Volume 10, Number 4—April 2004
Research
Pneumocystis jiroveci Dihydropteroate Synthase Genotypes in Immunocompetent Infants and Immunosuppressed Adults, Amiens, France
Cite This Article
Citation for Media
Abstract
To date, investigations of Pneumocystis jiroveci circulation in the human reservoir through the dihydropteroate synthase (DHPS) locus analysis have only been conducted by examining P. jirovecii isolates from immunosuppressed patients with Pneumocystis pneumonia (PCP). Our study identifies P. jirovecii genotypes at this locus in 33 immunocompetent infants colonized with P. jirovecii contemporaneously with a bronchiolitis episode and in 13 adults with PCP; both groups of patients were monitored in Amiens, France. The results have pointed out identical features of P. jirovecii DHPS genotypes in the two groups, suggesting that in these two groups, transmission cycles of P. jirovecii infections are linked. If these two groups represent sentinel populations for P. jirovecii infections, our results suggest that all persons parasitized by P. jirovecii, whatever their risk factor for infection and the form of parasitism they have, act as interwoven circulation networks of P. jirovecii.
Dihydropteroate synthase (DHPS) is the enzymatic target of sulfonamides, which are the major drugs for Pneumocystis pneumonia (PCP) prophylaxis or treatment (1). Pneumocystis jiroveci (human-specific Pneumocystis) organisms with nonsynonymous mutations at nucleotide positions 165 and 171 on the DHPS locus have been detected in HIV-infected patients with PCP who had previously been treated with sulfonamides (2–14). Prior exposure to sulfonamide drugs has been identified as a predictor of mutant genotypes (2–12). In addition, the city of patient residence has also been identified as an independent risk factor (6,8), a factor that supports the hypothesis that P. jirovecii is transmitted from infected treated patients to susceptible untreated patients, either directly or through a common environmental source. The analysis of P. jirovecii DHPS locus may thus serve as a useful circulation marker of the microorganism in the human reservoir.
To date, investigations of P. jirovecii circulation through the DHPS locus analysis have only been performed by examining isolates from immunosuppressed adults with PCP (8,15,16). However, Pneumocystis infections may cover a wide spectrum of clinical signs and symptoms; colonization with Pneumocystis in immunocompetent infants at risk for primary infection may constitute a large part of this spectrum (17,18). Genotyping of P. jirovecii at the internal transcribed spacer (ITS) locus showed that these infants were infected with similar genotypes as those previously reported in compromised hosts with PCP; this similarity is compatible with the hypothesis that both groups of patients make up a common human reservoir for the fungus (19).
The existence of similar genomic characteristics at another locus, in particular at the DHPS locus, among P. jirovecii isolates from these two groups would provide additional arguments in favor of the fungus’ circulating within a reservoir made up of persons with different clinical forms of P. jirovecii infection. For these reasons, we retrospectively investigated for DHPS genotyping archival P. jiroveici isolates from immunocompetent infants colonized with P. jirovecii and from immunocompromised adults with PCP. Both groups of patients lived in the same French city. The results of this study were reported in part in a conference report (20).
A total of 58 archival P. jirovecii isolates obtained from 58 patients (45 infants and 13 adults) were retrospectively analyzed for DHPS genotyping. All of these patients were monitored in the same University Hospital in Amiens, France.
Forty-five archival nasopharyngeal aspirates (NPA) obtained from 45 nonpremature, immunocompetent infants (median age 4.3 months [range 1.9–11.8]; sex ratio 26 boys and 19 girls) were examined. The 45 infants were hospitalized sometime in the period from November 1999 to April 2001. The specimens initially tested positive for P. jirovecii by a polymerase chain reaction (PCR) assay that amplifies a portion of the gene encoding the mitochondrial large sub-unit rRNA (mtLSUrRNA) (17). All infants initially had an acute respiratory syndrome compatible with a diagnosis of bronchiolitis and no patent immunodeficiency. The presence of P. jirovecii in these infants was considered to reflect merely a colonization. Indeed, clinical improvement was obtained with short-term hospitalization (1–12 days), despite the absence of specific treatment for the fungus. Furthermore, P. jirovecii was associated with the respiratory syncytial virus or with bacteria (Moraxella catarrhalis, Haemophilus influenzae, Streptococcus pneumoniae, Bordetella pertussis) in 35 of 45 infants. None had a past history of sulfonamide treatment. The infants’ characteristics are summarized in Table 1.
