Volume 23, Number 10—October 2017
Dispatch
Hantavirus Pulmonary Syndrome Caused by Maripa Virus in French Guiana, 2008–2016
Figure
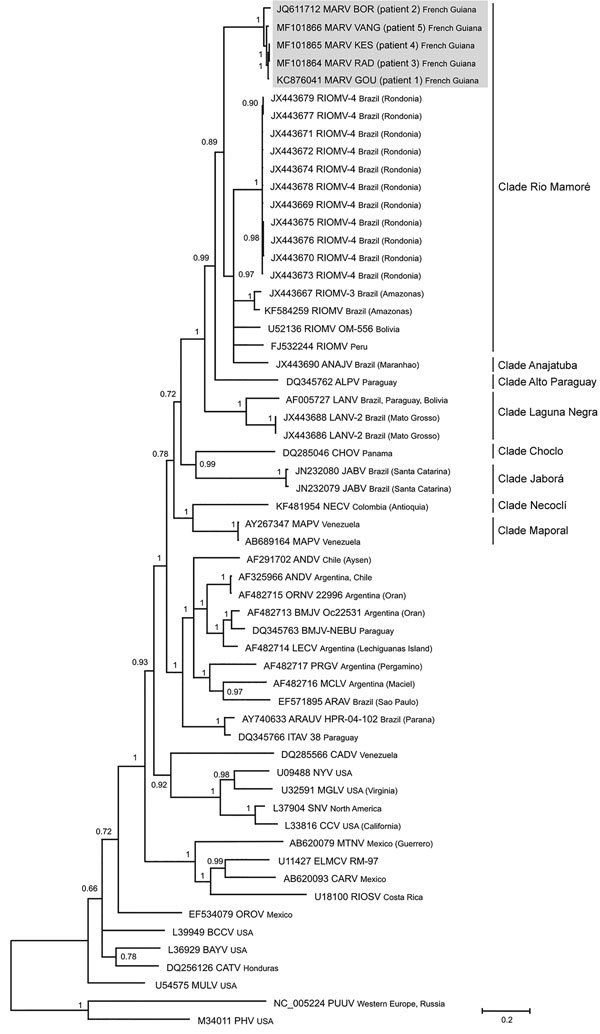
Figure. Phylogenetic tree based on the 1,308-bp fragment of the small (S) segment of 58 hantaviruses, including the 5 Maripa hantavirus isolates identified in French Guiana, 2008–2016 (gray shading). Tree was constructed by using the general time-reversible plus gamma distribution plus invariable site model of nucleotide evolution. GenBank accession numbers of viruses are indicated. Support for nodes was provided by the posterior probabilities of the corresponding clades. All resolved nodes have posterior probability >0.7. Scale bar indicates mean number of nucleotide substitutions per site.
Page created: September 18, 2017
Page updated: September 18, 2017
Page reviewed: September 18, 2017
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.