Volume 27, Number 11—November 2021
Dispatch
Isolation of Novel Mycobacterium Species from Skin Infection in an Immunocompromised Person
Cite This Article
Citation for Media
Abstract
We investigated a case of cutaneous infection in an immunocompromised patient in China that was caused by a novel species within the Mycobacterium gordonae complex. Results of whole-genome sequencing indicated that some strains considered to be M. gordonae complex are actually polyphyletic and should be designated as closely related species.
Mycobacterium gordonae was first described 50 years ago as a slow-growing scotochromogenic nontuberculous mycobacterium. Previous research revealed vague molecular typing results for M. gordonae–like strains. For example, the RNA polymerase-β (rpoB) PCR restriction analysis discriminates M. gordonae into 4 clusters even though cluster D does not hybridize well with the type strain (1). Two novel species, M. paragordonae and M. vicinigordonae, share >99% 16S rRNA gene similarity with M. gordonae, which might also lead to confusion about their classification (2,3).
M. gordonae is frequently isolated from water systems and clinical samples as a potential opportunistic pathogen (4,5); clinical infections ranging from skin and lung infections to disseminated systemic infections have been reported, especially in immunosuppressed patients (6,7). Both the M. paragordonae and M. vicinigordonae strains were first isolated as nonpathogenic organisms from pneumonia patients (2,3). M. paragordonae is often isolated from hospital water systems and devices, but only 1 case of iatrogenic M. paragordonae infection has been reported (8,9). These reports reveal the dissimilar effects produced by different M. gordonae–like strains.
The advent of whole-genome sequencing has brought genomewide analyses into common use to delineate species (10–12). The widely accepted cutoffs adopted for the average nucleotide identity (ANI) and in silico DNA–DNA hybridization (isDDH), 95%–97% for ANI and 70% for isDDH, strongly correlate with traditional DDH division values, providing more robust resolution than phenotyping or mycolic acid analysis for determining mycobacterial taxonomy (11,12). We report a case of cutaneous infection in Jiangsu Province, China, caused by a previously undescribed novel species belonging to the M. gordonae group.
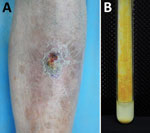
A 63-year-old man was admitted to the hospital for a 5-year history of a nodule on his left shin. The asymptomatic lesion initially appeared as a papule and gradually developed into a dull red verrucous nodule with scales (Figure 1, panel A). No trauma history before the onset was reported. The patient had received a diagnosis of lupus erythematosus 30 years earlier and had taken oral prednisone (20 mg/d) over the previous year. Laboratory test results indicated no remarkable findings. Histologic examination of a skin sample showed irregular epithelial hyperplasia and granulomatous infiltrations of a large number of epithelioid histocytes, neutrophil cells, plasma cells, and lymphocytes in the dermal layer. After 19 days of culture, orange colonies were observed on modified Löwenstein–Jensen slants at 32°C (Figure 1, panel B). The organism was scotochromogenic with a smooth appearance and grew well at 32°C and 37°C on both Löwenstein–Jensen slants and Middlebrook 7H9 with oleic acid dextrose citrate. The colonies were confirmed to be rod-shaped, acid-fast bacterium.
We extracted DNA from the colonies for PCR analysis and compared the sequences using BLAST (https://blast.ncbi.nlm.nih.gov/Blast.cgi). The 16S rRNA (1452 bp) gene shared greatest similarity (99.51%) with M. gordonae ASCr-1.2; gene sequencing showed rpoB (365 bp) shared 97.53% and hsp65 (765 bp) 95.53% similarity with M. paragordonae 49061. On the basis of these results, we diagnosed infection with a member of the M. gordonae complex. Drug sensitivity analysis revealed the bacterium to be sensitive to moxifloxacin, levofloxacin, ethambutol, and amikacin and resistant to clarithromycin, isoniazid, and rifampicin; we therefore prescribed a moxifloxacin regimen for the patient. Because the lesion had not healed over several years, we surgically resected it and applied photodynamic therapy after 2 months of antimicrobial drug treatment. One month later, the patient reported that the lesion had recovered well and refused further oral antimicrobial drugs. No recurrence was observed in the following year.
