of the genus Borrelia cause human and animal infections, including Lyme disease, relapsing fever, and epizootic bovine abortion. The borrelial genome is unique among bacterial genomes in that it is composed of a linear chromosome and a series of linear and circular plasmids. The plasmids exhibit significant genetic redundancy and carry 175 paralogous gene families, most of of unknown function. Homologous alleles on different plasmids could influence the organization and evolution of the Borrelia genome by serving as foci for interplasmid homologous recombination. The plasmid-carried Borrelia direct repeat (bdr) gene family encodes polymorphic, acidic proteins with putative phosphorylation sites and transmembrane domains. These proteins may play regulatory roles in Borrelia. We describe recent progress in the characterization of the Borrelia bdr genes and discuss the possible influence of this gene family on the biology, pathogenesis, and evolution of the Borrelia genome.
Species of the genus Borrelia cause human and animal infections (1). In North America, Lyme disease and endemic relapsing fever pose the greatest threat to human health and have received the most attention of the borrelial diseases. Approximately 14,000 cases of Lyme disease are reported in the United States each year; however, the actual number of cases may be 10-fold higher (2). Lyme disease was not recognized as a distinct clinical entity in North America until the 1970s (3). The causative agent, a previously uncharacterized spirochete transmitted through the bite of infected ticks of the Ixodes ricinus complex (I. scapularis in the Northeast and Midwest and I. pacificus on the West Coast) (4,5), was classified in the genus Borrelia and named B. burgdorferi. With the emergence of Lyme disease and the identification of its etiologic agent, Borrelia research focused on the development of reliable Lyme disease diagnostic assays and vaccines, and the phenotypic and genotypic diversity of Borrelia was thoroughly analyzed. Through modern molecular taxonomic techniques, several newly described species of Borrelia have emerged as possible causative agents of Lyme disease or at least as agents genetically related to B. burgdorferi (6-15). The B. burgdorferi sensu lato complex is composed of the following species: B. turdae, B. tanukii, B. bissettii, B. valaisiana, B. lusitaniae, B. bissettii, B. andersonii, B. japonica, B. garinii, and B. afzelii. Of these, B. burgdorferi, B. garinii, and B. afzelii are the dominant species associated with infection in humans.
Relapsing fever has been studied not only for its impact on human health but also as a model system for antigenic variation. There are two general forms of relapsing fever, epidemic (louse borne--Pediculus humanus) and endemic (tick borne--Ornithodoros spp.) (1). Epidemic relapsing fever tends to be associated with poor living conditions and social disruption (famine and war) and is rare in the United States. Endemic relapsing fever is more prevalent, predominantly in the western regions. Three closely related Borrelia species, B. hermsii, B. turicatae, and B. parkeri, are associated with this disease. Hallmark features of relapsing fever include cyclic fever and spirochetemia. The molecular basis for these features can be attributed to the differential production of dominant variable surface antigens of the Vmp protein families (16). The 40 or so plasmid-carried vmp related genes in the B. hermsii genome are expressed only one at a time. A single expression locus exists, and genes not at this site lack a promoter element and are therefore not transcribed (17). The expressed Vmp becomes primary target of a vigorous humoral immune response that kills most of the spirochetal population. However, at a frequency of approximately of 1 x 10-3 to 1 x 10-4 per generation, the identity of the expressed Vmp changes (18) through gene conversion (19). The net effect of this nonreciprocal event is to replace the gene located in the expression locus with one that was previously silent. The production of a new antigenically distinct Vmp allows evasion of the humoral immune response. This ongoing change in Vmp synthesis allows the relapsing fever spirochete population to reestablish itself in the host, thus leading to spirochetemia and the relapse of fever. Antigenic variation systems have also been identified in the Lyme disease spirochetes; however, they appear to exert a more subtle effect (20).
While clinical relapsing fever and Lyme disease differ from each other in many ways, their causative agents share many similarities at both the biologic and genetic levels. At the biologic level, they are host associated and undergo similar environmental transitions in the course of cycling between mammals and arthropods. In view of the distinctly different characteristics of these environments, the spirochetes must be able to adapt rapidly. Evidence suggests that the relapsing fever and Lyme disease spirochetes use related proteins to adapt to or carry out similar functions in changing environments. For example, homologs of the plasmid-carried ospC gene of the Lyme disease spirochetes are carried by several other Borrelia species, including the relapsing fever spirochetes (21). Both ospC and its relapsing fever spirochete homolog (vmp33) are selectively expressed during the early stages of infection, which suggests that they play a common functional role (22,23). The B. burgdorferi Rep or Bdr protein family is also distributed genuswide. Members of this polymorphic protein family possess highly conserved putative functional motifs and structural properties, which suggests that they may also carry out an important genuswide role (24,25).
