Volume 18, Number 11—November 2012
Research
Lack of Cross-protection against Bordetella holmesii after Pertussis Vaccination
Cite This Article
Citation for Media
Abstract
Bordetella holmesii, a species closely related to B. pertussis, has been reported sporadically as a cause of whooping cough–like symptoms. To investigate whether B. pertussis–induced immunity is protective against infection with B. holmesii, we conducted an analysis using 11 human respiratory B. holmesii isolates collected during 2005–2009 from a highly B. pertussis–vaccinated population in Massachusetts. Neither whole-cell (wP) nor acellular (aP) B. pertussis vaccination conferred protection against these B. holmesii isolates in mice. Although T-cell responses induced by wP or aP cross-reacted with B. holmesii, vaccine-induced antibodies failed to efficiently bind B. holmesii. B. holmesii–specific antibodies provided in addition to wP were sufficient to rapidly reduce B. holmesii numbers in mouse lungs. Our findings suggest the established presence of B. holmesii in Massachusetts and that failure to induce cross-reactive antibodies may explain poor vaccine-induced cross-protection.
Bordetella pertussis and B. parapertussis commonly cause whooping cough, a highly contagious, acute coughing illness, in humans (1,2). Licensed in the mid-1940s, the first whooping cough vaccines consisted of whole-cell inactivated B. pertussis (wP), and their use led to a dramatic decrease in disease incidence by the mid-1960s (2,3). The potential for health risks related to wP vaccine, however, led to adaptation of acellular (aP) vaccines, which contain combinations of purified B. pertussis proteins. Despite high vaccine coverage, reported whooping cough incidence in industrialized countries has been increasing during the past 20 years, although this could be the result of greater awareness and improved analytical tools (4–9).
In November 1983, the Centers for Disease Control and Prevention (CDC) received a gram-negative bacterium isolated from an asplenic patient (10). During the following decade, additional clinical isolates with the same microbiological characteristics (slow-growing, gram-negative, small coccoid, asaccharolytic, oxidase negative, nonmotile and brown-soluble-pigment-producing) were submitted to the CDC for identification (10). Subsequent biochemical analysis, 16S rRNA sequencing, and DNA relatedness studies revealed that these strains were new Bordetella species, which was named Bordetella holmesii to honor Barry Holmes (10). Since then, this bacterium has been isolated from numerous countries, including Australia, Canada, Chile, France, Germany, Japan, Netherlands, Switzerland, the United Kingdom, and the United States (6–15). These findings indicate that B. holmesii is a widespread pathogen among populations that are highly vaccinated against B. pertussis.
Comparative analysis of B. holmesii and B. pertussis by using 16S rRNA suggests that B. holmesii is closely related to B. pertussis, but further analysis of cellular fatty acid composition, housekeeping genes, and the BvgAS locus suggests that B. holmesii may not share many of the highly conserved virulence factors of B. pertussis (16). Antibodies against B. pertussis pertactin, pertussis toxin, fimbriae, adenylate cyclase toxin, and filamentous hemagglutinin recognize few, if any, proteins from multiple B. holmesii isolates (14), results that suggest B. holmesii may be antigenically distinct from B. pertussis.
Although B. holmesii has been isolated primarily from immunocompromised hosts (asplenic or sickle cell disease patients and transplant recipients) (14,17–20) and was first isolated from blood, the bacterium has also been found to cause respiratory diseases (11,12,21–23). B. holmesii was isolated from pleural fluid and lung biopsy specimens from an immunocompetent adolescent who had fever and pulmonary fibrosis (12) and from sputum of patients with respiratory failure (22). Moreover, B. holmesii was isolated from nasopharyngeal specimens of previously healthy persons who had whooping cough–like symptoms, including paroxysms, whooping, or post-tussive vomiting (11,21,23). Therefore, B. holmesii appears to be able to colonize the respiratory tract in the same manner as other Bordetella species. A case study in Japan also found epidemiologic links between 5 persons colonized with B. holmesii, which indicates the ability of this pathogen to transmit from person to person (6).
In collaboration with the Massachusetts Department of Public Health (MDPH), we reviewed B. holmesii surveillance data collected in Massachusetts during 2005–2009. B. holmesii was isolated from several patients experiencing whooping cough–like symptoms. By using a murine infection model, we examined the effects of B. pertussis vaccination on B. holmesii infection susceptibility.