Thirteen archival bronchoalveolar lavage (BAL) specimens obtained from 13 immunosuppressed adults in whom PCP was diagnosed were also examined. The 13 patients were hospitalized at some point during the period from June 1996 to November 2001. The specimens initially tested positive for P. jiroveici by microscopy examination that used methanol-Giemsa stain and an immunofluorescence assay (MonofluoKit Pneumocystis; Bio-RAD, Marnes la Coquette, France), and by the PCR at mtLSUrRNA. The underlying conditions were HIV infection (nine patients), renal transplantation (two patients), and long-term corticosteroid treatment for systemic lupus erythematosus (one patient) and for hepatic granulomatosis (one patient). None of the patients had P. jirovecii prophylaxis with sulfonamide drugs in the 3 months preceding the BAL retrieval. The patients’ characteristics are summarized in Table 1. DNAs extracted from NPA and BAL were stored at –20°C until they were typed.
The P. jirovecii DHPS locus was analyzed by PCR–restriction fragment length polymorphism (RFLP). The DHPS sequence was first amplified by a nested PCR assay. The two rounds of PCR were performed under the same conditions (21). Each reaction mixture contained the following reagents at the indicated final concentrations: 10 mM Tris-HCl (pH 8.8), 0.1% Tween 20 (vol/vol), 2.5 mM MgCl2, 200 μM each deoxynucleoside triphosphate (dNTP set, Eurogentec, Seraing, Belgium), 0.6 μM each oligonucleotide primer, and 0.02 U DNA polymerase (Red Goldstar DNA polymerase, Eurogentec)/μL. The first PCR round was conducted with primer pair AHUM (5′- GCG CCT ACA CAT ATT ATG GCC ATT TTA AAT C-3′) and BHUM (5′- CAT AAA CAT CAT GAA CCC G -3′) (14) by using a “touch-down” program. In the first cycle, the denaturation step was 92°C for 30 s, the annealing step was 52°C for 1 min, and the extension step was 72°C for 1 min. This cycle was repeated 10 times but with each annealing step at 1°C lower temperature than the preceding cycle. Subsequently, the last cycle, with an annealing at 42°C, was repeated 20 times. The second PCR round was performed with primer pair CPRIM (5′- CCC CCA CTT ATA TCA-3′) and DPRIM (5′- GGG GGT GTT CAT TCA -3′) (21), for 30 cycles consisting of denaturation at 94°C for 30 s, annealing at 50°C for 1 min, and extension at 72°C for 1 min. The PCR products from the first and the second rounds underwent electrophoresis on a 1.5% agarose gel containing ethidium bromide to visualize the expected bands of 766 bp and 269 bp, respectively. To avoid contamination, each step (reagent preparation, extraction, and amplification) was performed in different rooms with different sets of micropipettes and using barrier tips. PCR mixtures and the extraction step were prepared in a laminar-flow cabinet. Rooms required for amplified DNA manipulation were continuously submitted to an airflow with UV decontamination (SPRW 30 GR4; Paragerm, Inc., Paris, France).
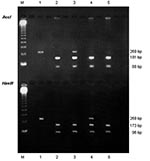
To monitor for possible contamination, a negative control (ultrapure distilled water) was included in each PCR step. The RFLP assay was performed with two restriction enzymes, according to the manufacturer’s recommendations (Promega Corporation, Madison, WI). One part of the nested PCR products was digested with the restriction enzyme AccI, and another part with HaeIII, which make possible the detection of mutations at nucleotide positions 165 and 171, respectively (5) The restriction profiles were visualized by electrophoresis of each digested product on a 1.5% agarose gel with ethidium bromide, as described in Figure 1. The mutations inhibit the restriction enzyme activity. Thus, a wild genotype was shown, after digestion with AccI, by two fragments of 181 bp and 88 bp, and after digestion with HaeIII, by two other fragments of 173 bp and 96 bp. A mutant genotype that has a mutation at nucleotide position 165 (change from A to G, corresponding to a change from Thr to Ala at aminoacid position 55) was shown, after digestion with AccI, by only one uncut fragment of 269 bp, and after digestion with HaeIII, by the two fragments of 173 bp and 96 bp. A mutant genotype that has a mutation at nucleotide position 171 (change from C to T, corresponding to a change from Pro to Ser at aminoacid position 57) was shown after digestion with AccI, by the two fragments of 181 bp and 88 bp, and after digestion with HaeIII, by only one uncut fragment of 269 bp. A double mutant genotype, which has mutations at nucleotide positions 165 and 171, was shown, after digestion with either AccI or HaeIII, by an uncut fragment of 269 bp.
