Volume 16, Number 12—December 2010
Research
Reassortant Group A Rotavirus from Straw-colored Fruit Bat (Eidolon helvum)
Figure 4
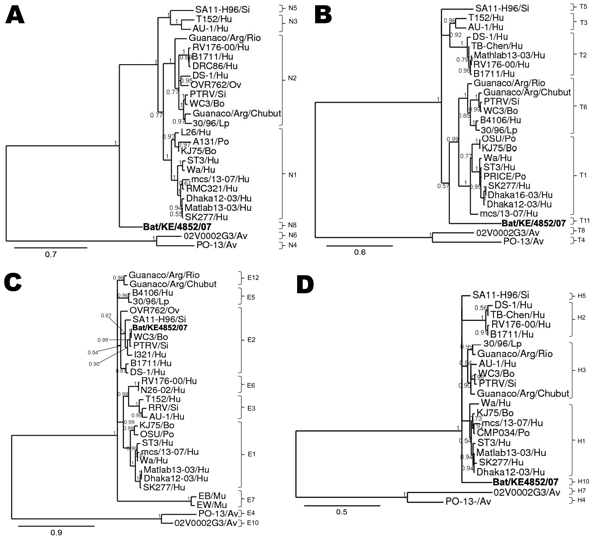
Figure 4. Phylograms indicating genetic relationships of partial or complete nucleotide sequences of A) nonstructural protein 2 (NSP2), B) NSP3, C) NSP4, and D) NSP5 of bat rotavirus strain Bat/KE4852/07 (boldface) from Kenya with representatives of known human and animal rotavirus genotypes. Posterior probability values are indicated at each branch node. Scale bars indicate nucleotide substitutions per site. GenBank accession numbers of all strains used are listed in the Technical Appendix. Genotypes of each gene segment characterized in this study are listed to the right of each tree. Si, simian; Hu, human; Ov, ovine; Bo, bovine; Lp, lapine; PO, porcine; Av, avian; Mu, murine.
Page created: August 29, 2011
Page updated: August 29, 2011
Page reviewed: August 29, 2011
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.