Volume 17, Number 10—October 2011
Dispatch
Plasmodium knowlesi Infection in Humans, Cambodia, 2007–2010
Cite This Article
Citation for Media
Abstract
Two cases of Plasmodium knowlesi infection in humans were identified in Cambodia by 3 molecular detection assays and sequencing. This finding confirms the widespread distribution of P. knowlesi malaria in humans in Southeast Asia. Further wide-scale studies are required to assess the public health relevance of this zoonotic malaria parasite.
In Cambodia, malaria ranks among the leading causes of illness and death. Mostly affecting the ≈3 million persons (23% of Cambodia’s population) who live near forested areas, malaria remains an occupational disease in specific high-risk groups, such as forestry workers and migrant populations who have come into forested areas. However, for the past decade, the number of reported malaria cases has generally decreased but in a sawtooth pattern of periodic increases (1).
Four of the 5 Plasmodium species known to cause malaria in humans have already been described in Cambodia (2,3). P. falciparum remains the most frequent cause of malaria (83,777 cases in 2009, prevalence of 70%) (1). However, distributions of Plasmodium species are changing, with a particularly substantial increase of P. vivax malaria cases, from 4,105 (8%) cases in 2000 to 6,250 (25%) in 2009. In several areas of low transmission, the proportion of P. vivax infections has increased up to 50% (2). This trend is probably related to various effective control strategies implemented in Cambodia against P. falciparum malaria.
No studies in humans (3) and monkeys in Cambodia have identified the simian malaria parasite, P. knowlesi, which is causing human disease in some other countries in Southeast Asia (4). P. knowlesi parasites were not detected in blood samples collected during 2004–2007 for 138 monkeys (102 Macaca fascicularis monkeys; 13 M. leonina monkeys; 20 Hyobates pileatus monkeys; 2 Presbytis cristata monkeys; and 1 Pongo pygmaeus monkey) by using PCR (cytb and cox1 genes) (L. Duval, unpub. data). Because these animals were confiscated by Wildlife Alliance rescue patrols from illegal traders, the locations where they were trapped in Cambodia are unknown.
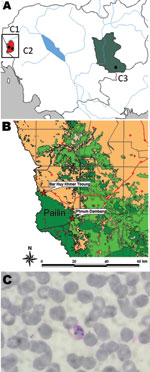
We undertook a cross-sectional prospective study in 3 sites in Cambodia (Figure, panel A) (5). The main objective of this study was to develop evidence to guide the management of malaria parasite–negative persons with acute febrile illness and to determine whether such persons were infected with P. knowlesi.
We enrolled all patients >7 years of age with acute (<8 days) febrile illness in selected outpatient clinics, where they had been tested for malaria by using rapid diagnostic test (CareStart, Access Bio Inc, Somerset, NJ, USA). A subset of nonfebrile patients were used as controls; patients in a critical clinical condition were excluded. After we obtained informed consent, each patient’s history was taken, a physical examination was conducted, and blood and throat swab samples were collected. The defined test battery, including the following pathogens, was performed by using a PCR/sequencing approach: Plasmodium spp., Leptospira spp., Rickettsia spp. (including Orientia tsutsugamushi), dengue viruses, Japanese encephalitis virus, and influenza viruses. HIV infection and tuberculosis were not evaluated. DNA and RNA from the erythrocyte pellet or throat swab were extracted by using the QIAamp kit (QIAGEN, Hilden, Germany), according to the manufacturer’s instructions. Malaria parasites were detected by using a Plasmodium spp.–specific nested PCR based on the cytb gene followed by sequencing (6).
During December 2007–December 2010, a total of 1,475 patients were enrolled (621 in Soun Kouma; 650 in Ou Chra, Pailin Province; and 204 in Snoul, Kratié Province), comprising 1,193 febrile and 282 nonfebrile persons. For 564 (38.2%) patients, no pathogens were detected. A total of 754 patients (51.1%; 676 cases, 78 controls) were infected with malaria parasites, and the distribution of Plasmodium spp. was as follows: P. vivax, 51.6%; P. falciparum, 40.6%; P. vivax/P. falciparum, 7.4%; P. knowlesi, 0.3%; and P. ovale, 0.1%.
We detected P. knowlesi infections in 2 patients from Ou Chra health center in Pailin Province (Figure, panel B). The first patient was a 41-year-old man from Borhuytbong village. He came in April 2010 to the health center with fever (38.5°C), chills, and headache. Rapid diagnostic test and microscopy were negative for malaria; PCR/sequencing was positive for P. knowlesi. No other pathogen was found. The patient was originally from Thailand and reported that he had not returned to Thailand since getting married in Cambodia in 1989. He reported self-treatment with mefloquine (Lariam [Roche, Basel, Switzerland]) and was cured in 3 days. He further added that he used to hunt and spent most of his time in forests around Pailin, where long-tailed macaques, the natural hosts of P. knowlesi (7), are usually found.
The second patient was a 40-year-old Khmer man from Phnomdambang village. He came in September 2010 to the health center because of chills and headache. No fever (36°C) was detected by the medical staff, but the patient reported a history of fever and had self-treated with acetaminophen. Results of rapid diagnostic test were negative, but microscopy results were positive (33 parasites/µL) (Figure, panel C). PCR amplification followed by sequencing confirmed P. knowlesi infection. No other pathogen was found. The patient reported having lived in this same village since 1995, and he frequently went to the forests around Pailin for hunting and collecting valued wood.