The amplification of P. jirovecii DNA by using the DHPS-based PCR assay was positive for 33 of 45 NPA from infants that initially tested positive with the PCR directed at the mtLSUrRNA gene, whereas it was successful for 13 of 13 BAL specimens from adults. In each positive specimen with the DHPS-based PCR assay, the RFLP technique led to identification of a wild P. jirovecii DHPS genotype. However, mixed infections were diagnosed in three infants and one adult. Indeed, in three (9%) of the 33 NPA from infants (E123, E164, and E181)], the wild genotype was associated with a mutant genotype. In infant E164, the mutant genotype had a mutation at nucleotide position 165, whereas in infants E123 and E181, it had a mutation at nucleotide position 171. In one (8%) of the 13 BAL from adults (N61), the wild genotype was also associated with a mutant genotype, which had a mutation at nucleotide position 171. No infants or adults were infected with a mutant genotype either singly or with a double mutant genotype. The results are detailed in Table 2.
Most studies on P. jirovecii DHPS genotyping have focused on the relationship between P. jirovecii DHPS mutants and prior sulfonamide exposure on the one hand, and PCP outcome on the other hand (2–13,22–24). We have used the DHPS locus analysis differently, as a marker for studying the potential circulation of the fungus in the human reservoir, as it was recently used by Beard et al. and Huang et al. (8,15,16). Although a multilocus genotyping was recently reported as an efficient system for P. jirovecii characterization (25), in this study, we only analyzed the DHPS locus because it still remains the sole marker of circulation. We also obtained the first data on the analysis of P. jiroveici DHPS locus in immunocompetent infants at risk for primary Pneumocystis infection.
The first step of this analysis required a PCR assay. The amplification failed to give positive results for 12 of the 45 NPA from infants who initially tested positive for P. jirovecii by using the PCR at mtLSUrRNA. This difference in sensitivity between the two PCR assays can be explained by the fact the mtLSUrRNA gene is present in many copies within each P. jirovecii genome, whereas the folic acid synthesis gene, encoding the DHPS, is thought to be present in only one copy (26). This difference is particularly manifest on specimens collected by noninvasive means, such as NPA, in which the amount of P. jirovecii is usually low. Indeed, NPA essentially recover cells from the upper respiratory tract, whereas the fungus primarily infects the alveolar spaces. Despite these difficulties, the identification of P. jirovecii DHPS genotypes was successful for three fourths of the samples we examined.
Most investigations of mutations on the P. jiroveici DHPS locus have used the direct sequencing of PCR products (3,4,6–8,10,11,14–16,22,23). More recently, a single-strand conformation polymorphism assay has been described as an alternative method for detecting DHPS mutations (12,27). We used the RFLP assay in this study since this method has a lower cost, is less time-consuming (5,28), and is more efficient for detecting mixed infections than direct sequencing (L. Diop Santos, pers. comm.). The use of restriction enzymes AccI and HaeIII for the digestion of the PCR products showed two mutations at nucleotide positions 165 and 171, as described above. The RFLP assay of P. jirovecii DHPS gene was assessed by Helweg-Larsen et al., who have examined 27 BAL specimens containing a mixture of wild and mutant DHPS genotypes, previously determined by direct sequencing (28). For detecting mutations at nucleotide positions 165 and 171 on P. jirovecii DHPS sequence, these researchers found a 100% concordance between DHPS genotypes determined by AccI and HaeIII restriction enzyme cleavage and by sequencing. Thus, the RFLP assay appears to be a reliable method for discriminating wild and mutant DHPS genotypes.