At the molecular level, a unique feature of Borrelia is the unusual organization and structure of their genome. Unlike most bacteria, which carry their genetic material in the form of a single, circular DNA molecule, Borrelia have a segmented genome (26-28). Most genetic elements carried by these bacteria are linear with covalently closed termini or telomeres (27). The telomeres are characterized by short hairpin loops of DNA (29). If heat denatured, these linear molecules relax to form a single-stranded circular molecule. If reannealed, they base-pair upon themselves to form a double-stranded linear molecule that by physical necessity possesses a short single-stranded hairpin loop at each telomere. Genetic elements of this structure are rare in bacteria and are reminiscent of certain viral genomes. In B. burgdorferi (isolate B31), the largest of the linear genomic elements is the 911-kb chromosome (30). The chromosome carries 853 putative ORFs, most of which are thought to encode housekeeping functions. The remaining 12 linear and 8 circular genetic elements are plasmids. The plasmids might best be thought of as mini-chromosomes, since as a group they are indispensable in situ and may carry genes encoding proteins involved in housekeeping functions (31). In addition, they may further deviate from the true definition of a plasmid in that their replication may not be independent and may instead be tightly coordinated with the replication of the chromosome (32,33).
Nearly 50% of the plasmid-carried ORFs lack homology with known sequences, which suggests that their encoded proteins may define the unique biologic and pathogenetic aspects of Borrelia (30). Several of the proteins derived from these plasmid-carried genes of unknown function are antigenic or selectively expressed during infection, which indicates that they function in the mammalian environment (20,34-37). A striking feature of the plasmid-carried ORFs is that they are organized into 175 paralogous gene families of two or more members (30). Hence, the DNA content of the plasmids is highly redundant. Since the maintenance of DNA is energetically expensive, it is likely that this redundant DNA is of biologic importance to Borrelia. The paralogous gene families of Borrelia have been the focus of intensive research as they are thought to play important roles in pathogenesis and to influence genome organization and evolution (20,30,35,38-40).
The bdr gene family is a large, polymorphic, plasmid-carried, paralogous gene family of unknown function that was originally identified in B. burgdorferi (41,42). Members of this gene family have been characterized in several Borrelia species and isolates (Table 1) and have been assigned various gene names (25,41-44) (Table 2).We have adopted the bdr designation in the context of a nomenclature system (25), summarized below. Genes belonging to the bdr gene family were first identified through the analysis of repeated DNA sequences in B. burgdorferi sensu lato complex isolates (41,42). Seven nonidentical but closely related copies of a plasmid-carried repeated element were identified in B. burgdorferi 297 (42). Three additional copies of this repeated sequence were further identified in B. burgdorferi 297 (45). These loci carry several ORFs that were designated as rep+, rep-, LPA, LPB (the LP genes have recently been redesignated as mlp for multicopy lipoprotein [45]), rev, and the orfABCD operon (note: ORFs A and B have been redesignated as blyA and blyB). Some of these genes, particularly rep and mlp, exhibit allelic variation and encode polymorphic proteins, the functions of which are under investigation. Focusing specifically on the rep or bdr genes, the rep designation was originally chosen to reflect a central repeat motif carrying domains in the deduced amino acid sequences. The + and - designations were assigned to indicate that the overlapping rep+ and rep- genes are located on opposing DNA strands. Plasmid-carried repeated DNA sequences were also identified in B. burgdorferi B31 and found to carry either all or a subset of seven ORFs, designated A through G (41). Of relevance to this discussion are the ORF-E sequences that are rep or bdr homologs. A bdr-related gene was also identified in B. afzelii DK1 and designated as p21 (43). B. afzelii causes Lyme disease in Europe and Asia. The rep+, ORF-E, and p21 designations have recently been replaced with bdr gene designations (24,25,44).
To assess and compare the composition and complexity of the bdr gene family among species and isolates of the B. burgdorferi sensu lato complex, restriction fragment length polymorphism (RFLP) patterns were determined (Appendix ). Genomic DNA digested with Xba1 was Southern blotted and probed with an oligonucleotide targeting the bdr genes (Figure 1). A variable number of hybridizing bands of different size were detected. These analyses demonstrate that extensive bdr gene families are carried by B. burgdorferi sensu lato complex isolates and that the RFLP patterns vary at the inter- and intraspecies level. Hybridization analyses of other Borrelia species showed that they also carry bdr-related gene families (24,25,46). bdr-related genes have been detected by hybridization in B. turicatae, B. hermsii, B. parkeri, B. coriaceae, and B. anserina (25,46). Isolates of these species also exhibit substantial variation in their bdr RFLP patterns at the intraspecies level. Table 1 lists the Borrelia species that carry bdr-related genes and indicates the methods by which these genes or proteins were detected.