Identification of B. holmesii Cases in Massachusetts
Culture-confirmed B. holmesii cases identified during 2005–2009 by the State Laboratory Institute at the MDPH were included in our analysis. According to MDPH guidelines, a nasopharyngeal swab was cultured if the patient was <11 years of age or had a cough for <14 days. For all other patients (>11 years of age and >14 days of cough), a serum test was performed. Details on culturing methods and Bordetella spp. identification tests performed have been described (21). A total of 41 B. holmesii infections were reported; the case records, including symptomology for 26 of these, are maintained in the Massachusetts Virtual Epidemiologic Network.
Bacterial Strains and Growth
B. pertussis strain 536 (24) and B. parapertussis strain CN2591 (25) have been described. B. holmesii strain P3421 was isolated in Massachusetts and used for animal experiments. Bacteria were maintained on Bordet-Gengou agar (Difco, Sparks, MD, USA) supplemented with 10% sheep’s blood (Hema Resources, Aurora, OR, USA) without antimicrobial drugs (B. holmesii) or with 20 µg/mL streptomycin (Sigma-Aldrich, St. Louis, MO, USA) (B. pertussis or B. parapertussis). Liquid cultures were grown overnight in Stainer-Scholte broth at 37°C to mid-log phase (26,27).
Phylogenetic Analysis
Phylogenetic analyses were performed on the basis of atpD, rpoB, tuf, and rnpB gene sequences as described (16). Gene amplifications were completed on 30 B. holmesii isolates obtained from MDPH or the CDC, B. pertussis strain 536, B. parapertussis strain 2591, B. bronchiseptica strain RB50, B. avium strain 197N, and B. hinzii strain BC304. Concatenated sequences were aligned and used to construct unweighted pair group method using average linkages trees in MEGA4 software (www.megasoftware.net/mega4/mega.html).
Animal Experiments
All protocols were approved by Institutional Animal Care and Use Committee (IACUC). All animals were C57BL/6 mice and were handled in accordance with institutional guidelines (IACUC approval no. 31297). Animal experiments were performed as described, with 4 mice per group, and were performed in replicate (28–33). For vaccinations, sedated 4–6-week-old mice were vaccinated by intraperitoneal injection of 108 CFU of heat-inactivated (65°C for 30 min) bacteria in 200 µL of phosphate-buffered saline (PBS; Omnipur, Gibbstown, NJ, USA) for whole-cell B. holmesii vaccine (wH) or wP B. pertussis vaccine; one fifth human dose of Adacel (Sanofi-Pasteur, Swiftwater, PA, USA) (0.5 µg PT, 1 µg FHA, 0.6 µg pertactin, 5 µg fimbriae 2 and 3 per mouse) with Imject Alum (Thermo Scientific) (aP); or only Imject Alum in 200 µL PBS on days 14 and 28 before challenge (28,33).
For challenge, 50 µL PBS containing 5 × 105 CFU of B. pertussis or B. parapertussis or 107 CFU of B. holmesii was added by pipetting onto the external nares of sedated mice (29). A larger inoculum of B. holmesii was used to achieve reproducibility and detect B. holmesii from the respiratory tract at later time points, because it is cleared more rapidly from the lower respiratory tract than are B. pertussis or B. parapertussis. For adoptive transfer of serum antibodies, mice were vaccinated with the indicated bacteria on days 0 and 14, and serum samples were collected on day 28 from vaccinated or naive animals. Serum samples of 200 µL were intraperitoneally injected immediately before mice were inoculated with 5 × 106 CFU of B. holmesii (30,32). Bacterial numbers were quantified as described (30).
Splenocyte Restimulations
Splenocytes were isolated from vaccinated mice as described (31,33) and stimulated with either media alone or media containing 107 CFU (multiplicities of infection of 5) of the indicated heat-killed bacteria (28,31). After 3 days, the supernatants were collected and analyzed for interferon (IFN) γ and interleukin-10 (IL-10) production by using ELISA (R&D Systems, Minneapolis, MN, USA) according to the manufacturer’s instructions.
Titer ELISAs
Antibody titers were determined as described (34–36). In brief, wP- or wH-induced/naive serum samples (1:200 dilution) or aP/adjuvant-induced serum samples (1:50 dilution) from each individual mouse were added to and serially diluted 1:2 across plates coated with heat-inactivated exponential-phase bacteria. After incubation, samples were probed with 1:4,000 dilution of goat anti-mouse Ig horseradish peroxidase–conjugated antibodies (Southern Biotech, Birmingham, AL, USA). Titers were determined by using the endpoint method (33).