Mutations at nucleotide positions 165 and 171 have been correlated with prior sulfonamide treatment or prophylaxis (2–12). In our study, since none of the infants or adults had this medical history, the presence of P. jirovecii DHPS mutants has to be discussed. Because of the young age and, consequently, the short medical history of the infants, we could easily ensure that none had had prior exposure to sulfonamides. Conversely, this exposure throughout the adults’ lifetimes cannot strictly be ruled out. These difficulties have previously been raised by Huang et al., who have pointed out the need for a standardized definition of exposure to sulfonamides (29). In particular, the period during which sulfonamides have not been used, preceding patient sampling, to define the absence of selective pressure, varies according to the experience of each medical team. At any rate, in our study, no adults were treated with sulfonamides in the 3 months before BAL retrieval. In this group of patients, we detected P. jirovecii DHPS mutants with a frequency of 8%. This finding may reflect a basic level of infections caused by mutants in the absence of direct selective pressure; their presence is related to an incidental acquisition of the microorganism from humans treated with sulfonamides, either directly or through hypothetical environmental stages. In the same way, this hypothesis may explain the presence of DHPS mutants in the infant group.
Airborne transmission of the fungus from host to host has been demonstrated in rodent models (30), and several observations suggest that interindividual transmission occurs in humans (31,32). Moreover, Pneumocystis organisms infecting each mammalian species are host-specific, and the hypothesis of an animal reservoir for P. jirovecii can be excluded (33). Although an exosaprophytic form of the fungus cannot be ruled out, these data point toward the potential for the specific host to serve as its own reservoir and for PCP in humans as an anthroponosis with humans as a reservoir for P. jirovecii.
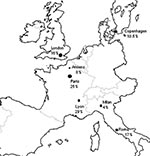
The 8% frequency with which we have detected mutants in PCP patients from Amiens who had no sulfonamide exposure is close to the figure reported for a similar group of patients in Milan, Italy (4% [2]) and in Copenhagen (10.5% [4]), while it appears lower than the rate in Rome (17% [11]), Tokyo (25% [23]), and various U.S. cities (15%–81% [3,6–8,16]). In France, data on DHPS genotypes concern patients living in Paris or Lyon, as recently reported by Latouche et al. and Nahimana et al., respectively (12,13). The frequency of mutants in PCP patients who had no prior sulfonamide treatment or prophylaxis reaches 25% in Paris (13), and 29% in Lyon (12). The low proportion of mutants in Amiens in comparison to Paris and Lyon may be related to different features of P. jirovecii epidemiology in these cities. Amiens (population 150,000) is characterized by a low incidence of AIDS and PCP (34). Conversely, in Paris and its suburbs, a megalopolis of 10 million people, the incidence of these two infections is 30 times as high (34). In Lyon, the second largest city in France, this incidence is 10 times higher than in Amiens (34). Consequently, use of sulfonamides is widespread in Paris and Lyon, favoring the emergence of mutants and provoking a high risk for incidental acquisition of these mutants, even in patients not directly exposed to sulfonamides. This hypothesis is strengthened by a recent report of Miller et al., concerning patients in London, which showed that the decrease of sulfonamide prophylaxis use, related to the introduction of high-active antiretroviral therapy since 1996 conversely generated a reduction of mutant DHPS genotypes in London (36% compared to 17%) (35). Available frequencies of mutants in PCP patients living in Europe who had no prior sulfonamide exposure are shown in Figure 2.
We detected mutants in immunocompetent infants colonized with P. jirovecii and in immunossuppressed adults with PCP with frequencies of 9% and 8%, respectively. Besides these similar frequencies, the most frequent P. jirovecii DHPS genotype was the wild genotype. Mutant genotypes have only been detected within mixed infections. On the whole, genomic characteristics of P. jirovecii organisms at the DHPS locus in the two patient populations living in the same city are similar. In the United States, Beard et al. observed different genomic features at this locus among P. jiroveici isolates from adults and deceased infants (36). For these reasons, those researchers suggested that transmission cycles for P. jirovecii infection in infants and adults were independent. However, whether the two individual groups lived in the same American city was not specified. Conversely, our results of genotyping based on DHPS locus analysis suggest that these transmission cycles are linked, the two patient groups being part of a common reservoir in which the fungus may circulate.