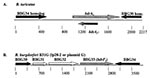
Sequences flanking some bdr alleles also appear to be distributed genus wide. Some bdr alleles of B. turicatae, B. parkeri, and B. hermsii are flanked by genes that are homologs of genes carried by the Lyme disease spirochetes (24,25). As a specific example, the B. turicatae bdrA1 gene is flanked by ORFs that are homologs of the BBG34 and BBG30 genes of B. burgdorferi (24,25). In the Lyme disease spirochetes, BBG34 is part of a three-member paralogous gene family, while BBG30 is a single-copy gene (30). Located between BBG30 and BBG34 is BBG33, a member of the bdr gene family (recently redesignated as bdrF2) (25). Although these divergent Borrelia species carry related genes, their organization differs (24), which indicates that rearrangement has taken place in the ancestral plasmid that carried these homologs. Figure 2 compares the organization of two bdr loci from B. turicatae and B. burgdorferi.
To simplify the complicated nomenclature of bdr-related genes, a bdr nomenclature system has been developed that assigns gene names on the basis of phylogenetic relationships inferred from comparative analysis of genetically stable regions of the bdr genes (25). This system, which is applicable genuswide, allows for a ready assessment of relationships among bdr paralogs and orthologs. The rationale for this system stemmed from the results of a comprehensive evolutionary analysis of >50 bdr-related sequences from five Borrelia species that demonstrated that bdr sequences are organized into six distinct subfamilies, designated A through F (25). Subfamilies are not necessarily species specific; some contain bdr alleles from different Borrelia species (25). Since members of a given subfamily are closely related to one another with identity values for the N terminal domain being >95%, each member is assigned the same gene name designation, and paralogs are distinguished by a numerical subscript. In B. turicatae OZ-1, two bdr subfamilies, bdrA and bdrB, contain at least four and five members, respectively (24). Members of the bdrA subfamily are designated bdrA1, bdrA2, bdrA3, and bdrA4, while members of the bdrB family are designated bdrB1 through bdrB5. This revised Bdr nomenclature scheme was modeled after that proposed for bacterial polysaccharide synthesis genes (47) and is in accordance with the nomenclature guidelines established by Demerec (48).
The subfamily affiliation of bdr genes can be readily determined through comparative sequence analyses of the amino acid segment preceding the polymorphic repeat motif region of these proteins (described in detail below) (25). Relationship assessments based on the genetically stable N terminal domain (vs. complete sequences) are preferable because the calculated evolutionary distances and clustering relationships are not artificially skewed by the variable number of repeat motifs present in the repeat motif domain. Since the genetically unstable repeat motif domain comprises as much as 50% of the total coding sequence in some alleles, it can have a substatial impact on inferred relationships. In addition, extensive sequence variation in the carboxyl termini of the Bdr proteins at the inter-species level makes it difficult to align this domain with confidence, which further influences the inferred relationships.
bdr evolutionary analyses show that Borrelia species carry members of at least two bdr subfamilies (25,44). In fact, B. burgdorferi carries three distinct subfamilies. Multiple Bdr subfamilies in diverse Borrelia species suggest that there has been selective pressure to maintain multiple bdr alleles and bdr genetic diversity. This genetic diversity may increase the functional diversity of the Bdr proteins.
While early analyses of Borrelia bdr genes demonstrated their multicopy nature (41,42,46), the full extent of the complexity of the bdr gene family in the Lyme disease spirochetes was not fully recognized until the B. burgdorferi genome sequence was determined (30). B. burgdorferi B31 was found to carry 17 distinct bdr-related genes (and one truncated variant) distributed among different linear and circular plasmids. B. turicatae, which carries at least nine different bdr alleles, carries these genes exclusively on linear plasmids (24,25,46). Other relapsing fever spirochete species (B. parkeri and B. hermsii) are similar to the Lyme disease bacteria in that they carry bdr genes on both linear and circular plasmids (25). In the Lyme disease spirochetes each of the 32-kb circular plasmids, with the exception of plasmids M and P, carry two different bdr genes separated by seven or eight ORFs. Each of these circular plasmids carries one bdrD subfamily member and one bdrE subfamily member. The maintenance of genes belonging to different subfamilies on a single plasmid is consistent with the possibility that each carries out a different function. In contrast, in the Lyme disease spirochetes, the bdrF subfamily members are localized to linear plasmids with only a single bdr gene per plasmid. These observations suggest that there has been selective pressure to maintain the association of specific subfamilies with specific types of plasmids. Less is known about the bdr-carrying plasmids and the organization of the bdr genes and subfamilies in the relapsing fever borreliae. However, as in the Lyme disease spirochetes, in B. turicatae most bdr-carrying plasmids carry two bdr genes, one from subfamily bdrA and one from subfamily bdrB (24).