Western Blot Analysis
Lysates containing 107 CFU of indicated heat-killed bacteria were subjected to 10% sodium dodecyl sulfate–polyacrylamide gel electrophoresis under denaturing conditions. Polyvinylidene fluoride membranes (Millipore, Billerica, MA, USA) were probed overnight with either naive serum (1:100 dilution) or wH- (1:500 dilution), wP- (1:500 dilution), aP- (1:100 dilution) induced serum. A 1:10,000 dilution of goat anti-mouse Ig horseradish peroxidase–conjugated antibodies was used as the detector antibody (35,37). The membrane was visualized with ECL Western Blotting Detection Reagent (Pierce Biotechnology, Rockford, IL, USA).
Statistical Analysis
Mean ± SE values were determined for all appropriate data. Two-tailed, unpaired Student t tests, analysis of variance and Tukey’s simultaneous test in Minitab (www.minitab.com) with similar significance were used to determine statistical significance between groups.
B. holmesii Endemicity in Massachusetts
In 1999, Yih et al. reported an increase in culture-positive B. holmesii cases from 1995 to 1998 (0.2% to 0.6%) (23). Here, collaborating with the same MDPH research team, we report the numbers of B. holmesii culture-positive nasopharyngeal specimens submitted to the MDPH during 2005–2009. Over these 5 years, B. holmesii was isolated from the nasopharyngeal swabs of 41 patients who had similar respiratory symptoms, which is 8 more total cases than observed by Yih et al. during 1994–1998 (33 total cases) (23). At least 2 isolates were recovered each year, and 17 cases were identified in 2006, the highest number observed (Figure 1, panel A). The rate of B. holmesii–positive nasopharyngeal swabs ranged from 0.1% to 0.4%, in line with previous results (Table). Similar to observations made in prior years, 71% of cases occurred in persons 10–19 years of age (Figure 1, panel B), compared with >80% of cases during 1994–1998.
Symptom documentation was obtained for 26 of the 41 cases. All 26 of these patients had a cough; 17 (65%) had a paroxysmal cough, 6 (23%) had post-tussive vomiting, and 4 (15%) had an inspiratory whoop. Nineteen patients (73%) exhibited >1 of these classic symptoms of whooping cough and met the World Health Organization clinical case definition for pertussis (www.who.int/immunization_monitoring/diseases/pertussis_surveillance/en/index.html). No data were collected regarding any previous underlying diseases or potential co-infections among these patients. Although we have no evidence that B. holmesii is the causative agent of these pertussis-like illness, these data suggest that B. holmesii is consistently present in the nasopharynx of a small number of patients who have respiratory infections in Massachusetts.
Phylogenetic Relationships among B. holmesii Isolates
To evaluate the phylogenetic relationships among B. holmesii isolates, an unweighted pair group method using average linkages tree was constructed on the basis of concatenated nucleotide sequences amplified from regions of atpD, rpoB, tuf, and rnpB genes (16). Twelve isolates were obtained from CDC (designated with the letter ‘G’), while the remaining isolates were obtained from the MDPH. Consistent with previous findings (16), all the B. holmesii isolates tested were more closely related to B. hinzii and B. avium than to the classical bordetellae (Figure 2). Although single-nucleotide polymorphisms exist among B. holmesii isolates, pairwise comparisons among the 10 nasopharyngeal isolates from Massachusetts showed >99% sequence identity among 2,958 aligned bases, indicating their close relatedness. By comparison, the CDC isolates were more diverse, with the lowest percent sequence identity 97.5% between CDC strains G7851 and G4363. Moreover, B. holmesii isolates from blood and nasopharyngeal specimens do not cluster separately, which suggests that evolutionary relationship among B. holmesii isolates is not associated with the anatomic site of isolation.
B. holmesii Susceptibility to B. pertussis Vaccine-induced Immunity
Most records for the identified B. holmesii culture-positive cases did not include information regarding vaccination history; however, 16 patients that were culture positive for B. holmesii received >3 doses of pertussis vaccine. The 2010 coverage among children in Massachusetts for DTaP (diphtheria toxoid–tetanus toxoid–acellular pertussis vaccine) 4-dose vaccine was estimated to be >91%, ranking third among the 50 United States (www.cdc.gov/vaccines/stats-surv/imz-coverage.htm). However, adolescents and adults have lower B. pertussis vaccine coverage and waning immunity against B. pertussis. More adolescents and adults compared with children were culture-positive for B. holmesii in Massachusetts, which suggests that B. pertussis vaccine may confer some level of protection against B. holmesii in recently vaccinated persons, similar to the cross-protection against B. bronchiseptica in a murine model of infection (38). Although DTaP was not designed to prevent B. holmesii infections, it is critical to evaluate whether B. pertussis vaccines confer cross-protection against B. holmesii.
