Volume 26, Number 8—August 2020
Research Letter
mcr-Positive Escherichia coli ST131-H22 from Poultry in Brazil
Cite This Article
Citation for Media
Abstract
Escherichia coli sequence type (ST) 131 is of concern because it can acquire antimicrobial resistance and cause extraintestinal infections. E. coli ST131-H22 sublineage appears capable of being transmitted to humans through poultry. We report on multidrug-resistant ST131-H22 poultry isolates in Brazil closely related to international human and poultry isolates.
The pandemic, extraintestinal, pathogenic Escherichia coli multilocus sequence type (MLST) 131 lineage has emerged extensively, gaining notoriety for its extensively multidrug-resistant ST131-H30 sublineage (1). Whereas ST131-H30 appears to be transmitted primarily from person to person, the H22 sublineage may be transmitted zoonotically through poultry and cause urinary tract infections and urosepsis (2,3). We report isolating ST131-H22 strains that are multidrug resistant (MDR), meaning that they are resistant to >3 classes of antimicrobials (4), carrying mobile colistin-resistance (mcr) determinants from poultry in Brazil, the largest poultry-exporting country in the world.
We collected 64 E. coli strains from poultry with colibacillosis cases from 2 different farms in the same geographic region of Brazil and screened them by PCR for the ST131 clonal group (5). PCR detected 6 ST131 isolates (2 from the first farm, 4 from the second), which we whole-genome sequenced (BioProject no. PRJNA398035). We determined phenotypic antimicrobial susceptibility with disk diffusion testing, except for isolates carrying the mcr gene, which we tested using broth microdilution (6).
We trimmed the reads and used QUAST (http://quast.sourceforge.net) to evaluate the quality of assemblies (contig lengths and expected genome sizes). We assembled DNA sequences with SPAdes (http://cab.spbu.ru/software/spades), then determined the serotype, phylogroup, MLST, fimH protein type, virulence gene profile, plasmid replicons, and markers of antimicrobial resistance for each isolate in silico using the ABRicate virulence factors database (https://github.com/tseemann/abricate) and ResFinder/PlasmidFinder tools from CGE (https://cge.cbs.dtu.dk/services). Genes were identified with a minimum of >95% of identity and coverage.
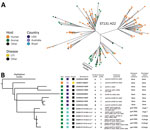
We identified all isolates as O25:H4-ST131-H22, all belonging to phylogroup B2. We generated a maximum-likelihood phylogeny tree on the basis of core-genome single-nucleotide polymorphisms, including the 6 isolates from Brazil and 140 ST131-H22 sequences from EnteroBase (http://enterobase.warwick.ac.uk) and a previous study (2), using the Northern Arizona SNP Pipeline (https://tgennorth.github.io/NASP/) aligned against E. coli JJ1886 ST131-H30 (GenBank accession no. CP006784) (Appendix). The 6 isolates from poultry were nested within a clade of intermingled poultry and human clinical isolates within the overall international isolates (Figure, panel A). The isolates from Brazil were closely related to ST131-H22 avian pathogenic E. coli isolates from poultry in the United States and those from a human urinary tract infection in Australia (Figure, panel B). Identical virulence factors and plasmid replicons were observed among 4 β-lactamase positive isolates and between 2 isolates missing the β-lactamase genes but carrying mcr colistin–resistance determinants. All 6 isolates had MDR profiles, phenotypically confirmed (data not shown except for those from colistin microdilution method) (Figure, panel B).
The ST131-H22 lineage, while currently not as common as the H30 sublineage as a cause of community-acquired infections, does present a public health challenge because it colonizes poultry flocks, contaminating retail poultry products, and carries mcr colistin–resistance genes (3). The enormity and rapid growth of poultry production, in which many developing countries use antimicrobials extensively (5), and its zoonotic potential, make ST131-H22 worthy of specific attention (2).
Findings from our phylogenetic analyses of a global collection of ST131-H22 isolates from humans and poultry support findings from previous studies (2,3) and underscore the zoonotic potential of this virulent sublineage. Given that Brazil annually processes 13.8 million poultry products and exports 3.8 million kilograms (4), these findings warrant further examination to assess potential zoonotic spillover in Brazil and poultry-importing countries. Until such studies are conducted, the zoonotic potential of ST131-H22 in flocks in Brazil cannot be quantified.
The discovery of mcr mobile colistin resistance determinants in food animals has renewed attention to the potential risks of widespread antimicrobial use in livestock. In Latin America, mcr-5 has been found in poultry in Paraguay (9). The description of the mcr-9 homologue from humans in the United States and horses in Sweden has raised attention to another mcr gene with potential for global spread (10). Both mcr variants in this study, 153_Br and 157_Br, showed phenotypic resistance (6) and came from the same farm (Figure, panel B). Interestingly, 153_Br carried both mcr-5.1 and mcr-9 variants. These isolates may portend a more widespread problem within poultry flocks in Brazil.