The sequence of more than 50 bdr alleles from five different Borrelia species has been determined (Table 2) (24,25,41-43,46). These extensive comparative sequence analyses led to the identification of conserved features that provide insight into the possible biologic roles of the Bdr proteins. For example, all bdr alleles carry centrally located repeat motif domains (Figure 3). Although conserved in sequence, these domains vary in length among alleles as a result of varying numbers of the repeat motif. The core tripeptide of the repeat is the sequence KID. The repeat motifs encode consensus casein kinase 2 phosphorylation (CK2P) motifs of the sequence T/SKID/E (43). While it may appear somewhat paradoxical for bacteria to carry casein kinases, casein kinase is a descriptive term broadly applied to at least two classes of ubiquitous protein kinases for which the substrates may include various enzymes and noncatalytic proteins involved in important cellular regulatory functions (49). Most proteins phosphorylated by CK2-like kinases are highly acidic, as are the Borrelia Bdr proteins (isoelectric points between 5 and 6). The phosphorylation site in CK2P motifs is either the Ser or Thr residue of the motif. Although histidine kinases have been known to exist in some bacteria, it has been widely held that bacteria lack Ser - Thr kinases. However, Ser -Thr kinases have recently been identified in several bacterial species, including Myxococcus, Anabeana, Freymella, Yersinia, and Streptomyces (50). Most importantly, analysis of the B. burgdorferi genome sequence identified a putative Ser - Thr kinase designated BB0648 (30,50). This ORF carries a domain that exhibits homology with the active site of Ser - Thr kinases. B. burgdorferi also carries a homolog of the PPM family of eucaryotic protein Ser - Thr phosphatases (30,50). The presence of these genes in B. burgdorferi suggests that the Borrelia possess the machinery necessary for Ser - Thr phosphorylation and dephosphorylation.
Another important conserved feature identified through sequence analyses is the hydrophobic carboxyl terminal domain of approximately 20 amino acids. Computer analyses conducted with the TMpred program indicate that this domain has a high propensity to form a transmembrane helix (24,25). The Tmpred values for the 20 aa C-terminal domains are 2,000 to 2,600. A value of 500 or greater is considered significant (24,25). Comparison of the Bdr putative transmembrane domain sequences from the Lyme disease spirochetes with those from the relapsing fever spirochetes indicates that, while there is conservation in physical properties, there is essentially no conservation of primary sequence. However, sequence conservation does exist at the subfamily level (24,25). Since the Bdr proteins lack an obvious export signal, membrane association would most likely be with the spirochetal inner membrane, with the rest of the protein, which is hydrophilic, extending into the cytoplasm. The terminal residue of the protein is in almost all cases a positively charged amino acid (lysine or asparagine). This residue could extend into the periplasm and serve to anchor the Bdr proteins to other cellular components, such as the peptidoglycan.
The presence of multiple bdr alleles and bdr subfamilies within isogeneic populations has prompted speculation that there may be differential expression at either the subfamily or individual allele level, possibly in response to environmental stimuli (46). Limited studies of bdr expression and production, based on either mRNA detection or immunoblot analyses, have been performed. Porcella et al. (42) used Northern hybridization to determine if expression of B. burgdorferi bdr-related genes occurs during cultivation in the laboratory under standard culture conditions (33°C in BSK media). Bdr transcripts were not detected by this approach. Similarly, in an earlier analysis, we also conducted Northern hybridization experiments to assess bdr expression (46). We detected expression of B. turicatae OZ1 bdrA subfamily members in bacteria cultivated under standard laboratory growth conditions (46). However, when reverse transcriptase (RT)-PCR methods were applied, transcription of a single bdrA allele was detected (46). B. turicatae OZ-1 was later demonstrated to carry at least nine bdr alleles, four of which belong to the bdrA subfamily. Analysis of the sequence of these alleles showed that all four should have been readily amplified by the RT-PCR primer set because of the conservation of the primer binding sites (24). The lack of detection of transcript derived from these alleles suggested that only a subset of the bdr A subfamily alleles is expressed. This raised the possibility that other bdr alleles are either nonfunctional genes or their expression requires different environmental stimuli. The transcriptional expression of the bdrB subfamily has not been specifically assessed. Thorough transcriptional analyses using allele-specific probes and primers are an important step, since they allow specific assessment of the expression of individual bdr alleles under differing environmental conditions. In addition, analyses of the upstream DNA sequences of individual bdr alleles and their genomic location may elucidate the molecular basis for bdr transcriptional regulation.
