Volume 16, Number 9—September 2010
Dispatch
Co-infections with Plasmodium knowlesi and Other Malaria Parasites, Myanmar
Cite This Article
Citation for Media
Abstract
To determine the frequency of co-infections with Plasmodium species in southern Myanmar, we investigated the prevalence of P. knowlesi. More than 20% of patients with malaria had P. knowlesi infection, which occurred predominantly as a co-infection with either P. falciparum or P. vivax.
Plasmodium species are co-endemic to regions of Southeast Asia (1,2). This finding is believed to be underestimated because of insufficient sensitivity of microscopic detection of parasites. The prevalence of mixed infections with malaria parasites in the border regions between Thailand and Myanmar was recently found to be <24% (3). Identification of P. knowlesi as the fifth human malaria pathogen, which is prevalent in countries in Southeast Asia, has complicated this situation. P. knowlesi is a parasite that infects mainly long-tailed macaques (Macaca fascicularis) and pig-tailed macaques (M. nemestrina) in Southeast Asia (4). The parasite has developed the capacity to naturally infect humans, and infections in some persons have been life-threatening (5,6). Furthermore, infections with P. knowlesi in travelers to this region have been increasing (7,8).
P. knowlesi isolates obtained from humans have been frequently misidentified as P. falciparum or P. malariae because of the morphologic similarities of these parasites (2). Use of PCRs specific for 18S small subunit (SSU) rRNA genes of malaria parasites has identified suspected cases (2,9). P. knowlesi infection in humans in the border area between the People’s Republic of China and Myanmar has been reported (10), but the prevalence is unknown. We investigated the frequency of co-infections with P. knowlesi and other Plasmodium spp. in this region.
The study was reviewed and approved by the Ethic Committee of the Institute for Parasitic Disease Control of Yunnan Province and local administration authority in Myanmar. One hundred forty-six blood samples were obtained in 2008 from randomly selected patients with uncomplicated malaria in southern Myanmar near Yunnan Province of China, where pig-tailed macaques are also present. Written consent was obtained from each person before blood samples were obtained. A drop (20 µL–50 µL) of fingerprick blood was placed directly on premarked filter paper. Malaria infection was identified by microscopic analysis of Giemsa-stained blood films made from blood spotted on the paper.
DNA templates for a nested PCR were prepared from whole blood spots on filter paper according to a previously reported method (11). Genomic DNA of P. falciparum was obtained from in vitro–proliferated 3D7 clone. Genomic DNA of P. vivax was obtained from a patient from the study region previously identified by using PCR and the primer sets used in this study. As reported (2,12), DNA (3 μL/sample and 1 ng of each positive control DNA) from each sample was amplified with the Plasmodium genus–specific primer pair rPLU1 and rPLU5.
Two microliters of PCR product from each amplification was subjected to a second PCR amplification with species-specific primer pairs rFAL1 and rFAL2 for P. falciparum, rMAL1 and rMAL2 for P. malariae, rVIV1 and rVIV2 for P. vivax, rOVA1 and rOVA2 for P. ovale, and Pmk8 and Pmkr9 for P. knowlesi. PCR products amplified with nested primers were analyzed by agarose gel electrophoresis. DNA bands were removed from the gel, purified by using the QIAquick Gel Extraction Kit (QIAGEN, Valencia, CA, USA) and ligated to T-cloning vector (Invitrogen, Carlsbad, CA, USA) according to protocols provided by the manufacturers. Plasmid inserts were then sequenced.
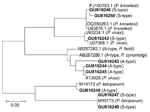
Sequence identity was confirmed by random basic local alignment search tool analysis of sequences in GenBank (http://blast.ncbi.nlm.nih.gov/). Novel sequences were deposited in GenBank with accession nos. GU816242–GU816250. Phylogenetic relationships of unique sequences amplified by using nested primers with corresponding reference sequences were constructed by using the neighbor-joining method in MEGA version 4.0 (13). All sequences clustered with reference sequences of P. falciparum, P. vivax, or P. knowlesi, which suggested that all sequences were species specific (Figure).
Three parasite species (P. falciparum, P. vivax, and P. knowlesi) were identified in 146 infected persons. Phylogenetic analysis showed that amplified products were species specific (Figure). Monoinfection with P. falciparum, P. vivax, and P. knowlesi accounted for 34.9% (51/146), 36.3% (53/146), and 2.7% (4/146), respectively, of the infections. Mixed infections of P. knowlesi with P. falciparum or P. vivax accounted for 6.9% (10/146) of the infections, and mixed infections with P. knowlesi and either P. falciparum or P. vivax accounted for 8.9% (n = 13 in both groups) of the infections. Only 2 samples (1.4%) had mixed infections with P. falciparum, P. vivax, and P. knowlesi (Table). Thus, the prevalence of mixed infections in southern Myanmar was lower than that in northern Myanmar near the border with Thailand (3). The prevalence of P. knowlesi was 21.9%. In most cases, this parasite showed co-infection with either P. falciparum or P. vivax, which indicated that P. knowlesi may have not fully adapted to the human host or that humans who were infected with other malaria parasites may be more vulnerable to P. knowlesi infection.