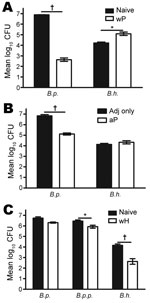
To test whether B. pertussis vaccines provide cross-protection against B. holmesii, we vaccinated C57BL/6 mice with wP or aP. These mice or vaccine-naive mice were then challenged with B. holmesii or B. pertussis and euthanized 3 days later. Because B. holmesii colonization efficiency in the murine respiratory tract is low compared with the classical bordetellae, possibly because of the decreased attachment to mouse respiratory epithelium (A.T. Karanikas and E. T. Harvill, unpub. data), a higher challenge dose was used than for B. pertussis. wP vaccination reduced B. pertussis numbers in the lungs by >99.99% compared with naive mice; however, wP failed to reduce B. holmesii numbers (Figure 3, panel A). In fact, wP vaccination appeared to increase B. holmesii numbers in the lungs compared with naive mice. Although aP-vaccinated mice also reduced B. pertussis numbers in their lungs by ≈98% (Figure 3, panel B), they were not capable of reducing B. holmesii numbers. Together, these data indicate that wP- or aP-induced immunity does not protect against B. holmesii infections.
To determine whether B. holmesii immunization induces protection against itself or cross-protection against B. pertussis and/or B. parapertussis, we vaccinated C57BL/6 mice with heat-killed B. holmesii (wH) and challenged these or naive mice with B. pertussis, B. parapertussis, or B. holmesii; the mice were then euthanized on day 3 postinoculation for bacterial quantification. Similar numbers of B. pertussis were recovered from the lungs of naive and wH-vaccinated mice (Figure 3, panel C), an indication that B. holmesii vaccination failed to reduce B. pertussis numbers within 3 days. However, B. holmesii vaccination reduced the B. parapertussis load by ≈70% in the lungs, which indicates a modest cross-protection provided by B. holmesii against B. parapertussis. Compared with naive mice, mice who received B. holmesii vaccination had a ≈97% reduction of B. holmesii in the lungs (Figure 3, panel A). This experiment indicates that B. holmesii antigens induce efficient protective immunity against B. holmesii but have little effect against B. parapertussis and are even less protective against B. pertussis.
T-cell Responses to wH and wP/aP for B. holmesii
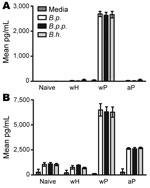
Because wP and aP fail to reduce B. holmesii numbers, we hypothesized that the pertussis vaccines may induce a different T-cell response than the wH vaccine does. To determine whether T-cell responses after vaccination are cross-reactive, splenocytes from naive or wH-, wP-, or aP-vaccinated mice were stimulated with media or heat-killed B. pertussis, B. holmesii, or B. parapertussis for 72 hours. Cell culture supernatant cytokine concentrations of IFN-γ and IL-10, representing Th1 and Treg cytokines, respectively, were determined. wH vaccination did not induce IFN-γ and IL-10 production by splenocytes at greater levels than those produced by naive splenocytes (Figure 4). However, splenocytes from wP-vaccinated mice produced high levels of IFN-γ and IL-10 on stimulation with heat-killed B. pertussis, B. parapertussis, or B. holmesii, which indicates that wP induces a strong cross-reactive T-cell response to B. holmesii. Although splenocytes from aP-vaccinated mice produced little IFN-γ, they produced ≈2,500 pg/mL IL-10 on stimulation with heat-killed B. pertussis, B. holmesii, or B. parapertussis, which indicates cross-reactive T-cell responses following aP vaccination. These data indicate that B. pertussis vaccines can induce cross-reactive T-cell responses to B. holmesii and thus do not explain the lack of cross-protection between these species.
