Volume 20, Number 3—March 2014
Dispatch
Nontoxigenic tox-bearing Corynebacterium ulcerans Infection among Game Animals, Germany
Cite This Article
Citation for Media
Abstract
Corynebacterium ulcerans may cause diphtheria in humans and caseous lymphadenitis in animals. We isolated nontoxigenic tox-bearing C. ulcerans from 13 game animals in Germany. Our results indicate a role for game animals as reservoirs for zoonotic C. ulcerans.
The Corynebacterium species C. diphtheriae, C. ulcerans, and C. pseudotuberculosis form the C. diphtheriae group, as shown by 16S rRNA gene sequence analysis (1). Strains of this group carrying lysogenic β-corynephages might produce the tox-encoded diphtheria toxin (DT) (2). Moreover, C. ulcerans and C. pseudotuberculosis may produce phospholipase D, the major virulence factor involved in caseous lymphadenitis, which is a disease that mainly affects sheep, goats, and horses (3).
From a public health perspective, diphtheria is the most critical human disease attributed to coryneform bacteria (3). In recent years, cases of diphtheria caused by C. ulcerans have outnumbered those caused by C. diphtheriae (4). C. diphtheriae carriage is nearly exclusively restricted to humans; C. ulcerans is a zoonotic pathogen and has been found in various animal species that have contact with humans (5). C. ulcerans is most closely related to C. pseudotuberculosis, and distinction between these species is often difficult when using standard bacteriological methods (5). The aim of this study was to comprehensively characterize 13 C. ulcerans strains isolated from game animals in Germany.
Strains of C. ulcerans were isolated during routine bacteriological investigations in conjunction with necropsies of wild animals that were found dead or that had suspicious lesions during 1997–2013. Isolates of coryneform bacteria were subjected to conventional biochemical tests (3), and were evaluated after prolonged incubation at 37°C for as long as 14 days. For further characterization, commercial tests API Coryne and VITEK2-compact with cards for coryneform bacteria and corynebacteria and anaerobes (bioMérieux, Nürtingen, Germany) were used according to the manufacturer’s instructions.
We conducted the reverse CAMP test by using Staphylococcus aureus American Type Culture Collection (ATCC [Manassas, VA, USA]) 25923 and the CAMP test by using Rhodococcus equi ATCC 33701 according to standard procedures on Columbia sheep blood agar (Oxoid, Wesel, Germany) (3). We determined DT production using a modified Elek test (6); we used C. diphtheriae NCTC 10648 and C. diphtheriae NCTC 10356 as positive and negative controls, respectively; and performed a cytotoxicity assay using Vero cells (7). The rpoB and tox genes were partially amplified by using primer pairs C2700F/C3130R and DT1/DT2, respectively, as described (5).
PCR products were purified for sequence analysis by using the Double Pure Combi Kit (Bio&SELL, Nürnberg, Germany). Both strands of the rpoB and tox PCR products were sequenced by Microsynth (Balgach, Switzerland) by using the amplification primers. Sequence analysis was performed by using the BLAST (http://blast.ncbi.nlm.nih.gov/Blast.cgi) sequence analysis tool. Additionally, coryneform isolates in which C. ulcerans was suspected were analyzed by using Matrix-assisted laser desorption-ionization time-of-flight mass spectrometry (MALDI-TOF MS) and by using Biotyper version 3.3.1.0 (BrukerBiotyper; BrukerDaltonics, Bremen, Germany). The database used (DB 4613) comprised spectra from 71 Corynebacterium species including C. diphtheriae, C. ulcerans, and C. pseudotuberculosis. For Fourier-transform infrared (FT-IR) spectroscopy, bacterial isolates were harvested and prepared as described (5). IR spectra were recorded by using an FT-IR spectrometer (Tensor 27 with High Throughput Screening eXTension HTS-XT module) and OPUS software version 4.2 (BrukerOptics, Ettlingen, Germany). IR spectra of isolates from game animals and selected C. ulcerans and C. pseudotuberculosis strains were compared by cluster analysis by using the second derivation of vector normalized spectra (8). The dendrogram obtained depicts the arrangement of isolates in groups according to their spectral differences (Figure 1).
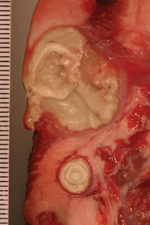
In total, 13 strains of C. ulcerans were isolated from 12 wild boars and 1 roe deer in 4 states of Germany (Table 1). The bacteria grew from >1 delimited pseudotuberculosis-like caseous abscess, arranged in concentric layers and ranging from 0.1 to 10 cm in diameter (Figure 2). All strains had positive reactions by using a traditional CAMP test inoculated with R. equi and a reverse CAMP test inoculated with S. aureus, indicating phospholipase D activity (3).
