Volume 22, Number 1—January 2016
Letter
Widespread Bat White-Nose Syndrome Fungus, Northeastern China
Cite This Article
Citation for Media
To the Editor: Emerging infectious diseases have caused catastrophic declines in wildlife populations, and the introductions of many pathogen have been linked to increases in global trade and travel (1). Mapping the distribution of pathogens is necessary to identify species and populations at risk and identify sources of pathogen spillover and introduction. Once pathogen distributions are known, management actions can be taken to reduce the risk for future global spread (2).
Bats with symptoms of white-nose syndrome (WNS) were first detected in the United States in 2006, and the disease has subsequently caused precipitous declines in temperate bat populations across eastern North America (3,4). Pseudogymnoascus destructans, the causative agent of WNS, is a cold-growing fungus that infects bats’ skin during hibernation, leading to more frequent arousals from torpor and death (3). P. destructans is widespread throughout Europe (5), but, to our knowledge, its presence in Asia has not been documented.
We sampled bats and hibernacula surfaces (cave walls and ceilings) across northeastern China during 2 visits (June–July 2014 and March 2015) using a previously described swab-sampling technique (6). Bats were captured inside caves and at their entrances. DNA was extracted from samples by using a modified QIAGEN DNeasy blood and tissue kit (QIAGEN, Valencia, CA, USA) and tested in duplicate for the presence of P. destructans with a quantitative real-time PCR (qPCR) (6,7).
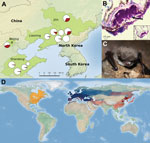
In the summer of 2014 and winter of 2015, we collected 385 samples from hibernacula surfaces at 12 sites in 3 provinces and 1 municipality (Figure, panel A) and 215 samples from 9 species of bats at 10 sites (summer: Rhinolophus ferrumequinum, Rhinolophus pusillus, Myotis adversus, Myotis macrodactylus, Myotis pilosus, Myotis chinensis, Murina usseriensis; winter: R. ferrumequinum, Murina leucogaster, Myotis petax). During the summer, P. destructans was widely distributed across the study region with positive samples (determined on the basis of qPCR results) obtained from cave surfaces at 9 of 12 sites and from bats at 2 of the 9 sites where bats were sampled (Figure, panel A).
Prevalence of P. destructans was low during summer in the environment (mean prevalence across sites 0.06 ± 0.03) and in bats. Bats of 3 species tested positive for P. destructans in the summer: M. macrodactylus (1/10), M. chinensis (1/1), and M. ussuriensis (1/1). P. destructans was not detected in bats of 4 other species, of which >20 individual animals of each species were sampled (R. ferrumequinum, R. pusillus, M. pilosus, and M. adversus). The low prevalence of P. destructans in bats and on hibernacula surfaces in China during the summer was similar to comparable results from studies in North America (6).
In winter, prevalence at the 2 sites we revisited was much higher; 75% of 85 samples from 3 species tested positive, including samples from 16/17 M. petax bats. We also detected P. destructans in bats from 2 additional species (R. ferrumequinum [11/19 bats] and M. leucogaster [11/16 bats]).
In addition, during March 2015, we observed visual evidence of P. destructans in bats (M. petax; Figure, panel C) and obtained 2 fungal cultures from swab specimens taken from these bats. To isolate P. destructans from these samples, we plated swab specimens from visibly infected bats on Sabouraud dextrose agar at 10°C. We identified potential P. destructans isolates on the basis of morphologic characteristics. DNA was then extracted from 2 suspected fungal cultures and tested for P. destructans by qPCR, as previously described.
To further confirm the presence of P. destructans, we prepared the fungal isolates for Sanger sequencing (Technical Appendix). The 600-nt amplification products from these 2 isolates were sequenced and found to be 100% identical to the P. destructans rRNA gene region targeted for amplification. In addition, using BLAST (http://www.ncbi.nlm.nih.gov/blast.cgi), we found that sequences were a 100% match with isolates from Europe (GenBank accession no. GQ489024) and North America (GenBank accession no. EU884924). This result confirms that the same species of fungus occurs on all 3 continents. We also obtained wing biopsy punches from these bats and found lesions characteristic of WNS by histopathologic examination (Figure, panel B; Technical Appendix).