Our study also emphasizes the need for improvement of current methods for detecting P. knowlesi infection (14,15). A recent report found that the primer pair Pmk8 and Pmkr9, which is specific for the 18S SSU rRNA gene of P. knowlesi, can cross-hybridize with the corresponding sequence of P. vivax (14). We observed weak amplifications in 16 samples (11%); all were from P. vivax–infected blood. All amplicons made by using Pmk8 and Pmkr9 primers were sequenced and compared with the homologous sequences. Because of the high similarity of 18S SSU rRNA gene sequences among these parasites, more specific sequences are needed for establishing a reliable PCR-based method for routine diagnosis of P. knowlesi infection (14).
Dr Jiang is a research scientist at the Key Laboratory of Zoonosis, Jilin University, Changchun, China. Her research interests are malaria epidemiology and host–parasite interactions.
Acknowledgments
We thank Clemens H. Kocken for providing P. knowlesi DNA and the volunteers for providing blood samples for this study.
This study was supported by the National Basic Research Program of China (973 Program, no. 2007CB513100 to Q.C.), a grant from the Young Distinguished Scientist Program of the National Natural Science Foundation of China (30625029), National Science and Technology project (2008zc10004-011), the Swedish Institute for Infectious Disease Control, and the Swedish International Development Cooperation Agency.
References
- McKenzie FE, Bossert WH. Mixed-species Plasmodium infections of humans. J Parasitol. 1997;83:593–600. DOIPubMedGoogle Scholar
- Singh B, Kim Sung L, Matusop A, Radhakrishnan A, Shamsul SS, Cox-Singh J, A large focus of naturally acquired Plasmodium knowlesi infections in human beings. Lancet. 2004;363:1017–24. DOIPubMedGoogle Scholar
- Putaporntip C, Hongsrimuang T, Seethamchai S, Kobasa T, Limkittikul K, Cui L, Differential prevalence of Plasmodium infections and cryptic Plasmodium knowlesi malaria in humans in Thailand. J Infect Dis. 2009;199:1143–50. DOIPubMedGoogle Scholar
- Butcher GA, Mitchell GH, Cohen S. Plasmodium knowlesi infections in a small number of non-immune natural hosts (Macaca fascicularis) and in rhesus monkeys (M. mulatta). Trans R Soc Trop Med Hyg. 2010;104:75–7. DOIPubMedGoogle Scholar
- Cox-Singh J, Singh B. Knowlesi malaria: newly emergent and of public health importance? Trends Parasitol. 2008;24:406–10. DOIPubMedGoogle Scholar
- Cox-Singh J, Davis TM, Lee KS, Shamsul SS, Matusop A, Ratnam S, Plasmodium knowlesi malaria in humans is widely distributed and potentially life threatening. Clin Infect Dis. 2008;46:165–71. DOIPubMedGoogle Scholar
- Bronner U, Divis PC, Färnert A, Singh B. Swedish traveler with Plasmodium knowlesi malaria after visiting Malaysian Borneo. Malar J. 2009;8:15. DOIPubMedGoogle Scholar
- van Hellemond JJ, Rutten M, Koelewijn R, Zeeman AM, Verweij JJ, Wismans PJ, Human Plasmodium knowlesi infection detected by rapid diagnostic tests for malaria. Emerg Infect Dis. 2009;15:1478–80. DOIPubMedGoogle Scholar
- Lee KS, Cox-Singh J, Brooke G, Matusop A, Singh B. Plasmodium knowlesi from archival blood films: further evidence that human infections are widely distributed and not newly emergent in Malaysian Borneo. Int J Parasitol. 2009;39:1125–8. DOIPubMedGoogle Scholar
- Zhu HM, Li J, Zheng H. Human natural infection of Plasmodium knowlesi [in Chinese]. Zhongguo Ji Sheng Chong Xue Yu Ji Sheng Chong Bing Za Zhi. 2006;24:70–1. PubMedGoogle Scholar
- Bereczky S, Mårtensson A, Gil JP, Färnert A. Short report: Rapid DNA extraction from archive blood spots on filter paper for genotyping of Plasmodium falciparum. Am J Trop Med Hyg. 2005;72:249–51. PubMedGoogle Scholar
- Singh B, Bobogare A, Cox-Singh J, Snounou G, Abdullah MS, Rahman HA. A genus- and species-specific nested polymerase chain reaction malaria detection assay for epidemiologic studies. Am J Trop Med Hyg. 1999;60:687–92. PubMedGoogle Scholar
- Ng OT, Ooi EE, Lee CC, Lee PJ, Ng LC, Pei SW, Naturally acquired human Plasmodium knowlesi infection, Singapore. Emerg Infect Dis. 2008;14:814–6. DOIPubMedGoogle Scholar
- Imwong M, Tanomsing N, Pukrittayakamee S, Day NP, White NJ, Snounou G. Spurious amplification of a Plasmodium vivax small-subunit RNA gene by use of primers currently used to detect P. knowlesi. J Clin Microbiol. 2009;47:4173–5. DOIPubMedGoogle Scholar
- Kawai S, Hirai M, Haruki K, Tanabe K, Chigusa Y. Cross-reactivity in rapid diagnostic tests between human malaria and zoonotic simian malaria parasite Plasmodium knowlesi infections. Parasitol Int. 2009;58:300–2. DOIPubMedGoogle Scholar
Figure
Table
Cite This Article1These authors contributed equally to this article.
Table of Contents – Volume 16, Number 9—September 2010
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Qijun Chen, Institute for Infectious Disease Control, Karolinska Institutet, 171 71 Stockholm, Sweden
Top