Conventional biochemical tests showed congruent results for catalase activity, urea hydrolysis, and glucose acidification (positive) and for esculin hydrolysis and nitrate reduction (negative). Additional reactions and API and VITEK test results are shown in Table 2. All isolates were nontoxigenic tox-bearing (NTTB) strains as shown by positive tox-PCR, and negative Elek test and Vero cell cytotoxicity results. Partial rpoB and partial tox sequences for all 13 isolates were identical to those submitted to GenBank for C. ulcerans strain CVUAS 4292 (accession nos. GU818735 and GU818742, respectively [5]).
By using MALDI-TOF MS, all isolates were identified to the species level as C. ulcerans because they had score levels of 2.0–2.2. The comparison of the IR spectra of the 13 strains from game animals with a collection of reference strains showed a clear separation in 2 main branches for the 2 species C. pseudotuberculosis and C. ulcerans (Figure 1). Inside the C. ulcerans branch, all isolates from game animals clustered compactly together and were closely adjacent to a group of spectra formed by reference strains from humans.
With respect to its zoonotic potential, C. ulcerans is one of the most notable members of the genus and was referred to as an emerging pathogen in 2011 (9). Numerous reports state there is zoonotic potential for contact with companion or farm animals, but proven transmission of tox-positive C. ulcerans strains is documented for only 4 cases, involving 2 dogs, 1 cat, and 1 pig (8).
Limited information is available concerning C. ulcerans infection in wild animals. To our knowledge, 3 reports regarding tox-positive C. ulcerans infection in wildlife have been published: 1 involved 2 European otters from 2 widely separated regions within the United Kingdom (10), and the other 2 reports described NTTB strains in 2 wild boars in 1 report and 1 roe deer in the other report; these 3 cases were in the same area of Germany (5,8). An additional report on C. ulcerans with unknown toxigenicity in wildlife pertains to an outbreak among 350 squirrels from Canada, 63 of which had clinical disease (11).
Here, we provide comprehensive data on 13 NTTB C. ulcerans strains from game animals in Germany. The finding of infected game in the center of Middle Europe suggests an even wider occurrence and distribution in other European countries. Misdiagnoses of C. ulcerans isolates as C. pseudotuberculosis in the past because of similar pathology and similar phenotype cannot be excluded. Our finding of C. ulcerans in a wild boar specimen from 1997 could indicate that this pathogen has not only recently infected wildlife.
As also shown in this study, biochemical differentiation between C. ulcerans and C. pseudotuberculosis might be problematic, and basic conventional tests may not properly discriminate between the 2 species (3). By using the standardized systems API Coryne and VITEK2-compact for coryneform bacteria, erroneous identification was made of most isolates (10 and 11 cases, respectively) from game animals as C. pseudotuberculosis. For correct understanding of epidemiology and host range and for unequivocal determination of the involved pathogen to species level, additional methods such as FT-IR and MALDI-TOF MS or DNA sequencing should be used. Because partial rpoB sequencing is more discriminatory than 16S rDNA sequencing, a cutoff value of ≤95% similarity proved suitable for species identification within Corynebacterium (12) and also clearly enabled species identification in this study. Furthermore, partial tox- and rpoB-gene sequencing demonstrated a very close relationship between the 13 strains because no variations in these sequences were found (8).
Concerning the zoonotic potential for C. ulcerans strains from wildlife, there is no information available. With respect to wild boars infected with C. ulcerans, however, it is noteworthy that 3 diphtheria cases occurred in humans who had occupational contact with pigs (13,14).
Lack of DT expression in tox-positive strains has been described (7). Nevertheless, it can be expected that DT-producing C. ulcerans strains might occur in game animals, providing a reservoir for this microorganism. Because the C. diphtheriae and C. ulcerans DT sequences are quite similar, it might be reasonable to offer diphtheria toxoid vaccination to persons who have direct contact with game animals to prevent diphtheria-like illness caused by tox-positive C. ulcerans (4).
Dr Eisenberg is a specialist in microbiology at the Hessian state laboratory in Gießen and team supervisor of the bacteriology department. He has special interests in infectious zoo and wildlife diseases and zoonoses.
Acknowledgments
We thank Asmahan Omar, Anna Mohr, Wolfgang Schmidt, Barbara Depner, Anna Katharina Schmid, and Mandy Hailer for excellent technical assistance and Anja Berger, Heribert Bischoff, and Regina Konrad for continuous support.
The Consiliary Laboratory on Diphtheria received grants for clinical research from the Robert Koch-Institute.
