Volume 22, Number 10—October 2016
Letter
Polymyxin B Resistance in Carbapenem-Resistant Klebsiella pneumoniae, São Paulo, Brazil
Cite This Article
Citation for Media
To the Editor: Infections caused by carbapenem-resistant Enterobacteriaceae have been associated with higher death rates than infections caused by carbapenem-susceptible strains, and resistant infections are mostly treated with polymyxins (1). Several outbreaks caused by carbapenem- and polymyxin-resistant Klebsiella pneumoniae (CPRKp) have been reported, mainly from Europe, and represent an emerging threat.
Carbapenem-resistant K. pneumoniae (CRKp) are endemic to Brazil, where polymyxin B (PMB) has been largely used against infections caused by these microorganisms. We evaluated PMB resistance rates and clonal diversity among CRKp isolates from patients in São Paulo, Brazil. The study was approved by the Research Review Board of Fleury Institute in São Paulo.
All K. pneumoniae isolates, except those from urine and active surveillance samples, recovered from inpatients during January 1, 2011–December 31, 2015, at 10 private tertiary-care hospitals in São Paulo were included in the study. K. pneumoniae isolates were identified by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry; we analyzed only the first isolate from each patient, unless other isolates were recovered after a 90-day interval. To determine antimicrobial drug MICs, we used nonautomated broth microdilution (2) polystyrene plates with cation-adjusted Mueller–Hinton broth (Becton Dickinson, Franklin Lakes, NJ, USA) for PMB and tigecycline; the Etest (bioMérieux, Marcy l’Etoile, France) for fosfomycin; and the disk-diffusion method (3) for all other antimicrobial drugs. Isolates with a MIC of <2 mg/L for PMB were considered susceptible; this value is the EUCAST (European Committee on Antimicrobial Susceptibility Testing) breakpoint for colistin in Enterobacteriaceae (2). Tigecycline and fosfomycin MICs were interpreted according to EUCAST guidelines (2). Imipenem and meropenem MICs were determined using the Etest for all isolates that were nonsusceptible to at least 1 carbapenem (ertapenem, meropenem, or imipenem) by disk-diffusion (3). We phenotypically detected class A carbapenemases as previously described (4).
We used convenience sampling to select 62 CPRKp isolates that were detected during 2014–2015 and used pulsed-field gel electrophoresis (PFGE) to evaluate their genomic DNA macrorestriction profiles after XbaI digestion. Dice similarity indexes were calculated using the UPGMA method with 1.25% tolerance and optimization (5). The minimal Dice index for a clonal group was defined as 80%.
We performed multilocus sequence typing as described (http://bigsdb.web.pasteur.fr/klebsiella/klebsiella.html) for 11 isolates that represented the 2 major PFGE clonal groups, CPRKp1 and CPRKp2. The full blaKPC nucleotide sequence was determined for these isolates (Technical Appendix Table 1).
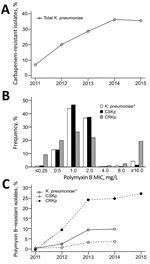
We included a total of 3,085 K. pneumoniae isolates in the analysis (Technical Appendix Table 2). A significant increase in carbapenem resistance (p<0.001) was seen from 2011 (6.8%) to 2015 (35.5%) (Figure, panel A). During the last year of analysis, we detected K. pneumoniae carbapenemase (KPC) in 96.2% of CRKp isolates.
PMB MICs showed a bimodal distribution that was clearly differentiated by a 2 mg/L MIC (Figure, panel B). When we stratified MICs by year and carbapenem resistance, a significantly increasing trend of resistance was seen among CRKp isolates from 2011 (0%) to 2014 (24.8%) to 2015 (27.1%) (p<0.001) (Figure, panel C). Resistance among carbapenem-susceptible K. pneumoniae varied from 0.7% in 2011 to 3.9% in 2014 (p = 0.002).
We did not evaluate the mechanism of PMB resistance. However, this resistance in KPC-producing K. pneumoniae is probably caused by the loss of mgrB function or the presence of nonsynonymous substitutions in pmrB that upregulate the pmrCAB and arnBCADTEF-pmrE operons, resulting in modification of lipid A. All these genes are located on the bacterial chromosome. Susceptibility testing showed that amikacin and tigecycline were the most active non–β-lactam antimicrobial agents against CPRKp isolates (amikacin 73.8%, tigecycline 69.4%) and CRKp isolates (amikacin 79.9%, tigecycline 72.2%) (Technical Appendix Table 3).