To accurately identify the pathogen to the species level, we performed whole-genome sequencing (8,509,558 reads, 110×) of the isolate X7091 using the Illumina Hiseq 4000 (https://www.illumina.com) and PacBio RS II (https://www.pacb.com) platforms at the Beijing Genomics Institute. Sequence data indicated a 7.1-Mb genome (7,319,570 bp) including a plasmid (216,348 bp) with a guanine-cytosine content of 64.6% (Genbank accession no. GCA_017086405.1). The complete genome had a guanine-cytosine content of 66.7%, similar to M. gordonae (66.8%) and M. paragordonae (67.0%). Functional annotation obtained through multiple databases revealed 6,704 coding sequences, 48 tRNA, 3 rRNA, and 35 small RNA genes.
We compared this isolate with all available genomes of the M. gordonae group using whole-genome-based computational strategies. ANI calculated by FastANI (https://github.com/ParBLiSS/FastANI) revealed that the closest matches, with M. gordonae HMC_M15 (87.80%) and M. gordonae DSM 44160 (87.79%), were well below the threshold for species delineation (Appendix Table) (11). Evaluating isDDH using the Type Strain Genome Server (https://tygs.dsmz.de) showed weak relations with M. gordonae DSM 44160 (34.5%) and M. paragordonae 49061 (31.3%) (13) (Appendix Table); we found no closely related genome in the database. For M. paragordonae strains, ANI was 97.8%–98.6% and isDDH 80.4%–99.9%; for M. gordonae strains, ANI was 99.1%–99.9% and isDDH 93.0%–99.3%.
The core-genome phylogeny of the M. gordonae complex, constructed using a previously described method, suggested that the isolated strain is a branch within the cluster but distant from other M. gordonae-like strains (14,15) (Figure 2, panel B). Integration of these highly concordant results strongly suggested that the isolate is distinct from present M. gordonae–like strains and represents a novel species within the M. gordonae complex. We proposed Mycobacterium camsnse sp. nov. as the name for this strain.
When comparing the similarity index of all available genomes of the M. gordonae group, we found clear demarcations among the M. camsnse X7091, M. gordonae CTRI 14-8773, M. vicinigordonae 24, 7 M. paragordonae, and 3 M. gordonae strains including the type strain DSM 44160 (Figure 2). Two strains recorded as M. gordonae ssp. in the cluster of M. paragordonae may have previously been misclassified. M. gordonae CTRI 14-8773, isolated in Russia, also represents a novel species of the M. gordonae group. These results confirmed the genomic diversity of M. gordonae–like strains, corroborating that the M. gordonae group is polyphyletic and should be divided into >5 closely related species.
M. gordonae is generally considered a minimally pathogenic mycobacteria. Nonetheless, clinical infections have been reported, even in immunocompetent individuals (6–7). We isolated a distinct strain within the M. gordonae group from a skin infection using whole genome–level approaches based on ANI, isDDH, and core gene phylogeny. We proposed the name Mycobacterium camsnse sp. nov. for this strain. We also provided genomic insights into the heterogeneity of the M. gordonae–like strains, including finding 2 strains potentially misclassified as M. gordonae ssp., and demonstrated that the present M. gordonae group should be designated as 5 closely related species.
Conventional routines for describing species, such as wet-lab DDH and phenotypic tests, which are tedious and restricted by laboratory capacity, often do not delineate closely related species. Genome-based analysis affords a more accurate alternative for delineating species. Although digital gene-expression analyses might not have provided enough conclusive authentication data, the low genomewide similarity between the strain X7091 and M. gordonae group strongly support it as a novel species. Only a few M. gordonae assembly models are available, and large-scale investigations are needed to better understand species diversity, geographic distribution, and clinical significance of M. camsnse sp. nov. infections.
The persistent but limited nonpurulent lesion of the immunosuppressed patient in our study reflects the attenuated nature of the pathogen. The various drug resistance properties of X7091 and other M. gordonae–like strains indicate the need for drug sensitivity testing before initiating drug treatment for M. gordonae–like strain infections (6,7). Our patient responded well, indicating that operation combined with antimicrobial therapy could be a good option for treating environmental Mycobacterium-induced skin infections.
This study was supported by grants from the Chinese Academy of Medical Sciences Innovation Fund for Medical Sciences (2016-I2M-1-005, 2017-I2M-B&R-14) and the National Natural Science Foundation of China (81972950).
Dr. Mei is a dermatologist at the Institute of Dermatology, Chinese Academy of Medical Sciences and Peking Union Medical College in Nanjing, China. Her primary research interest is mycobacterial infectious diseases.