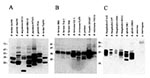
Immunologic analyses have provided a somewhat different overall picture regarding Bdr production. Immunologic analyses described in this report and elsewhere (44) demonstrate that several members of the bdr gene family are expressed during in vitro cultivation. We conducted a comprehensive analysis of the expression of Bdr proteins among Borrelia species. When antisera raised against recombinant B. afzelii BdrF1 (24) were used in immunoblot analyses, several immunoreactive proteins were detected in cell lysates of all Borrelia species tested (Figure 4). The only exception was B. anserina, a causative agent of avian spirochetosis. Although bdr-related sequences have been detected in B. anserina by hybridization techniques (46), immunoreactive proteins were not detected in immunoblot analyses. Additional analyses are required to determine if this indicates absence of translational expression or the lack of epitope conservation in this species. In any event, the fact that immunoreactive bands were not detected in this species attests to the specificity of the anti-Bdr antisera. As a further demonstration of the specificity of the antisera and to highlight the fact that the Bdr proteins are unique to Borrelia, a cell lysate of Leptospira interrogans was included in the immunoblot analyses. Immunoreactivity with proteins in the Bdr size range was not observed with the anti-Bdr antisera in this spirochete species. Borrelia species that expressed immunoreactive proteins included B. garinii, B. burgdorferi, B. turdae, B. tanukii, B. japonica, B. valaisiana, B. afzelii, B. coriaceae, B. bissettii, B. miyamotoi, B. parkeri, B. hermsii, and B. turicatae (Table 1). Particularly striking was the extensive variation in the number and molecular weight of the immunoreactive proteins expressed, with up to 12 distinct Bdr proteins detected. Variation in expression patterns was observed at both the inter- and intraspecies level. Analysis of three B. burgdorferi isolates (B31G, cN40, and CA12) demonstrated variability in both the size and number of expressed Bdr proteins. Isolate B31G has been demonstrated by genomic sequencing to carry 18 distinct bdr alleles. Immunoblot analyses show that not all alleles are expressed during in vitro cultivation; therefore, some alleles may be differentially regulated.
The broad immunoreactivity of the antisera with diverse Borrelia species indicates that some epitopes are conserved genuswide. In view of the sequence divergence in the N and C terminal domains of the Bdr proteins derived from different subfamilies, it is likely that the cross-reactive epitopes reside in the conserved repeat motif region. Consistent with this, computer analyses of the repeat domain of all determined Bdr protein sequences predict them to be alpha helical and to have a surface exposed on the protein and a positive Jameson-Wolf antigenic index (24,25,44). The conservation and synthesis of these polymorphic proteins in such a diverse group of Borrelia species suggest that they play an important role in Borrelia biology genuswide.
Bdr genes and extensive bdr gene families have now been identified and characterized in several diverse Borrelia species (24,25,42-44,46). Comparative sequence analyses, which have identified conserved putative functional domains, have provided the basis for the development of hypotheses regarding Bdr function and cellular location. The Bdr proteins, which lack known consensus export signals, are likely anchored to the cytoplasmic membrane through their conserved, hydrophobic, putative transmembrane spanning domain. The C-terminal positively charged amino acid may be exposed to the periplasm, where it may interact with other cellular components that may include the peptidoglycan. The repeat motif domain, which is predicted by computer analyses to be hydrophilic and surface exposed on the protein, likely extends into the cytoplasm. The conserved repeat motif domain that carries the putative Ser - Thr phosphorylation motifs may then be accessible for phosphorylation or to interact with other cytoplasmic proteins or DNA to form a membrane anchored complex. As with numerous other proteins, phosphorylation and dephosphorylation could play a regulatory role, perhaps in signaling or sensing.
Multiple polymorphic bdr alleles may increase the functional range and diversity of the Bdr proteins. Functional partitioning among Bdr proteins could offer a possible explanation of why Borrelia expend such biologic energy to maintain these genes in large gene families and express variants of these proteins. The homology among bdr alleles may also allow or lead to the continual modification of these genes through homologous recombination. In fact, the variable nature of the repeat motif region, which is clearly not evolutionarily stable, has likely arisen from slipped-strand mispairing, recombination, or rearrangement. In view of the extensive genetic redundancy of the plasmid component of the Borrelia genome, recombination in and among related sequences on different plasmids could affect the organization and evolution of the genome and ultimately host-pathogen interaction. Inter- or intra-plasmid exchange of DNA sequences could provide a mechanistic basis for the extensive genetic variability that has been widely described for Borrelia plasmids (28,29,51-59). In spite of the apparent necessity for at least most of the plasmids for survival, as inferred from their ubiquitous distribution among Borrelia isolates, these bacteria are able to tolerate remarkable genomic variability. Diversity in the plasmids and the genes they carry may actually be exploited as a tool for phenotypic diversity and rapid environmental adaptation.
Mr. Roberts is a 3rd-year Ph.D. student, Department of Microbiology and Immunology, Medical College of Virginia at Virginia Commonwealth University. His scientific interests are centered on the study of the molecular mechanisms of pathogenesis and adaptive responses of arthropod-carried spirochetes, specifically on the role of plasmid-carried gene families in Borrelia biology and pathogenesis.
Acknowledgments
We thank Michael Norgard, Steve Porcella, Justin Radolf, and the Molecular Pathogenesis research group at Virginia Commonwealth University for helpful discussions.
This work was supported in part by a grant from the Jeffress Memorial Trust.
References
- Barbour AG, Hayes SF. Biology of Borrelia species. Microbiol Rev. 1986;50:381–400.PubMedGoogle Scholar
- Yound JD. Underreporting of Lyme disease. N Engl J Med. 1998;:338.
