Volume 15, Number 10—October 2009
Dispatch
Orangutans Not Infected with Plasmodium vivax or P. cynomolgi, Indonesia
Cite This Article
Citation for Media
Abstract
After orangutans in Indonesia were reported as infected with Plasmodium cynomolgi and P. vivax, we conducted phylogenetic analyses of small subunit ribosomal RNA gene sequences of Plasmodium spp. We found that these orangutans are not hosts of P. cynomolgi and P. vivax. Analysis of >1 genes is needed to identify Plasmodium spp. infecting orangutans.
Parasites belonging to the genus Plasmodium cause malaria and are usually host specific. For example, humans are natural hosts for P. falciparum, P. vivax, P. malaria, and P. ovale, and orangutans are naturally infected with P. pitheci and P. silvaticum (1,2). However, simian malaria parasites can infect humans (1); for example, P. knowlesi, normally associated with infections in long-tailed and pig-tailed macaques, has recently been found to have caused malaria in humans in several countries in Southeast Asia (3–8). This finding raises the possibility that other zoonotic malaria parasites may emerge in humans.
Malaria parasites have distinct small subunit ribosomal RNA (SSU rRNA) genes that are developmentally regulated (9). Each Plasmodium species has at least 2 types of SSU rRNA genes, and the stage-specific expression of these genes varies among species. In general, the A-type genes are transcribed or expressed mainly during the asexual stages, and the S-type genes are transcribed mainly during the sporozoite stage. P. vivax also has O-type genes, which are expressed during ookinete and oocyst development. Phylogenetic analysis of the P. vivax and P. cynomolgi SSU rRNA genes has indicated that the genes appear to have evolved as a result of 2 gene duplication events (10). A more recent study, involving SSU rRNA sequence data from a much larger number of Plasmodium spp., demonstrated that gene duplication events giving rise to the A-type and S-type sequences took place independently at least 3 times during the evolution of Plasmodium spp. (11).
Reid et al. (12) analyzed the DNA sequences of SSU rRNA genes of Plasmodium spp. from blood of orangutans in Kalimantan, Indonesia. Using phylogenetic analysis, they concluded that, in addition to P. pitheci and P. silvaticum, the orangutans were infected with the human malaria parasite P. vivax and the macaque malaria parasite P. cynomolgi. Their report implies that human and macaque malaria parasites could be transmitted to orangutans and that orangutans could act as reservoir hosts for at least 1 of the human malaria parasites.
When taxonomic inferences of species within a genus are made from phylogenetic trees, trees must be reconstructed by using orthologous genes and must include as many species sequences as possible. However, Reid et al. used sequence data of only the S-type SSU rRNA genes for P. vivax, P. cynomolgi, and P. knowlesi and data of only the A-type genes for P. inui and P. fragile. Furthermore, they analyzed sequence data from only 4 simian malaria parasites. Nishimoto et al. recently included data from 10 simian malaria parasites (11). We therefore reanalyzed the SSU rRNA sequence data of malaria parasites of orangutans together with the A-type, S-type, and O-type SSUrRNA gene sequence data for various Plasmodium spp.
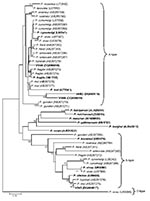
We used the neighbor-joining method, as described previously, to reconstruct the phylogenetic tree (3). Our phylogenetic analyses showed that SSU rRNA gene sequences VM88, VM82, and VM40 from orangutans (12) represent A-type SSU rRNA genes and that the VS63 sequence represents an S-type gene of Plasmodium spp. (Figure). No morphologic features of the malaria parasite stages in the blood were described for the Kalimantan orangutans by Reid et al. (12). Therefore, on the basis of SSU rRNA sequence data available for VM82 and VM88, whether these represent P. pitheci or P. silvaticum, previously described malaria parasites of orangutans, or some other species of Plasmodium cannot be determined with certainty.
The VS63 sequence is clearly not P. vivax, as previously reported by Reid et al. (12); it represents a Plasmodium sp. that is closely related to P. inui. It is most probably the S-type gene for either VM82 or VM88, which are A-type genes of P. pitheci and/or P. silvaticum. Furthermore, the VM40 sequence from orangutans represents a Plasmodium sp. closely related to the gibbon malaria parasite, P. hylobati (1), and is not the macaque malaria parasite, P. cynomolgi, as previously reported (12).