If one considers that both of these patient groups may represent sentinel populations for P. jirovecii infections, other persons infected with P. jirovecii may also be actively involved in the circulation of the fungus. Indeed, new detection tools such as PCR assays have shown that pulmonary colonization with P. jirovecii occurs in patients with diverse levels of immunodeficiency (37) and in immunocompetent patients with lung diseases (38,39). Such assays have also shown that P. jirovecii can transiently parasitize immunocompetent healthcare workers after contacts with PCP patients (40). Our positive results of DHPS genotyping on specimens collected by noninvasive means (NPA) ensure further investigations of P. jirovecii circulation involving such populations, for whom invasive sampling cannot easily be performed. In fact, all parasitized persons, whatever their predisposition to P. jirovecii acquisition and the clinical form of P. jirovecii infection they have, may reflect a wide human reservoir of which all components are not yet characterized. New insights into the P. jirovecii reservoir could provide better prophylactic measures against P. jirovecii transmission and, consequently, PCP.
Dr. Totet is a specialist in medical biology (Department of Parasitology, Mycology, and Travel Medicine, University Hospital of Amiens, France) with an advanced degree in parasitology. This study is a part of the work for her doctoral degree from the University of Picardy.
Acknowledgment
This study was supported by the French Ministry of Education, Research and Technology, “Programme de Recherche Fondamentale en Microbiologie et Maladies Infectieuses et Parasitaires (PRFMMIP),” and the fifth Framework Program of the European Commission (contract number QLK2-CT-2000-01369).
References
- U.S. Public Health Service (USPHS) and Infectious Diseases Society of America. (IDSA). USPHS/IDSA guidelines for the prevention of opportunistic infections in persons infected with human immunodeficiency virus. MMWR Morb Mortal Wkly Rep. 1999;48:1–59.
- Ma L, Kovacs JA, Cargnel A, Valerio A, Fantoni G, Atzori C. Mutations in the dihydropteroate synthase gene of human-derived Pneumocystis carinii isolates from Italy are infrequent but correlate with prior sulfa prophylaxis. J Infect Dis. 2002;185:1530–2. DOIPubMedGoogle Scholar
- Ma L, Borio L, Masur H, Kovacs JA. Pneumocystis carinii dihydropteroate synthase but not dihydrofolate reductase gene mutations correlate with prior trimethoprim-sulfamethoxazole or dapsone use. J Infect Dis. 1999;180:1969–78. DOIPubMedGoogle Scholar
- Helweg-Larsen J, Benfield TL, Eugen-Olsen J, Lundgren JD, Lundgren B. Effects of mutations in Pneumocystis carinii dihydropteroate synthase gene on outcome of AIDS-associated P. carinii pneumonia. Lancet. 1999;354:1347–51. DOIPubMedGoogle Scholar
- Diop Santos L, Lacube P, Latouche S, Kac G, Mayaud C, Marteau M, Contribution of dihydropteroate synthase gene typing for Pneumocystis carinii f.sp. hominis epidemiology. J Eukaryot Microbiol. 1999;46(Suppl):S133–4.
- Kazanjian P, Armstrong W, Hossler PA, Burman W, Richardson J, Lee CH, Pneumocystis carinii mutations are associated with duration of sulfa or sulfone prophylaxis exposure in AIDS patients. J Infect Dis. 2000;182:551–7. DOIPubMedGoogle Scholar
- Kazanjian P, Locke AB, Hossler PA, Lane BR, Bartlett MS, Smith JW, Pneumocystis carinii mutations associated with sulfa and sulfone prophylaxis failures in AIDS patients. AIDS. 1998;12:873–8. DOIPubMedGoogle Scholar
- Huang L, Beard CB, Creasman J, Levy D, Duchin JS, Lee S, Sulfa or sulfone prophylaxis and geographic region predict mutations in the Pneumocystis carinii dihydropteroate synthase gene. J Infect Dis. 2000;182:1192–8. DOIPubMedGoogle Scholar