wP/aP-induced Antibodies for B. holmesii
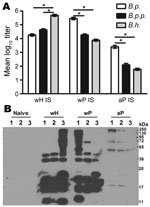
Njamkepo et al. observed that proteins in B. holmesii cell lysates are not recognized by antibodies specific for B. pertussis vaccine antigens (14). To determine whether B. holmesii and B. pertussis vaccine–induced antibodies are cross-reactive, Ig titers of wH-, wP-, or aP-induced serum samples were examined by using ELISA with heat-inactivated bacteria as antigens. The B. holmesii–specific Ig titer of wH-induced serum was >450,000, which is ≈10-fold and 25-fold higher than B. parapertussis– and B. pertussis–specific serum titers, respectively (Figure 5, panel A); this result indicates partial cross-reactivity. B. pertussis–specific Ig titers of wP- and aP-induced serum antibodies were 290,000 and 2,500, respectively. wP- and aP-induced antibodies bind less well to B. parapertussis (Figure 5, panel A) and confer little protection against B. parapertussis in vivo (33). wP- or aP-induced antibodies bound even less well to B. holmesii. A similar trend was observed when live bacteria were used to coat the ELISA plates (data not shown), which rules out the possibility that heat inactivation selectively destroys cross-reactive antigens.
To compare the antigens recognized by serum samples from different groups, Western blot analyses were performed on B. pertussis, B. parapertussis, or B. holmesii lysates probed with naive serum or wH-, wP-, or aP-induced serum. wH-induced serum antibodies recognized B. holmesii antigens of various molecular masses but lacked cross-recognition of some higher molecular mass B. pertussis and B. parapertussis antigens (Figure 5, panel B). The wP- and aP-induced serum antibodies bound less efficiently to B. parapertussis antigens than to B. pertussis antigens, consistent with published data (33). Although wP-induced antibodies recognized some B. holmesii antigens, they lacked recognition of higher molecular mass (>60 kDa) B. holmesii antigens. aP-induced antibodies only poorly recognized a single B. holmesii antigen. Together, these data suggest that antibodies generated following B. pertussis vaccination do not efficiently recognize B. holmesii antigens.
B. holmesii–specific Antibodies and wP-induced Immunity against B. holmesii.
On the basis of our data, we discerned that wP induces sufficient T-cell responses but that antibody responses are not sufficient to confer cross-protection against B. holmesii. If the lack of antibody cross-recognition is the only reason wP and aP vaccines are ineffective against B. holmesii infection, then adding B. holmesii–specific antibodies to wP should render the vaccination effective against B. holmesii.
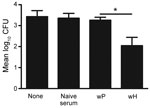
To test this hypothesis, C57BL/6 mice were wP vaccinated or left untreated and later received either naive serum or wP- or wH-vaccinated mouse serum before B. holmesii challenge. B. holmesii lung colonization was determined 3 days postinoculation. Neither naive serum nor wP-induced serum reduced B. holmesii numbers, but vaccinated mice that received B. holmesii–immune serum had substantially lowered B. holmesii numbers in the lungs (Figure 6). The finding indicates that the addition of B. holmesii–specific antibodies to wP increases its efficacy against B. holmesii.
The epidemiologic data collected by the MDPH suggest that B. holmesii is endemic in Massachusetts and is associated with classic whooping cough–like symptoms (Figure 1). Since the establishment of B. holmesii as a species in 1995 (10), infections have been sporadically reported worldwide (6–9,12,13,15,17,18,20,21,23). Increasing awareness of pertussis and improved analytical tools in industrialized countries may have contributed to the increased numbers of reported B. holmesii cases. However, nasopharyngeal B. holmesii specimens submitted to the MDPH each year during 2005–2009 likely represent a small fraction of B. holmesii infections in Massachusetts. Serologic testing and PCR are the dominant diagnostic Bordetella identification assays because of their high sensitivity and time efficiency, but no serologic test or PCR specific for B. holmesii is widely accepted or used by the MDPH. Less than 25% of B. pertussis cases reported in Massachusetts during 1990–2008 were identified by culture, the only test that currently detects B. holmesii. Therefore, larger numbers of B. holmesii cases might be identified if additional B. holmesii–specific serologic or PCR diagnostic tests are used.
When B. holmesii has been clinically identified, several reports have demonstrated little heterogeneity among isolates on the basis of pulsed-field gel electrophoresis banding patterns (21) and 16S rRNA heterogeneity (10,17). By using a sequence-based approach, Diavatopoulos et al. analyzed 7 B. holmesii isolates and observed only 1 nonsynonymous polymorphism among 3,666 bases (16). Using a similar method, we further analyzed 20 isolates from CDC and 10 from MDPH and identified 174 variable sites among the 2,958 aligned bases; this finding suggests more genetic variation among B. holmesii than previously recognized. Similar analyses on a wider range of B. pertussis, B. parapertussis, and B. holmesii isolates could better elucidate the evolutionary history among these human-adapted bordetellae.