The occurrence of P. destructans at most sites sampled indicates that this pathogen is widespread in eastern Asia (Figure, panel A). The presence of P. destructans in bats from 6 species in China and on bats in 13 species in Europe (8) confirms the generalist nature of this fungus and suggests that it may occur throughout Eurasia (Figure, panel D).
Decontamination and restrictions on the use of equipment that has been used in caves in Asia would help reduce the probability of introducing P. destructans to uninfected bat populations (e.g., western North America, New Zealand, southern Australia, and temperate areas of South America). These measures would also reduce the risk of introducing new strains of P. destructans to regions where bats are already infected (e.g., eastern North America and Europe). These measures are necessary to prevent the devastating effects this pathogen has had on bats in North America and would help maintain the ecosystem services that bats provide (9,10).
Acknowledgments
We thank the members of J.F.’s laboratory at Northeast Normal University for their help and support.
Financial support was provided by the National Science Foundation (NSF) East Asian Pacific Summer Institute program IIA-1415092, NSF grant DEB-1115895 and DEB-1336290, National Speleological Society Rapid Response Fund, US Fish and Wildlife Service, National Science and Technology Foundation grant no. 2013FY113600, The Robert and Patricia Switzer Foundation, and the crowd-funding platform of Experiment.com.
References
- Daszak P, Cunningham AA, Hyatt AD. Emerging infectious diseases of wildlife—threats to biodiversity and human health. Science. 2000;287:443–9. DOIPubMedGoogle Scholar
- St John RK, King A, de Jong D, Bodie-Collins M, Squires SG, Tam TW. Border screening for SARS. Emerg Infect Dis. 2005;11:6–10. DOIPubMedGoogle Scholar
- Warnecke L, Turner JM, Bollinger TK, Lorch JM, Misra V, Cryan PM, Inoculation of bats with European Geomyces destructans supports the novel pathogen hypothesis for the origin of white-nose syndrome. Proc Natl Acad Sci U S A. 2012;109:6999–7003. DOIPubMedGoogle Scholar
- Langwig KE, Hoyt JR, Parise KL, Kath J, Kirk D, Frick WF, Disease dynamics of white-nose syndrome invasion, midwestern United States, 2012–2014. Emerg Infect Dis. 2015;21:1023–6. DOIPubMedGoogle Scholar
- Puechmaille SJ, Wibbelt G, Korn V, Fuller H, Forget F, Muhldorfer K, Pan-European distribution of white-nose syndrome fungus (Geomyces destructans) not associated with mass mortality. PLoS ONE. 2011;6:e19167. DOIPubMedGoogle Scholar
- Langwig KE, Frick WF, Reynolds R, Parise KL, Drees KP, Hoyt JR, Host and pathogen ecology drive the seasonal dynamics of a fungal disease, white-nose syndrome. Proc Biol Sci. 2015;282:20142335. DOIPubMedGoogle Scholar
- Muller LK, Lorch JM, Lindner DL, O’Connor M, Gargas A, Blehert DS. Bat white-nose syndrome: a real-time TaqMan polymerase chain reaction test targeting the intergenic spacer region of Geomyces destructans. Mycologia. 2013;105:253–9. DOIPubMedGoogle Scholar
- Zukal J, Bandouchova H, Bartonicka T, Berkova H, Brack V, Brichta J, White-nose syndrome fungus: a generalist pathogen of hibernating bats. PLoS ONE. 2014;9:e97224. DOIPubMedGoogle Scholar
- Maine JJ, Boyles JG. Bats initiate vital agroecological interactions in corn. [Epub ahead of print]. Proc Natl Acad Sci U S A. 2015;•••:201505413.PubMedGoogle Scholar
- Langwig KE, Frick WF, Bried JT, Hicks AC, Kunz TH, Kilpatrick AM, Sociality, density-dependence and microclimates determine the persistence of populations suffering from a novel fungal disease, white-nose syndrome. Ecol Lett. 2012;15:1050–7. DOIPubMedGoogle Scholar
Figure
Cite This Article1Current affiliation: University of New Hampshire, Durham, New Hampshire, USA.
Related Links
Table of Contents – Volume 22, Number 1—January 2016
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Joseph R. Hoyt, University of California, Santa Cruz–Ecology and Evolutionary Biology, 1156 High St, Santa Cruz, CA 95064, USA; ; and Jiang Feng, Northeast Normal University, Changchun-School of Environment, 5268 Renmin St., Changchun, Jilin, People’s Republic of China
Top