Both patients’ P. knowlesi infections were confirmed by additional PCR amplification and sequencing of the ldh, tufA, and cox genes (8) at the Genomic Platform, Institut Pasteur, Paris, France. Blood spots on filter paper were also sent blinded to the Malaria Research Centre, Faculty of Medicine and Health Sciences, University Malaysia, Sarawak, Malaysia, where they were identified as P. knowlesi single infections by real-time PCR (9) and by nested PCR with the primers Pmk8 and Pmkr9 (10) and PkF1140 and PkR1550 (11). The nucleotide sequence determined in this study has been deposited in the GenBank database and assigned accession nos. JF419317–JF419323.
Our findings confirm that P. knowlesi infections occur in humans in Cambodia, thereby increasing the number of countries in Southeast Asia with cases in humans (10). However, further wide-scale studies are required to assess the prevalence and distribution of P. knowlesi malaria cases in humans and monkeys. Such studies would enable an assessment of the public health relevance of this zoonotic malaria parasite and characterization of P. knowlesi malaria epidemiology in this region. They would be particularly useful in determining whether this simian species has been imported from neighboring countries by humans or whether P. knowlesi parasites are already circulating and are transmitted from monkey reservoir hosts to humans. Moreover, to address this issue, development of new tools, such as specific serologic markers, is urgently needed, in addition to molecular biology methods.
Dr Khim is an engineer and head of the molecular biology platform in the Malaria Molecular Epidemiology Unit at Pasteur Institute of Cambodia. Her research interests include developing molecular tools for improving the surveillance of resistance to antimalarial drugs in Cambodia.
Acknowledgments
We thank all the patients and their parents or guardians for participating in the study. We also thank all the health workers at the health centers of Soun Kouma, Ou Chra, and Snoul and the staff of the Ministry of Health of Cambodia. We are grateful to Iveth J. González, David Bell, Mark Perkins, Eva-Maria Christophel, Bayo Fatunmbi, Franz-Josef von Sonnenburg, Philippe Buchy, Bertrand Guillard, Abdur Rashid, and Steven Bjorge for their assistance with this study. We thank the Wildlife Alliance for their support and for providing the primate blood specimens.
This work was supported by grants from World Health Organization Regional Office for the Western Pacific, Foundation for Innovative New Diagnostics, and University of Munich.
References
- World Health Organization. Malaria morbidity and mortality by province in Cambodia. 2011 [cited 2011 Aug 3]. http://www.wpro.who.int/sites/mvp/epidemiology/malaria/cam_graphs.htm
- Incardona S, Vong S, Chiv L, Lim P, Nhem S, Sem R, Large-scale malaria survey in Cambodia: novel insights on species distribution and risk factors. Malar J. 2007;6:37. DOIPubMedGoogle Scholar
- Steenkeste N, Rogers WO, Okell L, Jeanne I, Incardona S, Duval L, Sub-microscopic malaria cases and mixed malaria infection in a remote area of high malaria endemicity in Rattanakiri province, Cambodia: implication for malaria elimination. Malar J. 2010;9:108. DOIPubMedGoogle Scholar
- Cox-Singh J, Singh B. Knowlesi malaria: newly emergent and of public health importance? Trends Parasitol. 2008;24:406–10. DOIPubMedGoogle Scholar
- Foundation for Innovative New Diagnostics. Non-malaria Febrile Illness Study Final Review Workshop. 2011 [cited 2011 Aug 3]. http://www.wpro.who.int/sites/mvp/meetings/Non-Malaria+Febrile+Illness+Study+Final+Review+Workshop.htm
- Steenkeste N, Incardona S, Chy S, Duval L, Ekala MT, Lim P, Towards high-throughput molecular detection of Plasmodium: new approaches and molecular markers. Malar J. 2009;8:86. DOIPubMedGoogle Scholar
- Garnham PCC. Malaria parasites and other haemosporidia. Oxford (UK): Blackwell Scientific Publications; 1966.
- Duval L, Fourment M, Nerrienet E, Rousset D, Sadeuh SA, Goodman SM, African apes as reservoirs of Plasmodium falciparum and the origin and diversification of the Laverania subgenus. Proc Natl Acad Sci U S A. 2010;107:10561–6. DOIPubMedGoogle Scholar
- Divis PC, Shokoples SE, Singh B, Yanow SK. A TaqMan real-time PCR assay for the detection and quantitation of Plasmodium knowlesi. Malar J. 2010;9:344. DOIPubMedGoogle Scholar
- Singh B, Kim Sung L, Matusop A, Radhakrishnan A, Shamsul SS, Cox-Singh J, A large focus of naturally acquired Plasmodium knowlesi infections in human beings. Lancet. 2004;363:1017–24. DOIPubMedGoogle Scholar
- Imwong M, Tanomsing N, Pukrittayakamee S, Day NP, White NJ, Snounou G. Spurious amplification of a Plasmodium vivax small-subunit RNA gene by use of primers currently used to detect P. knowlesi. J Clin Microbiol. 2009;47:4173–5. DOIPubMedGoogle Scholar
Figure
Cite This Article1These authors contributed equally to this article.
Table of Contents – Volume 17, Number 10—October 2011
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Didier Ménard, Molecular Epidemiology Unit–Pasteur Institute of Cambodia, 5 Blvd Monivong, PO Box 983, Phnom Penh, Cambodia
Top