- Armstrong W, Meshnick S, Kazanjian P. Pneumocystis carinii mutations associated with sulfa and sulfone prophylaxis failures in immunocompromised patients. Microbes Infect. 2000;2:61–7. DOIPubMedGoogle Scholar
- Mei Q, Gurunathan S, Masur H, Kovacs JA. Failure of co-trimoxazole in Pneumocystis carinii infection and mutations in dihydropteroate synthase gene. Lancet. 1998;351:1631–2. DOIPubMedGoogle Scholar
- Visconti E, Ortona E, Mencarini P, Margutti P, Marinaci S, Zolfo M, Mutations in dihydropteroate synthase gene of Pneumocystis carinii in HIV patients with Pneumocystis carinii pneumonia. Int J Antimicrob Agents. 2001;18:547–51. DOIPubMedGoogle Scholar
- Nahimana A, Rabodonirina M, Zannetti G, Meneau I, Franciolo P, Bille J, Association between a specific Pneumocystis jiroveci dihydropteroate synthase mutation and failure of pyrimethamine/sulfadoxine prophylaxis in HIV-positive and -negative patients. J Infect Dis. 2003;188:1017–23. DOIPubMedGoogle Scholar
- Latouche S, Lacube P, Maury E, Bolognini J, Develoux M, Girard PM, Pneumocystis jirovecii dihydropteroate synthase genotypes in French patients with pneumocystosis: a 1998–2001 prospective study. Med Mycol. 2003;41:533–7. DOIPubMedGoogle Scholar
- Lane BR, Ast JC, Hossler PA, Mindell DP, Bartlett MS, Smith JW, Dihydropteroate synthase polymorphisms in Pneumocystis carinii. J Infect Dis. 1997;175:482–5.PubMedGoogle Scholar
- Beard CB, Carter JL, Keely SP, Huang L, Pieniazek NJ, Moura IN, Genetic variation in Pneumocystis carinii isolates from different geographic regions: implications for transmission. Emerg Infect Dis. 2000;6:265–72. DOIPubMedGoogle Scholar
- Huang L, Friedly J, Morris AM, Carter JL, Turner JR, Merrifield C, Pneumocystis carinii dihydropteroate synthase genotypes in HIV-infected persons residing in San Francisco: possible implications for disease transmission. J Eukaryot Microbiol. 2001;48:137–8S. DOIPubMedGoogle Scholar
- Nevez G, Totet A, Pautard JC, Raccurt C. Pneumocystis carinii detection using nested-PCR in nasopharyngeal aspirates of immunocompetent infants with bronchiolitis. J Eukaryot Microbiol. 2001;48(Suppl):S122–3. DOIPubMedGoogle Scholar
- Vargas SL, Hughes WT, Santolaya ME, Ulloa AV, Ponce CA, Cabrera CE, Search for primary infection by Pneumocystis carinii in a cohort of normal, healthy infants. Clin Infect Dis. 2001;32:855–61. DOIPubMedGoogle Scholar
- Totet A, Pautard JC, Raccurt C, Roux P, Nevez G. Genotypes at the internal transcribed spacers of the nuclear rRNA operon of Pneumocystis jiroveci in nonimmunosuppressed infants without severe pneumonia. J Clin Microbiol. 2003;41:1173–80. DOIPubMedGoogle Scholar
- Totet A, Latouche S, Lacube P, Pautard JC, Jounieaux V, Roux P, Pneumocystis carinii dihydropteroate synthase genotypes in immunocompetent infants and immunosuppressed adults from the same French geographic region. [Suppl]. Am J Respir Crit Care Med. 2002;▪▪▪:165.PubMedGoogle Scholar
- Demanche C, Berthelemy M, Petit T, Polack B, Wakefield AE, Dei-Cas E, Phylogeny of Pneumocystis carinii from 18 primate species confirms host specificity and suggests coevolution. J Clin Microbiol. 2001;39:2126–33. DOIPubMedGoogle Scholar
- Navin TR, Beard CB, Huang L, del Rio C, Lee S, Pieniazek NJ, Effect of mutations in Pneumocystis carinii dihydropteroate synthase gene on outcome of P. carinii pneumonia in patients with HIV-1: a prospective study. Lancet. 2001;358:545–9. DOIPubMedGoogle Scholar
- Takahashi T, Hosoya N, Endo T, Nakamura T, Sakashita HK, Imura K, Relationship between mutations in dihydropteroate synthase of Pneumocystis carinii f. sp. hominis isolates in Japan and resistance to sulfonamide therapy. J Clin Microbiol. 2000;38:3161–4.PubMedGoogle Scholar
- Wakefield AE, Lindley AR, Ambrose HE, Denis MC, Miller RF. Limited asymptomatic carriage of Pneumocystis jiroveci in human immunodeficiency virus–infected patients. J Infect Dis. 2003;187:901–8. DOIPubMedGoogle Scholar