- Steere AC, Malawista SE, Snydman DR, Shope RE, Andiman WA, Ross MR, Lyme arthritis: an epidemic of oligoarticular arthritis in children and adults in three Connecticut communities. Arthritis Rheum. 1977;20:7–17. DOIPubMedGoogle Scholar
- Burgdorfer W, Barbour AG, Hayes SF, Benach JL, Grunwaldt E, Davis JP. Lyme disease--a tick-borne spirochetosis? Science. 1982;216:1317–9. DOIPubMedGoogle Scholar
- Benach JL, Bosler EM, Hanrahan JP, Coleman JL, Bast TF, Habicht GS, Spirochetes isolated from the blood of two patients with Lyme disease. N Engl J Med. 1983;308:740–2.PubMedGoogle Scholar
- Baranton G, Postic D, Saint Girons I, Boerlin P, Piffaretti J-C, Assous M, Delineation of Borrelia burgdorferi sensu stricto, Borrelia garinii sp. nov., and group VS461 associated with Lyme borreliosis. Int J Syst Bacteriol. 1992;42:378–83. DOIPubMedGoogle Scholar
- Marconi RT, Garon CF. Identification of a third genomic group of Borrelia burgdorferi through signature nucleotide analysis and 16S rRNA sequence determination. J Gen Microbiol. 1992;138:533–6.PubMedGoogle Scholar
- Marconi RT, Garon CF. Phylogenetic analysis of the genus Borrelia: a comparison of North American and European isolates of B. burgdorferi. J Bacteriol. 1992;174:241–4.PubMedGoogle Scholar
- Marconi RT, Liveris D, Schwartz I. Identification of novel insertion elements, restriction fragment length polymorphism patterns, and discontinuous 23S rRNA in Lyme Disease spirochetes: phylogenetic analyses of rRNA genes and their intergenic spacers in Borrelia japonica sp. nov. and genomic group 21038 (Borrelia andersonii sp. nov.) isolates. J Clin Microbiol. 1995;33:2427–34.PubMedGoogle Scholar
- Fukunaga M, Hamase A, Okada K, Inoue H, Tsuruta Y, Miyamoto K, Characterization of spirochetes isolated from ticks (Ixodes tanukin, I. turdus and Ixodes columnae) and comparision of the sequences with those of Borrelia burgdorferi sensu lato strains. Appl Environ Microbiol. 1996;62:2338–44.PubMedGoogle Scholar
- Fukunaga M, Hamase A, Okada K, Nakao M. sp. nov. and Borrelia turdae sp. nov. found from Ixodid ticks in Japan: rapid species identification by 16S rRNA gene--targeted PCR analysis. Microbiol Immunol. 1996;40:877–81.Borrelia tanukiPubMedGoogle Scholar
- Fukunaga M, Takahashi Y, Tsuruta Y, Matsushita O, Ralph D, McClelland M, Genetic and phenotypic analysis of Borrelia miyamotoi sp. nov., isolates from the Ixodid tick Ixodes persulcatus, the vector for Lyme disease in Japan. Int J Syst Bacteriol. 1995;45:804–10. DOIPubMedGoogle Scholar
- Wang G, van Dam AP, Le Flecha A, Postic D, Peter O, Baranton G, Genetic and phenotypic analysis of Borrelia valaisiana sp. nov. (Borrelia genomic groups VS116 and M19). Int J Syst Bacteriol. 1997;47:926–32. DOIPubMedGoogle Scholar
- Postic D, Belfazia J, Isogai E, Saint Girons I, Grimont PAD, Baranton G. A new genomic species in Borrelia burgdorferi sensu lato isolated from Japanese ticks. Res Microbiol. 1993;144:467–73. DOIPubMedGoogle Scholar
- Postic D, Edlinger C, Richaud C, Grimont F, Dufresne Y, Perolat P, Two genomic species in Borrelia burgdorferi. Res Microbiol. 1990;141:465–75. DOIPubMedGoogle Scholar
- Barbour AG. Antigenic variation of a relapsing fever Borrelia species. Annu Rev Microbiol. 1990;44:155–71. DOIPubMedGoogle Scholar
- Barbour AG, Byrman N, Carter CJ, Kitten T, Berstrom S. Variable antigen genes of the replasing fever agent Borrelia hermsii are activated by promoter addition. Mol Microbiol. 1991;5:489–93. DOIPubMedGoogle Scholar
- Stoenner HG, Dodd T, Larsen C. Antigenic variation of B. hermsii. J Exp Med. 1982;156:1297–311. DOIPubMedGoogle Scholar
- Plasterk RHA, Simon MI, Barbour AG. Transposition of structural genes to an expression sequence on a linear plasmid causes antigenic variation in the bacterium B. hermsii. Nature. 1985;318:257–63. DOIPubMedGoogle Scholar
- Zhang J-R, Hardham JM, Barbour AG, Norris AG. Antigenic variation in Lyme disease Borreliae by promiscuous recombination of vmp like sequence cassettes. Cell. 1997;89:275–85. DOIPubMedGoogle Scholar
- Marconi RT, Samuels DS, Schwan TG, Garon CF. Identification of a protein in several Borrelia species which is related to OspC of the Lyme disease spirochetes. J Clin Microbiol. 1993;31:2577–83.PubMedGoogle Scholar
- Schwan TG, Piesman J, Golde WT, Dolan MC, Rosa PA. Induction of an outer surface protein on Borrelia burgdorferi during tick feeding. Proc Natl Acad Sci U S A. 1995;92:2909–13. DOIPubMedGoogle Scholar
- Schwan TG, Hinnebusch BJ. Bloodstream- versus tick- associated variants of a Relapsing fever bacterium. Science. 1998;280:1938–40. DOIPubMedGoogle Scholar
- Carlyon JA, Roberts DM, Theisen M, Marconi RT. Molecular analyses of the B. turicatae bdr genes: a polymorphic, linear plasmid carried, paralogous gene family. In press 1999.
