Volume 18, Number 1—January 2012
Dispatch
Mutations I117V and I117M and Oseltamivir Sensitivity of Pandemic (H1N1) 2009 Viruses
Figure 2
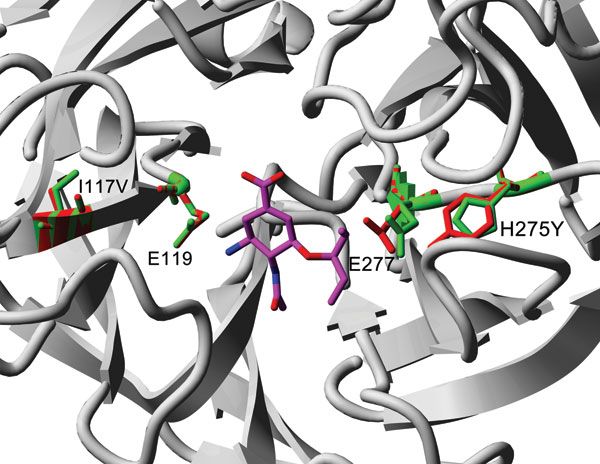
Figure 2. Comparison of wildtype I117 mutation from pandemic (H1N1) 2009 viruses (green residues; Protein Data Bank: 3nss) with FoldX/YASARA model of I117V + H275Y double mutant (red residues) (11,12). Residue numbering is according to pandemic (H1N1) 2009 neuraminadase. Oseltamivir is added as reference (Protein Data Bank: 3clO) and shown in magenta.
References
- Hurt AC, Lee R, Leang S, Cui L, Deng Y, Phuah S, Increased detection in Australia and Singapore of a novel influenza A(H1N1)2009 variant with reduced oseltamivir and zanamivir sensitivity due to a S247N neuraminidase mutation. Euro Surveill. 2011;16:pii:19884.
- Santos L, Correia V, Giria M, Pedro S, Santos M, Silvestre M, Genetic and antiviral drug susceptbility profiles of pandemic A(H1N1)v influenza virus circulating in Portugal. In: Proceedings of Options for the Control of Influenza VII, Hong Kong, SAR, China, September 1–3, 2010. Atlanta (GA): Intregress; 2010. Abstract no. O-851.
- van der Vries E, Stelma FF, Boucher CA. Emergence of a multidrug-resistant pandemic influenza A (H1N1) virus. N Engl J Med. 2010;363:1381–2. DOIPubMedGoogle Scholar
- Yi H, Lee JY, Hong EH, Kim MS, Kwon D, Choi JH, Oseltamivir-resistant pandemic (H1N1) 2009 virus, South Korea. Emerg Infect Dis. 2010;16:1938–42.PubMedGoogle Scholar
- Shin SY, Kang C, Gwack J, Kim JH, Kim HS, Kang YA, Drug-resistant pandemic (H1N1) 2009, South Korea. Emerg Infect Dis. 2011;17:702–4.PubMedGoogle Scholar
- Hurt AC, Selleck P, Komadina N, Shaw R, Brown L, Barr IG. Susceptibility of highly pathogenic A(H5N1) avian influenza viruses to the neuraminidase inhibitors and adamantanes. Antiviral Res. 2007;73:228–31. DOIPubMedGoogle Scholar
- Ilyushina NA, Seiler JP, Rehg JE, Webster RG, Govorkova EA. Effect of neuraminidase inhibitor–resistant mutations on pathogenicity of clade 2.2 A/Turkey/15/06 (H5N1) influenza virus in ferrets. PLoS Pathog. 2010;6:e1000933. DOIPubMedGoogle Scholar
- Hurt AC, Holien JK, Parker M, Barr IG. Oseltamivir resistance and the H274Y neuraminidase mutation in seasonal, pandemic and highly pathogenic influenza viruses. Drugs. 2009;69:2523–31. DOIPubMedGoogle Scholar
- Hurt AC, Holien JK, Barr IG. In vitro generation of neuraminidase inhibitor resistance in A(H5N1) influenza viruses. Antimicrob Agents Chemother. 2009;53:4433–40. DOIPubMedGoogle Scholar
- Hurt AC, Barr IG, Hartel G, Hampson AW. Susceptibility of human influenza viruses from Australasia and South East Asia to the neuraminidase inhibitors zanamivir and oseltamivir. Antiviral Res. 2004;62:37–45. DOIPubMedGoogle Scholar
- Van Durme J, Delgado J, Stricher F, Serrano L, Schymkowitz J, Rousseau F. A graphical interface for the FoldX forcefield. Bioinformatics. 2011;27:1711–2. DOIPubMedGoogle Scholar
- Krieger E, Koraimann G, Vriend G. Increasing the precision of comparative models with YASARA NOVA—a self-parameterizing force field. Proteins. 2002;47:393–402. DOIPubMedGoogle Scholar
- Li Q, Qi J, Zhang W, Vavricka CJ, Shi Y, Wei J, The 2009 pandemic H1N1 neuraminidase N1 lacks the 150-cavity in its active site. Nat Struct Mol Biol. 2010;17:1266–8. DOIPubMedGoogle Scholar
- Peng AW, Milleri S, Stein DS. Direct measurement of the anti-influenza agent zanamivir in the respiratory tract following inhalation. Antimicrob Agents Chemother. 2000;44:1974–6. DOIPubMedGoogle Scholar
- He G, Massarella J, Ward P. Clinical pharmacokinetics of the prodrug oseltamivir and its active metabolite Ro 64–0802. Clin Pharmacokinet. 1999;37:471–84. DOIPubMedGoogle Scholar
Page created: December 22, 2011
Page updated: December 22, 2011
Page reviewed: December 22, 2011
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.