Volume 15, Number 11—November 2009
Dispatch
Hemorrhagic Fever with Renal Syndrome in 4 US Soldiers, South Korea, 2005
Figure 2
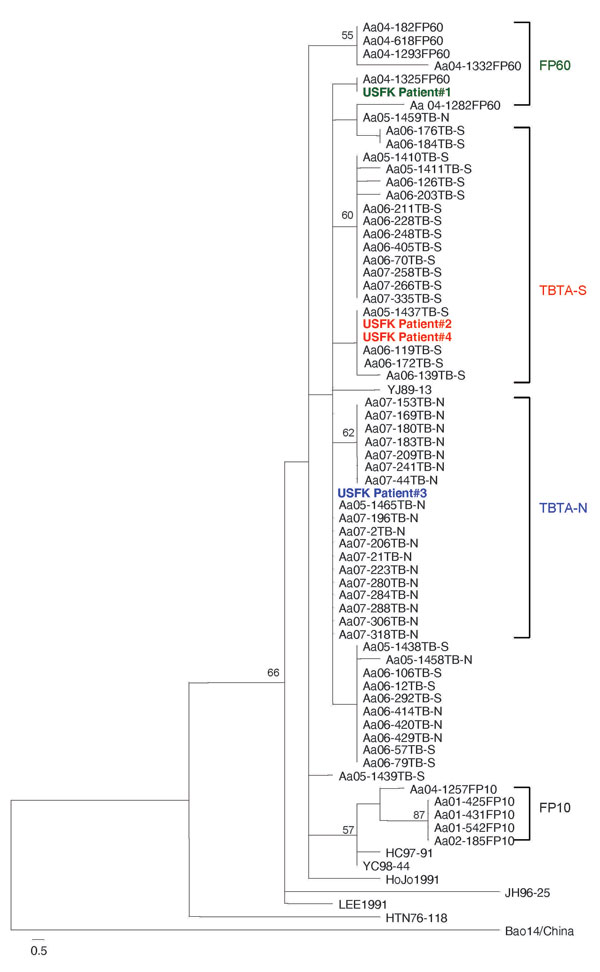
Figure 2. Phylogenetic tree by maximum parsimony method, rooted at the midpoint, based on the 320-bp region of G2 glycoprotein–encoding medium segment of 4 hemorrhagic fever with renal syndrome patients who were US soldiers in South Korea (patients 1–4), 2005 (GenBank accession nos. FJ561275–FJ561278) and field mice (Apodemus spp.)–borne Hantaan viruses (HTNV). HTNV sequence amplified from patient 1 was identical with a HTNV sequence (Aa04–1325) from A. agrarius mice captured at firing point (FP) 60. HTNV sequences from patients 2 and 4 were the same as 3 HTNV sequences (Aa05–1437, Aa06–119, Aa06–171) from A. agrarius mice captured at Twin Bridge Training Area–South (TBTA-S), and the HTNV sequence from patient 3 was identical to 11 HTNV sequences (Aa05–1465, Aa07–2, Aa07–21, Aa07–196, Aa07–206, Aa07–223, Aa07–280, Aa07–284, Aa07–288, Aa07–306 and Aa07–318) from A. agrarius mice at Twin Bridge Training Area-North (TBTA-N). Branch lengths are proportional to the number of nucleotide substitutions, while vertical distances are for clarity only. The numbers at each node are bootstrap probabilities (expressed as percentages), as determined for 100 iterations by using PAUP version 4.0b (http://paup.csit.fsu.poly). The colors indicate patients and corresponding training sites.
1Current affiliation: Centers for Disease Control and Prevention, Atlanta, Georgia, USA.
2Current affiliation: University of North Carolina at Chapel Hill, Chapel Hill, North Carolina, USA.