Although vaccine studies are better completed in a natural host of the pathogen, the murine model of human-adapted bordetellae infection is well-established. B. pertussis and B. parapertussis murine infections mimic the course of infection and the immune responses in humans (30,39,40), although an animal model of B. holmesii infection has not been previously described. Unlike the classic Bordetella species, B. holmesii colonizes the murine respiratory tract only when relatively large intranasal inoculums are delivered. This model may be improved, for example, by administering antimicrobial drugs or creating transgenic mice. In this study, reproducible colonization of the respiratory tract was achieved, and statistically significant differences were observed after wH vaccination, which suggests that the mechanisms guiding protective immunity in this model can be used to investigate rapid immune-mediated interactions within the respiratory tract.
We determined that B. pertussis vaccines confer little, if any, protection against B. holmesii, even though 16S rRNA comparative analysis of B. holmesii and B. pertussis suggested that B. holmesii is closely related to B. pertussis. Furthermore, antibodies against B. pertussis pertactin, pertussis toxin, fimbriae 2 and 3, adenylate cyclase toxin, and filamentous hemagglutinin recognized few, if any, proteins from multiple B. holmesii isolates (14), which suggests that these proteins are absent from B. holmesii or are antigenically distinct from B. pertussis. Although vaccine-induced T-cell responses are cross-reactive to B. holmesii (Figure 4), B. pertussis vaccine-induced antibodies poorly bind B. holmesii (Figure 5). Furthermore, B. holmesii–specific, but not B. pertussis–specific, antibody administration efficiently decreased B. holmesii numbers in the lungs of vaccinated mice (Figure 6), which suggests that the lack of cross-reactive antibody responses may result in poor cross-protection of B. pertussis vaccines against B. holmesii.
Our data indicate that B. holmesii is circulating in Massachusetts and that B. pertussis vaccination confers little protection against B. holmesii. Careful B. holmesii surveillance is required to better evaluate its prevalence and transmission. B. holmesii genome sequencing may identify novel virulence determinants to explain its emergence in the human population and guide effective vaccine design.
Dr Zhang is a postdoctoral fellow at Department of Microbiology and Immunobiology, Harvard Medical School, Cambridge, Massachusetts, USA. Her research interest is host and bacterial pathogen interaction.
Acknowledgment
We thank Linda Han and Noelle Cocoros for providing the epidemiological data used in this study and the Centers for Disease Control and Prevention and the Massachusetts Department of Public Health for providing the B. holmesii isolates.
References
- Hoppe JE. Update on respiratory infection caused by Bordetella parapertussis. Pediatr Infect Dis J. 1999;18:375–81. DOIPubMedGoogle Scholar
- Mattoo S, Cherry JD. Molecular pathogenesis, epidemiology, and clinical manifestations of respiratory infections due to Bordetella pertussis and other Bordetella subspecies. Clin Microbiol Rev. 2005;18:326–82. DOIPubMedGoogle Scholar
- Cherry JD, Brunell PA, Golden GS, Karson DT. Report of the task force on pertussis and pertussis immunization-1988. Pediatrics. 1988;81:939.
