Volume 20, Number 12—December 2014
Letter
Novel Divergent Rhabdovirus in Feces of Red Fox, Spain
Cite This Article
Citation for Media
To the Editor: Rhabdoviruses (family Rhabdoviridae) are enveloped single-stranded negative-sense RNA viruses belonging to the Mononegavirales order. The International Committee on Taxonomy of Viruses recognizes 11 genera (Cytorhabdovirus, Ephemerovirus, Lyssavirus, Novirhabdovirus, Nucleorhabdovirus, Perhabdovirus, Sigmavirus, Sprivivirus, Tibrovirus, Tupavirus, Vesiculovirus) (1). In addition, many recently described rhabdoviruses remain unassigned. Rhabdoviruses contain 5 major genes, encoding for nucleoprotein (N), phosphoprotein (P), matrix (M), glycoprotein (G), and RNA-dependent RNA polymerase (L). The Rhabdoviridae family includes pathogens of various animal species, humans, and plants. Viruses of the genus Lyssavirus are the most relevant to public health because they can cause rabies. Bats are the driving force within this genus; foxes and various other species of wild carnivores also can be infected with lyssaviruses and transmit them to humans and dogs (2).
During a viral metagenomic survey, conducted as described previously (3), of fecal samples collected from 4 red foxes (Vulpes vulpes) that were found dead in Álava, Basque Country, Spain, we identified the complete coding sequence and the partial leader and trailer sequence of a novel rhabdovirus, tentatively called red fox fecal rhabdovirus (RFFRV; 15,541 nt, GenBank accession no. KF823814; Technical Appendix) by mapping 8,287 of the 56,519 sequence reads in the sample of a red fox. A proportion of obtained reads contained sequences that were >99% identical to mitochondrial DNA of V. vulpes, which confirmed that the sample was collected from a red fox.
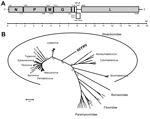
The obtained sequence of RFFRV was partially confirmed by specific primers and Sanger sequencing of PCR amplicons. Five major and 3 minor open reading frames (ORFs) were identified that had a genome organization similar to that of other rhabdoviruses (Figure, panel A). No significant hits were obtained by BLAST analysis (http://blast.ncbi.nlm.gov/Blast.cgi) of N, P, M, and G nucleotide and amino acid sequences, which was reported previously for novel divergent rhabdoviruses (4).
Predicted N, P, and M genes of RFFRV consist of 1,629, 2,490, and 813 nt, respectively, encoding for 543, 830, and 271 aa (Technical Appendix Table 1). In addition to the absence of significant hits observed by BLAST analysis, no significant sequence homology was observed with known rhabdovirus proteins in pairwise alignments. Furthermore, no conserved motifs were detected in N, P, and M genes of RFFRV that are commonly observed in rhabdoviruses. However, intergenic regions between all major ORFs contained relatively conserved motifs that could be transcription termination/polyadenylation sequences (A/U) CU7, similar to other rhabdoviruses (5). Adjacent to this termination signal was a stretch of conserved nucleotides that might function as a transcription initiation signal Technical Appendix Table 1).
The amino acid sequence of the G protein consisted of 669 aa and contained an N terminal signal peptide (1-MYHLIVLLVMLGQRAVA-17), a noncytoplasmic domain (aa 18–646), a transmembrane domain (647-ITAILMPLLSLAVVVGIIMCC-667), and a cytoplasmic tail of 2 aa, similar to other rhabdovirus G proteins as predicted by using Phobius and TMHMM (http://www.cbs.dtu.dk/services/TMHMM) (6,7). We predicted 3 potential glycosylation sites in the ectodomain at positions 38–40 (NKT), 554–556 (NAS), and 592–594 (NIS) using NetNGlyc 1.0 (http://www.cbs.dtu.dk/services/NetNGlyc).
Between the G and L genes, a complex intergenic region was present that contained 3 ORFs of 246 nt (7,413–7,658 aa), 231 nt (7,716–7,946 aa), and 459 nt (7,893–8,355 aa), of which 2 were overlapping frames (U1–3). Additional ORFs between G and L genes were detected previously in other rhabdoviruses (8,9). We detected transmembrane domains in the amino acid sequences of all 3 additional ORFs, suggesting they might act as viroporin (8,9).
