Volume 25, Number 5—May 2019
Dispatch
Genetic Characterization of Middle East Respiratory Syndrome Coronavirus, South Korea, 2018
Figure 2
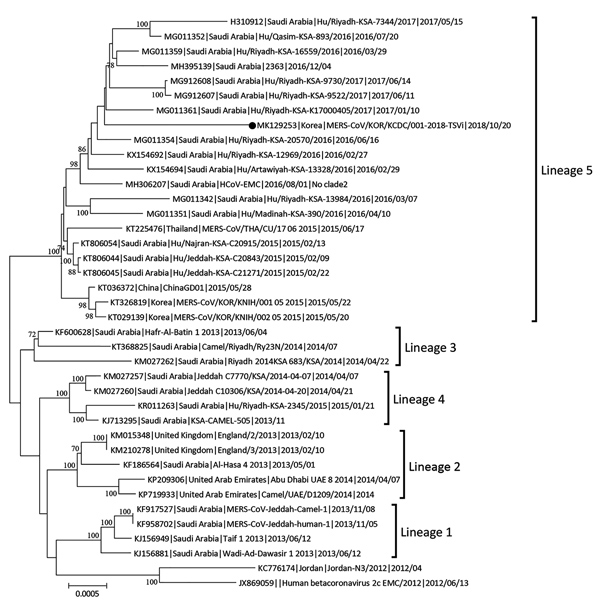
Figure 2. Maximum-likelihood tree showing MERS-CoV isolates from South Korea, September 2018 (black dot), and reference MERS-CoV genomes. Tree estimated using RAxML values (https://github.com/stamatak/standard-RAxML) on branches are shown as percentages based on 1,000 bootstrap replicates from the nucleotide sequences. MERS-CoV, Middle East respiratory syndrome coronavirus. Scale bar indicates nucleotide substitutions per site.
Page created: April 17, 2019
Page updated: April 17, 2019
Page reviewed: April 17, 2019
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.