- Volpe F, Dyer M, Scaife JG, Darby G, Stammers DK, Delves CJ. The multifunctional folic acid synthesis fas gene of Pneumocystis carinii appears to encode dihydropteroate synthase and hydroxymethyldihydropterin pyrophosphokinase. Gene. 1992;112:213–8. DOIPubMedGoogle Scholar
- Ma L, Kovacs JA. Rapid detection of mutations in the human-derived Pneumocystis carinii dihydropteroate synthase gene associated with sulfa resistance. Antimicrob Agents Chemother. 2001;45:776–80. DOIPubMedGoogle Scholar
- Helweg-Larsen J, Eugen-Olsen J, Lundgren B. Rapid detection of dihydropteroate polymorphism in AIDS-related Pneumocystis carinii pneumonia by restriction fragment length polymorphism. Scand J Infect Dis. 2000;32:481–3. DOIPubMedGoogle Scholar
- Huang L, Morris AM, Beard CB. Pneumocystis carinii dihydropteroate synthase mutations and treatment with sulfa or sulfone regimens: a proposal for standardized definitions for clinical evaluation. J Eukaryot Microbiol. 2001;48(Suppl):S180–1. DOIPubMedGoogle Scholar
- Dumoulin A, Mazars E, Seguy N, Gargallo-Viola D, Vargas S, Cailliez JC, Transmission of Pneumocystis carinii disease from immunocompetent contacts of infected hosts to susceptible hosts. Eur J Clin Microbiol Infect Dis. 2000;19:671–8. DOIPubMedGoogle Scholar
- Helweg-Larsen J, Tsolaki AG, Miller RF, Lundgren B, Wakefield AE. Clusters of Pneumocystis carinii pneumonia: analysis of person-to-person transmission by genotyping. Q J Med. 1998;91:813–20.PubMedGoogle Scholar
- Miller RF, Ambrose HE, Novelli V, Wakefield AE. Probable mother-to-infant transmission of Pneumocystis carinii f. sp. hominis infection. J Clin Microbiol. 2002;40:1555–7. DOIPubMedGoogle Scholar
- Durand-Joly I, Aliouat el M, Recourt C, Guyot K, Francois N, Wauquier M, et al. Pneumocystis carinii f. sp. hominis is not infectious for SCID mice. J Clin Microbiol. 2002;40:1862–5. DOIPubMedGoogle Scholar
- Institut de veille sanitaire. Surveillance du SIDA en France. Bulletin Epidémiologique Hebdomadaire. 2002;27:133–8.
- Miller RF, Lindley AR, Ambrose HE, Malin AS, Wakefield AE. Genotypes of Pneumocystis jiroveci isolates obtained in Harare, Zimbawe, and London, United Kingdom. Antimicrob Agents Chemother. 2003;47:3979–81. DOIPubMedGoogle Scholar
- Beard CB, Fox MR, Hanzlick RL, Guarner J, Carter JL, Del Rio C, Independent transmission cycles for Pneumocystis carinii pneumonia in immunocompetent infants and in adults with AIDS. In: Abstracts of the VII International Worskhops on Opportunistic Protists; Cincinnati, Ohio; 2001 June 13–16; Abstract PL94.
- Nevez G, Raccurt C, Vincent P, Jounieaux V, Dei-Cas E. Pulmonary colonization with Pneumocystis carinii in human immunodeficiency virus–negative patients: assessing risk with blood CD4+ T cell counts. Clin Infect Dis. 1999;29:1331–2. DOIPubMedGoogle Scholar
- Armbruster C, Hassl A, Kriwanek S. Pneumocystis carinii colonization in the absence of immunosuppression. Scand J Infect Dis. 1997;29:591–3. DOIPubMedGoogle Scholar
- Sing A, Roggenkamp A, Autenrieth IB, Heesemann J. Pneumocystis carinii carriage in immunocompetent patients with primary pulmonary disorders as detected by single or nested PCR. J Clin Microbiol. 1999;37:3409–10.PubMedGoogle Scholar
- Miller RF, Ambrose HE, Wakefield AE. Pneumocystis carinii f. sp. hominis DNA in immunocompetent health care workers in contact with patients with P. carinii pneumonia. J Clin Microbiol. 2001;39:3877–82. DOIPubMedGoogle Scholar
Figures
Tables
Cite This ArticleTable of Contents – Volume 10, Number 4—April 2004
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Gilles Nevez, Department of Parasitology, Mycology and Travel Medicine, University Hospital Centre, 1 avenue René Laennec, 80054 Amiens, France, EU; fax: 33-3-22-45-56-53
Top