- Carlyon JA, Roberts DM, Marconi RT. Evolutionary and molecular analyses of the Borrelia rep super gene family: delineation of six distinct sub-families and demonstration of the genus wide conservation of putative functional domains, structural properties and repeat motifs. Microb Pathog. 2000. In press.PubMedGoogle Scholar
- Ferdows MS, Barbour AG. Megabase-sized linear DNA in the bacterium Borrelia burgdorferi, the Lyme disease agent. Proc Natl Acad Sci U S A. 1989;86:5969–73. DOIPubMedGoogle Scholar
- Barbour AG, Garon CF. Linear plasmids of the bacterium Borrelia burgdorferi have covalently closed ends. Science. 1987;237:409–11. DOIPubMedGoogle Scholar
- Barbour AG. Plasmid analysis of Borrelia burgdorferi, the Lyme disease agent. J Clin Microbiol. 1988;26:475–8.PubMedGoogle Scholar
- Hinnebusch J, Barbour AG. Linear plasmids of Borrelia burgdorferi have a telomeric structure and sequence similar to those of a eukaryotic virus. J Bacteriol. 1991;173:7233–9.PubMedGoogle Scholar
- Fraser C, Casjens S, Huang WM, Sutton GG, Clayton R, Lathigra R, Genomic sequence of a Lyme disease spirochaete, Borrelia burgdorferi. Nature. 1997;390:580–6. DOIPubMedGoogle Scholar
- Barbour AG. Linear DNA of Borrelia species and antigenic variation. Trends Microbiol. 1993;:1.PubMedGoogle Scholar
- Hinnebusch J, Tilly K. Linear plasmids and chromosomes in bacteria. Mol Microbiol. 1993;10:917–22. DOIPubMedGoogle Scholar
- Hinnebusch J, Barbour AG. Linear- and circular-plasmid copy numbers in Borrelia burgdorferi. J Bacteriol. 1992;174:5251–7.PubMedGoogle Scholar
- Champion CI, Blanco DR, Skare JT, Haake DA, Giladi M, Foley D, A 9.0 kilobase-pair circular plasmid of Borrelia burgdorferi encodes an exported protein: evidence for expression only during infection. Infect Immun. 1994;62:2653–61.PubMedGoogle Scholar
- Akins D, Porcella SF, Popova TG, Shevchenko D, Baker SI, Li M, Evidence for in vivo but not in vitro expression of a Borrelia burgdorferi outer surface protein F (OspF) homologue. Mol Microbiol. 1995;18:507–20. DOIPubMedGoogle Scholar
- Suk K, Das S, Sun W, Jwang B, Barthold SW, Flavell RA, Borrelia burgdorferi genes selectively expressed in the infected host. Proc Natl Acad Sci U S A. 1995;92:4269–73. DOIPubMedGoogle Scholar
- Skare JT, Foley DM, Hernandez SR, Moore DC, Blanco DR, Miller JN, Cloning and molecular characterization of plasmid-encoded antigens of Borrelia burgdorferi. Infect Immun. 1999;67:4407–17.PubMedGoogle Scholar
- Marconi RT, Sung SY, Hughes CN, Carlyon JA. Molecular and evolutionary analyses of a variable series of genes in Borrelia burgdorferi that are related to ospE and ospF, constitute a gene family, and share a common upstream homology box. J Bacteriol. 1996;178:5615–26.PubMedGoogle Scholar
- Sung S-Y, LaVoie C, Carlyon JA, Marconi RT. Evolutionary instability of ospE related members of the UHB gene family in Borrelia burgdorferi sensu lato complex isolates. Infect Immun. 1998;66:4656–68.PubMedGoogle Scholar
- Zhang JR, Norris SJ. Genetic variation of the Borrelia burgdorferi gene vlsE involves cassette-specific, segmental gene conversion. Infect Immun. 1998;66:3698–704.PubMedGoogle Scholar
- Zuckert WR, Meyer J. Circular and linear plasmids of Lyme Disease spirochetes have extensive homology: characterization of a repeated DNA element. J Bacteriol. 1996;178:2287–98.PubMedGoogle Scholar
- Porcella SF, Popova TG, Akins DR, Li M, Radolf JR, Norgard MV. Borrelia burgdorferi supercoiled plasmids encode multicopy open reading frames and a lipoprotein gene family. J Bacteriol. 1996;178:3293–307.PubMedGoogle Scholar
- Theisen M. Molecular cloning and characterization of nlpH, encoding a novel surface exposed, polymorphic, plasmid-encoded 33kD lipoprotein of Borrelia afzelii. J Bacteriol. 1996;178:6435–42.PubMedGoogle Scholar