Phylogenetic analyses of the SSU rRNA genes indicate that none of the Plasmodium spp. isolated from orangutans in Kalimantan, Indonesia, are P. cynomolgi or P. vivax, as previously reported by Reid et al. (12). Before any conclusion about the identity of the malaria parasites infecting orangutans and their corresponding SSU rRNA gene sequences can be derived, a second or third gene of malaria parasites from these orangutans needs to be analyzed and the morphology of the corresponding blood stages needs to be described. Our study underscores the importance of using orthologous genes and sequence data from as many species as possible when inferring species within a genus from phylogenetic trees.
Prof Singh is director of the Malaria Research Centre, Universiti Malaysia Sarawak. His main research interests are centered on understanding the epidemiology of P. knowlesi and other primate malarias.
Mr Divis recently completed a Master of Science degree at Universiti Malaysia Sarawak, where he worked on the molecular characterization of malaria parasites in macaques. His research interests focus on the transmission of primate malarias.
Acknowledgment
This research was supported by grants from the Wellcome Trust, UK, and Universiti Malaysia Sarawak to B.S. and P.C.S.D.
References
- Garnham PCC. Malaria parasites and other haemosporidia. Oxford (UK): Blackwell Scientific Publications; 1966.
- Garnham PCC, Rajapaksa N, Peters W, Killick-Kendrick R. Malaria parasites of the orang-utan (Pongo pygmaeus). Ann Trop Med Parasitol. 1972;66:287–94.PubMedGoogle Scholar
- Singh B, Lee KS, Matusop A, Radhakrishnan A, Shamsul SSG, Cox-Singh J, A large focus of naturally acquired Plasmodium knowlesi infections in human beings. Lancet. 2004;363:1017–24. DOIPubMedGoogle Scholar
- Jongwutiwes S, Putaporntip C, Iwasaki T, Sata T, Kanbara H. Naturally acquired Plasmodium knowlesi malaria in human, Thailand. Emerg Infect Dis. 2004;10:2211–3.PubMedGoogle Scholar
- Zhu H, Li J, Zheng H. Human natural infection of Plasmodium knowlesi [in Chinese]. Chinese Journal of Parasitology and Parasitic Diseases. 2006;24:70–1.PubMedGoogle Scholar
- Luchavez J, Espino F, Curameng P, Espina R, Bell D, Chiodini P, Human infections with Plasmodium knowlesi, the Philippines. Emerg Infect Dis. 2008;14:811–3. DOIPubMedGoogle Scholar
- Ng OT, Ooi EE, Lee CC, Lee PJ, Ng LC, Pei SW, Naturally acquired human Plasmodium knowlesi infection, Singapore. Emerg Infect Dis. 2008;14:814–6. DOIPubMedGoogle Scholar
- Cox-Singh J, Davis TEM, Lee KS, Shamsul SSG, Matusop A, Ratnam S, Plasmodium knowlesi malaria in humans is widely distributed and potentially life threatening. Clin Infect Dis. 2008;46:165–71. DOIPubMedGoogle Scholar
- Rogers MJ, Li J, McCutchan F. The Plasmodium rRNA genes: developmental regulation and drug target. In: Sherman IW, editor. Malaria: parasite biology, pathogenesis and protection. Washington: American Society for Microbiology; 1998. p. 203–17.
- Li J, Gutell RR, Damberger SH, Wirtz RA, Kissinger JC, Rogers MJ, Regulation and trafficking of three distinct 18 S ribosomal RNAs during development of the malaria parasite. J Mol Biol. 1997;269:203–13. DOIPubMedGoogle Scholar
- Nishimoto Y, Arisue N, Kawai S, Escalante AA, Horii T, Tanabe K, Evolution and phylogeny of the heterogeneous cytosolic SSU rRNA genes in the genus Plasmodium. Mol Phylogenet Evol. 2008;47:45–53. DOIPubMedGoogle Scholar
- Reid MJ, Ursic R, Cooper D, Nazzari H, Griffiths M, Galdikas BM, Transmission of human and macaque Plasmodium spp. to ex-captive orangutans in Kalimantan, Indonesia. Emerg Infect Dis. 2006;12:1902–8.PubMedGoogle Scholar
Figure
Cite This ArticleTable of Contents – Volume 15, Number 10—October 2009
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Balbir Singh, Malaria Research Centre, Universiti Malaysia Sarawak, Jalan Tun Ahmad Zaidi Adruce, Kuching 93150, Malaysia
Top