PFGE identified 2 major clonal groups. The largest group, CPRKp1 (n = 30), belonged to sequence type (ST) 11, and the other group, CPRKp2 (n = 17), belonged to ST437. Both STs belonged to clonal complex (CC) 258. Interhospital and intrahospital dissemination among private and public hospitals was observed. All isolates tested had the blaKPC-2 gene (Technical Appendix Figure).
In a previous study, the PMB resistance rate was 27% among 22 CRKp isolates from patients at a tertiary hospital in São Paulo during 2008–2010 (6). This rate is much higher than the rate we obtained for 2010, possibly because the previous study had a small number of isolates. CPRKp has been reported in various European countries at rates similar to those we report (7).
The predominance of CC258 among KPC-2-producing K. pneumoniae, but not among CPRKp, was reported in Brazil (8), and ST11, a variant of ST258, has occasionally been detected in colistin-resistant KPC-producing isolates in Spain (9). The ST437 clone has been reported in KPC-2 producers in China (10), but we found no reports of CPRKp among this clonal group. Our findings show an alarming yearly increase in the rate of PMB resistance among CRKp isolates, mostly KPC-2 producing, and the occurrence of interhospital and intrahospital dissemination of CPRKp from CC258 in São Paulo.
Acknowledgment
This work was supported by a grant from the Fleury Institute, São Paulo, Brazil. F. Bartolleti was supported by a scholarship from the National Counsel of Technological and Scientific Development (CNPq).
References
- Falagas ME, Tansarli GS, Karageorgopoulos DE, Vardakas KZ. Deaths attributable to carbapenem-resistant Enterobacteriaceae infections. Emerg Infect Dis. 2014;20:1170–5.DOIPubMedGoogle Scholar
- European Committee on Antimicrobial Susceptibility Testing. Breakpoint tables for interpretation of MICs and zone diameters, version 5.0 [cited 2016 Aug 12]. http://www.eucast.org/fileadmin/src/media/PDFs/EUCAST_files/Breakpoint_tables/v_5.0_Breakpoint_Table_01.pdf
- Clinical and Laboratory Standards Institute. Performance standards for antimicrobial susceptibility testing; twenty-fifth informational supplement (M100–S25). Wayne (PA): The Institute; 2015.
- Tsakris A, Kristo I, Poulou A, Themeli-Digalaki K, Ikonomidis A, Petropoulou D, Evaluation of boronic acid disk tests for differentiating KPC-possessing Klebsiella pneumoniae isolates in the clinical laboratory. J Clin Microbiol. 2009;47:362–7.DOIPubMedGoogle Scholar
- Heras J, Domínguez C, Mata E, Pascual V, Lozano C, Torres C, GelJ—a tool for analyzing DNA fingerprint gel images. BMC Bioinformatics. 2015;16:270.DOIPubMedGoogle Scholar
- Pereira GH, Garcia DO, Mostardeiro M, Ogassavara CT, Levin AS. Spread of carbapenem-resistant Klebsiella pneumoniae in a tertiary hospital in Sao Paulo, Brazil. J Hosp Infect. 2011;79:182–3.DOIPubMedGoogle Scholar
- Giske CG. Contemporary resistance trends and mechanisms for the old antibiotics colistin, temocillin, fosfomycin, mecillinam and nitrofurantoin. Clin Microbiol Infect. 2015;21:899–905.DOIPubMedGoogle Scholar
- Andrade LN, Curiao T, Ferreira JC, Longo JM, Clímaco EC, Martinez R, Dissemination of blaKPC-2 by the spread of Klebsiella pneumoniae clonal complex 258 clones (ST258, ST11, ST437) and plasmids (IncFII, IncN, IncL/M) among Enterobacteriaceae species in Brazil. Antimicrob Agents Chemother. 2011;55:3579–83.DOIPubMedGoogle Scholar
- Ruiz-Garbajosa P, Curiao T, Tato M, Gijón D, Pintado V, Valverde A, Multiclonal dispersal of KPC genes following the emergence of non-ST258 KPC-producing Klebsiella pneumoniae clones in Madrid, Spain. J Antimicrob Chemother. 2013;68:2487–92.DOIPubMedGoogle Scholar
- Wang LH, Wei DD, Wan LG, Yu Y, Deng Q, Liu Y. Diversity of the genetic environment of the blaKPC-2 gene among Klebsiella pneumoniae clinical isolates in a Chinese hospital. Microb Drug Resist. 2016;22:15–21.DOIPubMedGoogle Scholar
Figure
Cite This ArticleRelated Links
Table of Contents – Volume 22, Number 10—October 2016
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Jorge Luiz Mello Sampaio, Faculdade de Ciências Farmacêuticas, Avenida Professor Lineu Prestes, 580, Butantã, São Paulo, SP, Brazil, 05508-900
Top