References
- Pascual C, Lawson PA, Farrow JA, Gimenez MN, Collins MD. Phylogenetic analysis of the genus Corynebacterium based on 16S rRNA gene sequences. Int J Syst Bacteriol. 1995;45:724–8. DOIPubMedGoogle Scholar
- Wong TP, Groman N. Production of diphtheria toxin by selected isolates of Corynebacterium ulcerans and Corynebacterium pseudotuberculosis. Infect Immun. 1984;43:1114–6 .PubMedGoogle Scholar
- Funke G, von Graevenitz A, Clarridge JE III, Bernard KA. Clinical microbiology of coryneform bacteria. Clin Microbiol Rev. 1997;10:125–59 .PubMedGoogle Scholar
- Wagner KS, White JM, Crowcroft NS, De Martin S, Mann G, Efstratiou A. Diphtheria in the United Kingdom, 1986–2008: the increasing role of Corynebacterium ulcerans. Epidemiol Infect. 2010;138:1519–30 . DOIPubMedGoogle Scholar
- Contzen M, Sting R, Blazey B, Rau J. Corynebacterium ulcerans from diseased wild boars. Zoonoses Public Health. 2011;58:479–88 . DOIPubMedGoogle Scholar
- Engler KH, Glushkevich T, Mazurova IK, George RC, Efstratiou A. A modified Elek test for detection of toxigenic corynebacteria in the diagnostic laboratory. J Clin Microbiol. 1997;35:495–8 .PubMedGoogle Scholar
- Sing A, Hogardt M, Bierschenk S, Heesemann J. Detection of differences in the nucleotide and amino acid sequences of diphtheria toxin from Corynebacterium diphtheriae and Corynebacterium ulcerans causing extrapharyngeal infections. J Clin Microbiol. 2003;41:4848–51 . DOIPubMedGoogle Scholar
- Rau J, Blazey B, Contzen M, Sting R. Corynebacterium ulcerans infection in roe deer (Capreolus capreolus). Berl Munch Tierarztl Wochenschr. 2012;125:159–62 .PubMedGoogle Scholar
- Sing A, Berger A, Schneider-Brachert W, Holzmann T, Reischl U. Rapid detection and molecular differentiation of toxigenic Corynebacterium diphtheriae and Corynebacterium ulcerans strains by LightCycler PCR. J Clin Microbiol. 2011;49:2485–9. DOIPubMedGoogle Scholar
- Foster G, Patterson T, Howie F, Simpson V, Davison N, Efstratiou A, Corynebacterium ulcerans in free-ranging otters. Vet Rec. 2002;150:524 .PubMedGoogle Scholar
- Olson ME, Goemans I, Bolingbroke D, Lundberg S. Gangrenous dermatitis caused by Corynebacterium ulcerans in Richardson ground squirrels. J Am Vet Med Assoc. 1988;193:367–8 .PubMedGoogle Scholar
- Khamis A, Raoult D, La Scola B. Comparison between rpoB and 16S rRNA gene sequencing for molecular identification of 168 clinical isolates of Corynebacterium. J Clin Microbiol. 2005;43:1934–6. DOIPubMedGoogle Scholar
- Schuhegger R, Schoerner C, Dlugaiczyk J, Lichtenfeld I, Trouillier A, Zeller-Peronnet V, Pigs as source for toxigenic Corynebacterium ulcerans. Emerg Infect Dis. 2009;15:1314–5. DOIPubMedGoogle Scholar
- Berger A, Boschert V, Konrad R, Schmidt-Wieland T, Hörmansdorfer S, Eddicks M, Two cases of cutaneous diphtheria associated with occupational pig contact in Germany. Zoonoses Public Health. 2013;60:539–42 . DOIPubMedGoogle Scholar
Figures
Tables
Cite This ArticleReference has only first page number. Please provide the last page number if article is longer than one page. (in reference 10 "Foster, Patterson, Howie, Simpson, Davison, Efstratiou, et al., 2002").
Table of Contents – Volume 20, Number 3—March 2014
| EID Search Options |
|---|
|
|
|
|
|
|

Please use the form below to submit correspondence to the authors or contact them at the following address:
Tobias Eisenberg, Landesbetrieb Hessisches Landeslabor, Abteilung Veterinärmedizin, Schubertstr. 60/ Haus 13, 35392 Gießen, GermanyTobias Eisenberg, Landesbetrieb Hessisches Landeslabor, Abteilung Veterinärmedizin, Schubertstr. 60/ Haus 13, 35392 Gießen, GermanyTobias Eisenberg, Landesbetrieb Hessisches Landeslabor, Abteilung Veterinärmedizin, Schubertstr. 60/ Haus 13, 35392 Gießen, Germany
Top