- Centers for Disease Control and Prevention. Pertussis–United States, 1997–2000. JAMA. 2002;287:977–9. DOIPubMedGoogle Scholar
- Celentano LPMM, Paramatti D, Salmaso S, Tozzi AE. EUVAC-NET Group. Resurgence of pertussis in Europe. Pediatr Infect Dis J. 2005;24:761–5. DOIPubMedGoogle Scholar
- Kamiya H, Otsuka N, Ando Y, Odaira F, Yoshino S, Kawano K, Transmission of Bordetella holmesii during pertussis outbreak, Japan. Emerg Infect Dis. 2012;18:1166–9. DOIPubMedGoogle Scholar
- Miranda C, Porte L, Garcia P. Bordetella holmesii in nasopharyngeal samples from Chilean patients with suspected Bordetella pertussis infection. J Clin Microbiol. 2012;50:1505, author reply 6. DOIPubMedGoogle Scholar
- Mooi FR, Bruisten S, Linde I, Reubsaet F, Heuvelman K, van der Lee S, Characterization of Bordetella holmesii isolates from patients with pertussis-like illness in The Netherlands. FEMS Immunol Med Microbiol. 2012;64:289–91. DOIPubMedGoogle Scholar
- Njamkepo E, Bonacorsi S, Debruyne M, Gibaud SA, Guillot S, Guiso N. Significant finding of Bordetella holmesii DNA in nasopharyngeal samples from French patients with suspected pertussis. J Clin Microbiol. 2011;49:4347–8. DOIPubMedGoogle Scholar
- Weyant RS, Hollis DG, Weaver RE, Amin MF, Steigerwalt AG, O’Connor SP, Bordetella holmesii sp. nov., a new gram-negative species associated with septicemia. J Clin Microbiol. 1995;33:1–7.PubMedGoogle Scholar
- Guthrie JL, Robertson AV, Tang P, Jamieson F, Drews SJ. Novel duplex real-time PCR assay detects Bordetella holmesii in specimens from patients with pertussis-like symptoms in Ontario, Canada. J Clin Microbiol. 2010;48:1435–7. DOIPubMedGoogle Scholar
- Russell FM, Davis JM, Whipp MJ, Janssen PH, Ward PB, Vyas JR, Severe Bordetella holmesii infection in a previously healthy adolescent confirmed by gene sequence analysis. Clin Infect Dis. 2001;33:129–30. DOIPubMedGoogle Scholar
- Dörbecker C, Licht C, Korber F, Plum G, Haefs C, Hoppe B, Community-acquired pneumonia due to Bordetella holmesii in a patient with frequently relapsing nephrotic syndrome. J Infect. 2007;54:e203–5. DOIPubMedGoogle Scholar
- Njamkepo E, Delisle F, Hagege I, Gerbaud G, Guiso N. Bordetella holmesii isolated from a patient with sickle cell anemia: analysis and comparison with other Bordetella holmesii isolates. Clin Microbiol Infect. 2000;6:131–6. DOIPubMedGoogle Scholar
- Greig JR, Gunda SS, Kwan JTC. Bordetella holmesii bacteraemia in an individual on haemodialysis. Scand J Infect Dis. 2001;33:716–7. DOIPubMedGoogle Scholar
- Diavatopoulos DA, Cummings CA, van der Heide HG, van Gent M, Liew S, Relman DA, Characterization of a highly conserved island in the otherwise divergent Bordetella holmesii and Bordetella pertussis genomes. J Bacteriol. 2006;188:8385–94. DOIPubMedGoogle Scholar
- Shepard CW, Daneshvar MI, Kaiser RM, Ashford DA, Lonsway D, Patel JB, Bordetella holmesii bacteremia: a newly recognized clinical entity among asplenic patients. Clin Infect Dis. 2004;38:799–804. DOIPubMedGoogle Scholar
- McCavit TL, Grube S, Revell P, Quinn CT. Bordetella holmesii bacteremia in sickle cell disease. Pediatr Blood Cancer. 2008;51:814–6. DOIPubMedGoogle Scholar
- Monnier S, Therby A, Couzon B, Doucet-Populaire F, Greder-Belan A. Bordetella holmesii bacteremia in a 26-year-old patient with sickle cell disease [article in French]. Med Mal Infect. Epub 2009 Jul 7.PubMedGoogle Scholar
- Lindquist SW, Weber DJ, Mangum ME, Hollis DG, Jordan J. Bordetella holmesii sepsis in an asplenic adolescent. Pediatr Infect Dis J. 1995;14:813–5. DOIPubMedGoogle Scholar
- Mazengia E, Silva EA, Peppe JA, Timperi R, George H. Recovery of Bordetella holmesii from patients with pertussis-like symptoms: use of pulsed-field gel electrophoresis to characterize circulating strains. J Clin Microbiol. 2000;38:2330–3.PubMedGoogle Scholar
- Tang YW, Hopkins MK, Kolbert CP, Hartley PA, Severance PJ, Persing DH. Bordetella holmesii–like organisms associated with septicemia, endocarditis, and respiratory failure. Clin Infect Dis. 1998;26:389–92. DOIPubMedGoogle Scholar