References
- Itoh S, Kazumi Y, Abe C, Takahashi M. Heterogeneity of RNA polymerase gene (rpoB) sequences of Mycobacterium gordonae clinical isolates identified with a DNA probe kit and by conventional methods. J Clin Microbiol. 2003;41:1656–63. DOIPubMedGoogle Scholar
- Kim BJ, Hong SH, Kook YH, Kim BJ. Mycobacterium paragordonae sp. nov., a slowly growing, scotochromogenic species closely related to Mycobacterium gordonae. Int J Syst Evol Microbiol. 2014;64:39–45. DOIPubMedGoogle Scholar
- Liu G, Yu X, Luo J, Hu Y, Dong L, Jiang G, et al. Mycobacterium vicinigordonae sp. nov., a slow-growing scotochromogenic species isolated from sputum. Int J Syst Evol Microbiol. 2021;71:71. DOIPubMedGoogle Scholar
- Loret JF, Dumoutier N. Non-tuberculous mycobacteria in drinking water systems: A review of prevalence data and control means. Int J Hyg Environ Health. 2019;222:628–34. DOIPubMedGoogle Scholar
- Barber TW, Craven DE, Farber HW. Mycobacterium gordonae: a possible opportunistic respiratory tract pathogen in patients with advanced human immunodeficiency virus, type 1 infection. Chest. 1991;100:716–20. DOIPubMedGoogle Scholar
- Al-Busaidi I, Wong D, Boggild AK. Cutaneous Mycobacterium gordonae infection in an elderly diabetic returned traveller. J Travel Med. 2017;24:24. DOIPubMedGoogle Scholar
- Chen Y, Jiang J, Jiang H, Chen J, Wang X, Liu W, et al. Mycobacterium gordonae in patient with facial ulcers, nosebleeds, and positive T-SPOT.TB test, China. Emerg Infect Dis. 2017;23:1204–6. DOIPubMedGoogle Scholar
- Cheung CY, Cheng NHY, Ting WM, Chak WL. Mycobacterium paragordonae: a rare cause of peritonitis in a peritoneal dialysis patient. Clin Nephrol. 2017;88:371–2. DOIPubMedGoogle Scholar
- Takajo I, Iwao C, Aratake M, Nakayama Y, Yamada A, Takeda N, et al. Pseudo-outbreak of Mycobacterium paragordonae in a hospital: possible role of the aerator/rectifier connected to the faucet of the water supply system. J Hosp Infect. 2020;104:545–51. DOIPubMedGoogle Scholar
- Jagielski T, Borówka P, Bakuła Z, Lach J, Marciniak B, Brzostek A, et al. Genomic insights into the Mycobacterium kansasii complex: an update. Front Microbiol. 2020;10:2918. DOIPubMedGoogle Scholar
- Jain C, Rodriguez-R LM, Phillippy AM, Konstantinidis KT, Aluru S. High throughput ANI analysis of 90K prokaryotic genomes reveals clear species boundaries. Nat Commun. 2018;9:5114. DOIPubMedGoogle Scholar
- Zong Z. Genome-based taxonomy for bacteria: a recent advance. Trends Microbiol. 2020;28:871–4. DOIPubMedGoogle Scholar
- Meier-Kolthoff JP, Göker M. TYGS is an automated high-throughput platform for state-of-the-art genome-based taxonomy. Nat Commun. 2019;10:2182. DOIPubMedGoogle Scholar
- Stamatakis A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics. 2014;30:1312–3. DOIPubMedGoogle Scholar
- Luo T, Xu P, Zhang Y, Porter JL, Ghanem M, Liu Q, et al. Population genomics provides insights into the evolution and adaptation to humans of the waterborne pathogen Mycobacterium kansasii. Nat Commun. 2021;12:2491. DOIPubMedGoogle Scholar
Figures
Cite This ArticleOriginal Publication Date: October 07, 2021
1These first authors contributed equally to this article.
Table of Contents – Volume 27, Number 11—November 2021
| EID Search Options |
|---|
|
|
|
|
|
|

Please use the form below to submit correspondence to the authors or contact them at the following address:
Hong-sheng Wang, Institute of Dermatology, Chinese Academy of Medical Sciences and Peking Union Medical College, St.12 Jiangwangmiao, Nanjing, Jiangsu, 210042, China
Top