The L gene of RFFRV contained 6,591 nt (2,197 aa). We detected several conserved domains and motifs, including RNA-dependent RNA polymerase, mRNA-capping region, mRNA capping enzyme, and virus-capping methyltransferase. Alignment of the deduced amino acid sequence of the L gene with the L gene of various other viruses belonging to the Mononegavirales order by using MAFFT version 7 (http://mafft.cbrc.jp/alignment/software/) and subsequent phylogenetic reconstruction by using a maximum-likelihood tree (WAG+F+I+G model with 100 bootstrap replicates in MEGA5 [http://www.megasoftware.net]) suggested that this virus belongs to a novel genus of the Rhabdoviridiae family. In addition, pairwise identities of the deduced amino acid sequence of the L gene of RFFRV with that of other rhabdoviruses of the Rhabdoviridae family were only <35% (Technical Appendix Table 2).
Because the fox was found dead and no tissue samples were collected, whether RFFRV played a role in the animal’s death is unknown. In addition, multiple attempts to isolate this virus on various cell lines of eukaryotes (Vero E6, MDCK, CRFK, N2a, and BHK cells, primary fox kidney cells) failed because of the absence of cytopathic effects and viral replication by quantitative reverse transcription PCR, despite a high number of reads in the original sample. The fox might have acquired the virus through spillover from a small prey, such as a bat, and additional studies are required to elucidate the prevalence, original host, and pathogenic potential of this novel virus.
Acknowledgments
We thank all researchers and institutions for their invaluable help during sampling and for providing the specimens used in this study, especially Patricia Lizarraga, Laura Elorza, Ricardo Gutierrez, and Luis Javier Chueca.
This work was funded by the European Community’s Seventh Framework Program (FP7/2007–2013) under the project “European Management Platform for Emerging and Re-emerging Infectious Disease Entities” European Community grant agreement no. 223498; the Virgo Consortium; and the Niedersachsen-Research Network on Neuroinfectiology of the Ministry of Science and Culture of Lower Saxony, Germany. In addition, this research was funded partially by the Basque Government through the research group on “Systematics, Biogeography and Population Dynamics” (ref. IT317-10; GIC10/76).
A.R.-G. holds a postdoctoral fellowship awarded by the Department of Education, Universities and Research of the Basque Government (ref. DKR-2012-64) and was awarded a short-visit research grant from the ConGenOmics Research networking program of the European Science Foundation to visit the Department of Viroscience, Erasmus Medical Centre and develop the current research project.
References
- International Committee on Taxonomy of Viruses. Virus taxonomy: 2013 release [cited 2014 Feb 9]. http://www.ictvonline.org/virusTaxonomy.asp
- Matha IS, Salunke SR. Immunogenicity of purified Vero cell rabies vaccine used in the treatment of fox-bite victims in India. Clin Infect Dis. 2005;40:611–3. DOIPubMedGoogle Scholar
- Bodewes R, Ruiz-Gonzalez A, Schapendonk CM, van den Brand JM, Osterhaus AD, Smits SL. Viral metagenomic analysis of feces of wild small carnivores. Virol J. 2014;11:89. DOIPubMedGoogle Scholar
- Palacios G, Forrester NL, Savji N, Travassos da Rosa AP, Guzman H, Detoy K, Characterization of Farmington virus, a novel virus from birds that is distantly related to members of the family Rhabdoviridae. Virol J. 2013;10:219. DOIPubMedGoogle Scholar
- Albertini AA, Ruigrok RW, Blondel D. Rabies virus transcription and replication. Adv Virus Res. 2011;79:1–22. DOIPubMedGoogle Scholar
- Käll L, Krogh A. Sonnhammer EL. A combined transmembrane topology and signal peptide prediction method. J Mol Biol. 2004;338:1027–36. DOIPubMedGoogle Scholar
- Gubala AJ, Proll DF, Barnard RT, Cowled CJ, Crameri SG, Hyatt AD, Genomic characterisation of Wongabel virus reveals novel genes within the Rhabdoviridae. Virology. 2008;376:13–23. DOIPubMedGoogle Scholar
- McWilliam SM, Kongsuwan K, Cowley JA, Byrne KA, Walker PJ. Genome organization and transcription strategy in the complex GNS-L intergenic region of bovine ephemeral fever rhabdovirus. J Gen Virol. 1997;78:1309–17 .PubMedGoogle Scholar
Figure
Cite This Article1These authors contributed equally to this article.
Related Links
Table of Contents – Volume 20, Number 12—December 2014
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Rogier Bodewes, Erasmus Medical Centre, Department of Viroscience Dr. Molewaterplein 50, 3015GE Rotterdam, the Netherlands
Top