- Yih WK, Silva EA, Ida J, Harrington N, Lett SM, George H. Bordetella holmesii–like organisms isolated from Massachusetts patients with pertussis-like symptoms. Emerg Infect Dis. 1999;5:441–3. DOIPubMedGoogle Scholar
- Stibitz S, Yang MS. Subcellular localization and immunological detection of proteins encoded by the vir locus of Bordetella pertussis. J Bacteriol. 1991;173:4288–96.PubMedGoogle Scholar
- Preston A, Allen AG, Cadisch J, Thomas R, Stevens K, Churcher CM, Genetic basis for lipopolysaccharide O-antigen biosynthesis in bordetellae. Infect Immun. 1999;67:3763–7.PubMedGoogle Scholar
- von Koenig CH, Tacken A, Finger H. Use of supplemented Stainer-Scholte broth for the isolation of Bordetella pertussis from clinical material. J Clin Microbiol. 1988;26:2558–60.PubMedGoogle Scholar
- Stainer DW, Scholte MJ. A simple chemically defined medium for the production of phase I Bordetella pertussis. J Gen Microbiol. 1970;63:211–20. DOIPubMedGoogle Scholar
- Gopinathan L, Kirimanjeswara GS, Wolfe DN, Kelley ML, Harvill ET. Different mechanisms of vaccine-induced and infection-induced immunity to Bordetella bronchiseptica. Microbes Infect. 2007;9:442–8. DOIPubMedGoogle Scholar
- Kirimanjeswara GS, Agosto LM, Kennett MJ, Bjornstad ON, Harvill ET. Pertussis toxin inhibits neutrophil recruitment to delay antibody-mediated clearance of Bordetella pertussis. J Clin Invest. 2005;115:3594–601. DOIPubMedGoogle Scholar
- Kirimanjeswara GS, Mann PB, Harvill ET. Role of antibodies in immunity to Bordetella infections. Infect Immun. 2003;71:1719–24. DOIPubMedGoogle Scholar
- Pilione MRHE. The Bordetella bronchiseptica type III secretion system inhibits gamma interferon production that is required for efficient antibody-mediated bacterial clearance. Infect Immun. 2006;74:1043–9. DOIPubMedGoogle Scholar
- Pishko EJ, Kirimanjeswara GS, Pilione MR, Gopinathan L, Kennett MJ, Harvill ET. Antibody-mediated bacterial clearance from the lower respiratory tract of mice requires complement component C3. Eur J Immunol. 2004;34:184–93. DOIPubMedGoogle Scholar
- Zhang X, Rodriguez ME, Harvill ET. O antigen allows B. parapertussis to evade B. pertussis vaccine-induced immunity by blocking binding and functions of cross-reactive antibodies. PLoS ONE. 2009;4:e6989. DOIPubMedGoogle Scholar
- Wolfe DNKG, Harvill ET. Clearance of Bordetella parapertussis from the lower respiratory tract requires humoral and cellular immunity. Infect Immun. 2005;73:6508–13. DOIPubMedGoogle Scholar
- Wolfe DN, Goebel EM, Bjornstad ON, Restif O, Harvill ET. The O antigen enables Bordetella parapertussis to avoid Bordetella pertussis–induced immunity. Infect Immun. 2007;75:4972–9. DOIPubMedGoogle Scholar
- Myc A, Buck J, Gonin J, Reynolds B, Hammerling U, Emanuel D. The level of lipopolysaccharide-binding protein is significantly increased in plasma in patients with the systemic inflammatory response syndrome. Clin Diagn Lab Immunol. 1997;4:113–6.PubMedGoogle Scholar
- Wolfe DN, Kirimanjeswara GS, Goebel EM, Harvill ET. Comparative role of immunoglobulin A in protective immunity against the Bordetellae. Infect Immun. 2007;75:4416–22. DOIPubMedGoogle Scholar
- Goebel EM, Zhang X, Harvill ET. Bordetella pertussis infection or vaccination substantially protects mice against B. bronchiseptica infection. PLoS ONE. 2009;4:e6778. DOIPubMedGoogle Scholar
- Harvill ET, Cotter PA, Miller JF. Pregenomic comparative analysis between Bordetella bronchiseptica RB50 and Bordetella pertussis Tohama I in murine models of respiratory tract infection. Infect Immun. 1999;67:6109–18.PubMedGoogle Scholar
Figures
Table
Cite This ArticleTable of Contents – Volume 18, Number 11—November 2012
| EID Search Options |
|---|
|
|
|
|
|
|


Please use the form below to submit correspondence to the authors or contact them at the following address:
Eric T. Harvill, Department of Veterinary and Biomedical Sciences, The Pennsylvania State University, 115 Henning Bldg, University Park, PA 16802, USA
Top