Volume 26, Number 10—October 2020
Dispatch
Rapid, Sensitive, Full-Genome Sequencing of Severe Acute Respiratory Syndrome Coronavirus 2
Figure 1
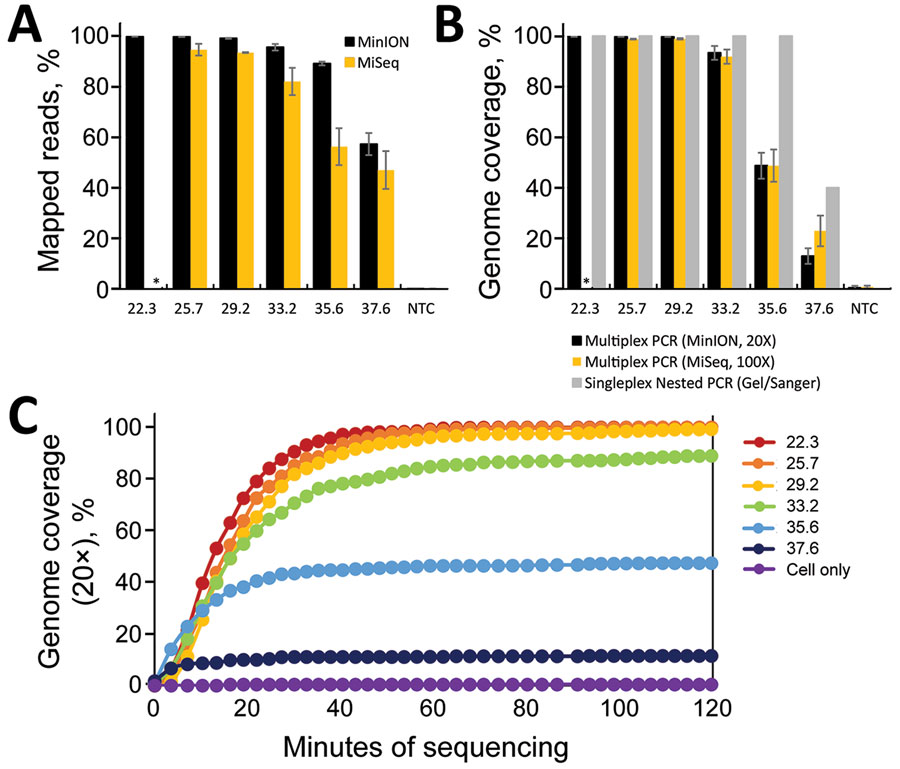
Figure 1. Limits of detection for sequencing severe acute respiratory syndrome coronavirus 2. Triplicate serial dilutions of virus isolate A12 (8) were amplified by using the singleplex or multiplex primer set. Multiplex amplicons were barcoded, library-prepped, and sequenced on an Oxford MinION apparatus (https://nanoporetech.com) or an Illumina MiSeq apparatus (https://www.illumina.com). A) Percentage of reads that map to the virus genome for each sample. B) Percentage of virus genome that is covered at >20× depth by the multiplex amplicons on the MinION (black) or >100× depth on the MiSeq (orange), or covered by the nested, singleplex amplicons (gray) (measured by presence or absence on a gel). C) Real-time analysis of MinION sequencing data. Each data point represents the average 20× genome coverage of three replicates. NTC, nontemplate controls (human cell nucleic acid carried through the PCR and library preparation). Asterisk (*) indicates that samples were not analyzed at that dilution.
References
- Holshue ML, DeBolt C, Lindquist S, Lofy KH, Wiesman J, Bruce H, et al.; Washington State 2019-nCoV Case Investigation Team. Washington State 2019-nCoV Case Investigation Team. First case of 2019 novel coronavirus in the United States. N Engl J Med. 2020;382:929–36. DOIPubMedGoogle Scholar
- Patel A, Jernigan DB, Abdirizak F, Abedi G, Aggarwal S, Albina D, et al.; 2019-nCoV CDC Response Team. 2019-nCoV CDC Response Team. Initial public health response and interim clinical guidance for the 2019 novel coronavirus outbreak—United States, December 31, 2019–February 4, 2020. MMWR Morb Mortal Wkly Rep. 2020;69:140–6. DOIPubMedGoogle Scholar
- Wang C, Horby PW, Hayden FG, Gao GF. A novel coronavirus outbreak of global health concern. Lancet. 2020;395:470–3. DOIPubMedGoogle Scholar
- World Health Organization. Coronavirus disease 2019 (COVID-19) situation report 141 [cited 2020 Jun 9]. https://www.who.int/emergencies/diseases/novel-coronavirus-2019/situation-reports
- Quick J, Grubaugh ND, Pullan ST, Claro IM, Smith AD, Gangavarapu K, et al. Multiplex PCR method for MinION and Illumina sequencing of Zika and other virus genomes directly from clinical samples. Nat Protoc. 2017;12:1261–76. DOIPubMedGoogle Scholar
- Quick J, Loman NJ, Duraffour S, Simpson JT, Severi E, Cowley L, et al. Real-time, portable genome sequencing for Ebola surveillance. Nature. 2016;530:228–32. DOIPubMedGoogle Scholar
- Holmes EC, Novel YZ. 2019 coronavirus genome, 2020 [cited 2020 Apr 5]. http://virological.org/t/novel-2019-coronavirus-genome/319
- Harcourt J, Tamin A, Lu X, Kamili S, Sakthivel SK, Murray J, et al. Severe Acute Respiratory Syndrome Coronavirus 2 from Patient with Coronavirus Disease, United States. Emerg Infect Dis. 2020;26:1266–73. DOIPubMedGoogle Scholar
- COVID-19 Investigation Team. Clinical and virologic characteristics of the first 12 patients with coronavirus disease 2019 (COVID-19) in the United States. Nat Med. 2020;26:861–8. DOIPubMedGoogle Scholar
- Andersen K. Clock and TMRCA based on 27 genomes, 2020 [cited 2020 Jan 25]. http://virological.org/t/clock-and-tmrca-based-on-27-genomes/347
- Lu R, Zhao X, Li J, Niu P, Yang B, Wu H, et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet. 2020;395:565–74. DOIPubMedGoogle Scholar
- Andersen KG, Rambaut A, Lipkin WI, Holmes EC, Garry RF. The proximal origin of SARS-CoV-2. Nat Med. 2020;26:450–2. DOIPubMedGoogle Scholar
- Deng X, Gu W, Federman S, du Plessis L, Pybus OG, Faria N, et al. Genomic surveillance reveals multiple introductions of SARS-CoV2 into northern California. Science. 2020 Jun 8:eabb9263.
1These authors contributed equally to this article.