- Zuckert WR, Meyer J, Barbour AG. Comparative analysis and immunological characterization of the Borrelia Bdr protein family. Infect Immun. 1999;67:3257–66.PubMedGoogle Scholar
- Yang X, Popova TG, Hagman KE, Wikel SK, Schoeler GB, Caimano MJ, Identification, characterization, and expression of three new members of the Borrelia burgdorferi Mlp (2.9) lipoprotein gene family. Infect Immun. 1999;67:6008–18.PubMedGoogle Scholar
- Carlyon JA, Marconi RT. Cloning and molecular characterization of a multi-copy, linear plasmid-carried, repeat motif-containing gene from Borrelia turicatae, a causative agent of relapsing fever. J Bacteriol. 1998;180:4974–81.PubMedGoogle Scholar
- Reeves PR, Hobbs M, Valvano MA, Kido N, Klena J, Maskell D, Bacterial polysaccharide synthesis and gene nomenclature. Trends Microbiol. 1996;4:498–503. DOIGoogle Scholar
- Demerec M. A proposal for a uniform nomenclature in bacterial genetics. Genetics. 1966;54:61–76.PubMedGoogle Scholar
- Pinna LA. Casein kinase 2: an "eminence grise" in cellular regulation. Biochim Biophys Acta. 1990;1054:267–84. DOIPubMedGoogle Scholar
- Shi L, Potts M, Kennelly PJ. The serine, threonine and/or tyrosine protein kinases and protein phosphatases of procaryotic organisms: a family portrait. FEMS Microbiol Rev. 1998;22:229–53. DOIPubMedGoogle Scholar
- Xu Y, Kodner C, Coleman L, Johnson RC. Correlation of plasmids with infectivity of Borrelia burgdorferi sensu stricto type strain B31. Infect Immun. 1996;64:3870–6.PubMedGoogle Scholar
- Xu Y, Johnson RC. Analysis and comparision of plasmid profiles of Borrelia burgdorferi sensu lato strains. J Clin Microbiol. 1995;33:2679–85.PubMedGoogle Scholar
- Stålhammar-Carlemalm M, Jenny E, Gern L, Aeschlimann A, Meyer J. Plasmid analysis and restriction fragment length polymorphisms of chromosomal DNA allow a distinction between Borrelia burgdorferi strains. Zentralbl Bakteriol. 1990;274:28–39.PubMedGoogle Scholar
- Simpson WJ, Garon CF, Schwan TG. Analysis of supercoiled circular plasmids in infectious and non-infectious Borrelia burgdorferi. Microb Pathog. 1990;8:109–18. DOIPubMedGoogle Scholar
- Schwan TG, Schrumpf ME, Karstens RH, Clover JR, Wong J, Daugherty M, Distribution and molecular analysis of Lyme disease spirochetes, Borrelia burgdorferi, isolated from ticks throughout California. J Clin Microbiol. 1993;31:3096–108.PubMedGoogle Scholar
- Marconi RT, Casjens S, Munderloh UG, Samuels DS. Analysis of linear plasmid dimers in Borrelia burgdorferi sensu lato isolates: implications concerning the potential mechanism of linear plasmid replication. J Bacteriol. 1996;178:3357–61.PubMedGoogle Scholar
- Marconi RT, Samuels DS, Landry RK, Garon CF. Analysis of the distribution and molecular heterogeneity of the ospD gene among the Lyme disease spirochetes: evidence for lateral gene exchange. J Bacteriol. 1994;176:4572–82.PubMedGoogle Scholar
- Ferdows MS, Serwer P, Griess GA, Norris SJ, Barbour AG. Conversion of a linear to a circular plasmid in the relapsing fever agent Borrelia hermsii. J Bacteriol. 1996;178:793–800.PubMedGoogle Scholar
- Carlyon JA, LaVoie C, Sung SY, Marconi RT. Analysis of the organization of multi-copy linear and circular plasmid carried open reading frames in Borrelia burgdorferi sensu lato isolates. Infect Immun. 1998;66:1149–58.PubMedGoogle Scholar
Figures
Tables
Cite This ArticleTable of Contents – Volume 6, Number 2—April 2000
| EID Search Options |
|---|
|
|
|
|
|
|


Please use the form below to submit correspondence to the authors or contact them at the following address:
Richard T. Marconi, Department of Microbiology and Immunology, Medical College of Virginia at VCU, Richmond, VA 23298-0678, USA; fax: 804-828-9946
Top