Volume 23, Number 10—October 2017
Research
Investigation of Outbreaks of Salmonella enterica Serovar Typhimurium and Its Monophasic Variants Using Whole-Genome Sequencing, Denmark
Figure 4
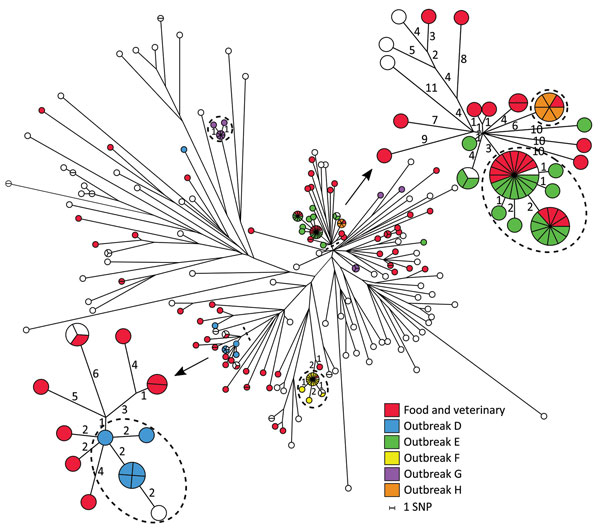
Figure 4. Maximum-parsimony tree of 169 human isolates and 73 linked food and veterinary isolates of Salmonella enterica serovar Typhimurium and the monophasic variants ST34 based on core-genome SNP analysis with an internal de novo assembled ST34 genome as the reference genome in an outbreak investigation of Salmonella Typhimurium and its monophasic variants, Denmark. Some branches are labeled with the number of SNP differences, and branch lengths correspond to the number of SNPs. Isolates belonging to outbreaks D, E, F, G, and H are as previously defined by MLVA. Isolates inside the dotted circles are outbreak isolates as defined by the SNP analysis. Two selected subgroups were reanalyzed separately with internal de novo assembled reference genomes (arrows). MLVA, multilocus variable-number tandem-repeat analysis; SNP, single-nucleotide polymorphism; ST, sequence type.