Isolates from this study showed resistance to all of the World Health Organization’s highest priority critically important antimicrobial classes (Figure, panel B) (8). Analysis of the absence of tetracycline resistance (tet[B]/[D]) in 1 of our isolates (Figure, panel B) indicates partial plasmid loss (data not shown).
Use of colistin as a growth promoter in livestock was banned in Brazil in November 2016, although it continued being therapeutically used in poultry up to 2018 (7). Therefore, mcr-encoding H22 strains could be selected out of the population over time. Further restrictions will have to be implemented to combat the growing resistance of E. coli in poultry in Brazil to critically important antimicrobial drugs (4).
Our findings suggest that poultry in Brazil may serve as a reservoir for MDR extraintestinal pathogenic E. coli carrying mobile colistin-resistance determinants. These findings highlight the need for better antimicrobial stewardship and surveillance systems to determine the prevalence of MDR E. coli ST131-H22 in these poultry flocks and clarify the risks posed to domestic and international poultry consumers.
Dr. Becker Saidenberg is a PhD student and researcher at the College of Veterinary Medicine, University of São Paulo, São Paulo, Brazil. He is engaged in microbiological studies focused on poultry and wildlife and the zoonotic aspects connected to the animal-human interface.
Acknowledgments
The authors wish to thank the invaluable support provided with the laboratory processing of samples for whole-genome sequencing by the Department of Bacteria, Parasites and Fungi of the Statens Serum Institut (SSI), Denmark.
This research was sponsored in part by FAPESP (2011/18204, and 2014/11523-7) and in part by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior - Brasil (CAPES - Finance Code 001- André Becker S. Saidenberg - PhD student).
References
- Manges AR. Escherichia coli and urinary tract infections: the role of poultry-meat. Clin Microbiol Infect. 2016;22:122–9. DOIPubMedGoogle Scholar
- Liu CM, Stegger M, Aziz M, Johnson TJ, Waits K, Nordstrom L, et al. Escherichia coli ST131-H22 as a foodborne uropathogen. MBio. 2018;9:e00470–18. DOIPubMedGoogle Scholar
- Roer L, Overballe-Petersen S, Hansen F, Johannesen TB, Stegger M, Bortolaia V, et al. ST131 fimH22 Escherichia coli isolate with a blaCMY-2/IncI1/ST12 plasmid obtained from a patient with bloodstream infection: highly similar to E. coli isolates of broiler origin. J Antimicrob Chemother. 2019;74:557–60. DOIPubMedGoogle Scholar
- Van Boeckel TP, Brower C, Gilbert M, Grenfell BT, Levin SA, Robinson TP, et al. Global trends in antimicrobial use in food animals. Proc Natl Acad Sci U S A. 2015;112:5649–54. DOIPubMedGoogle Scholar
- Doumith M, Day M, Ciesielczuk H, Hope R, Underwood A, Reynolds R, et al. Rapid identification of major Escherichia coli sequence types causing urinary tract and bloodstream infections. J Clin Microbiol. 2015;53:160–6. DOIPubMedGoogle Scholar
- Clinical and Laboratory Standards Institute. Performance standards for antimicrobial susceptibility testing; 28th informational supplement (M100–S28). Wayne (PA): The Institute; 2018.
- Brazil. Governmental Normative Instruction IN-45. Diario Oficial da Uniao. 2016. Nov 11 [cited 2020 Mar 20]. http://www.in.gov.br/materia/-/asset_publisher/Kujrw0TZC2Mb/content/id/22078290/do1-2016-11-30-instrucao-normativa-n-45-de-22-de-novembro-de-2016-22078259
- World Health Organization. Critically important antimicrobials for human medicine. 2011 [cited 2019 Oct 21]. http://apps.who.int/iris/bitstream/10665/77376/1/9789241504485%20eng.pdf
- Nesporova K, Jamborova I, Valcek A, Medvecky M, Literak I, Dolejska M. Various conjugative plasmids carrying the mcr-5 gene in Escherichia coli isolates from healthy chickens in Paraguay. J Antimicrob Chemother. 2019;74:3394–7. DOIPubMedGoogle Scholar
- Börjesson S, Greko C, Myrenås M, Landén A, Nilsson O, Pedersen K. A link between the newly described colistin resistance gene mcr-9 and clinical Enterobacteriaceae isolates carrying blaSHV-12 from horses in Sweden. J Glob Antimicrob Resist. 2020;20:285–9. DOIPubMedGoogle Scholar
Figure
Cite This ArticleOriginal Publication Date: July 03, 2020
Table of Contents – Volume 26, Number 8—August 2020
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Andre Becker S. Saidenberg, University of São Paulo, Av. Prof. Orlando Marques de Paiva, 87, 05508-270, São Paulo, Brazil
Top