Volume 27, Number 2—February 2021
Dispatch
Estimating Transmission Parameters for COVID-19 Clusters by Using Symptom Onset Data, Singapore, January–April 2020
Cite This Article
Citation for Media
Abstract
We estimated the generation interval distribution for coronavirus disease on the basis of serial intervals of observed infector–infectee pairs from established clusters in Singapore. The short mean generation interval and consequent high prevalence of presymptomatic transmission requires public health control measures to be responsive to these characteristics of the epidemic.
A systematic review estimated that the basic reproduction number (R0) for coronavirus disease (COVID-19) is 2–3 (1). However, R0 alone is insufficient to characterize an epidemic. The distribution of the serial interval (i.e., the length of time between symptom onset of 2 cases) has been estimated for COVID-19; mean intervals range from 3.1 to 7.5 days (2,3). Estimation of the generation interval (Tg) (i.e., the length of time between the points of infection for 2 linked cases) is less common. Although studies have reported means of 3.3 and 5.0 days (4; Li et al., unpub. data, ), Ganyani et al. (5) estimated the mean (+ SD) of Tg to be 3.9 (+ 2.7) days, on the basis of which they estimated that 66% (95% credible interval [CrI] 45%–84%) of transmission occurred before symptoms. Another study of 77 pairs estimated the same proportion to be 44% (95% CI 25%–69%) (6). Because conventional outbreak control measures are centered around isolation, contact tracing, and treatment of symptomatic case-patients, a high prevalence of presymptomatic transmission (p) would warrant shifting measures to address potential transmission among persons with no apparent symptoms (7). Hence, to inform control measures for the outbreak in Singapore, we generated estimates of Tg, R0, and p by using published symptom onset data for COVID-19 cases in Singapore.
We implemented a cross-sectional study design to estimate Tg, R0, and p for the COVID-19 outbreak in Singapore during January 23–April 6, 2020. Given that containment measures were initiated over the duration of the study, we considered R0 𝑡o be the effective reproduction number of the outbreak. All confirmed COVID-19 cases classified by the Ministry of Health of Singapore (MOH) as linked to a local cluster were included in this analysis. Information on case number, cluster, patient age and sex, imported status, date of symptom onset (DOO), and known contacts who have also been confirmed as case-patients were extracted from daily press releases published by MOH. DOOs for cases that were not available from press releases were extracted from a similar anonymized dataset of COVID-19 admissions to the National Centre for Infectious Diseases, Singapore. Cases with DOOs not available from that dataset were subsequently excluded from analysis. Our study was approved by the ethics review board of National Healthcare Group, Singapore.
We identified index cases and potential infectors of each case-patient on the basis of available information of the case-patients’ known contacts, published case links, and a heuristic to sensibly include potential infectors who could have transmitted the infection to the case-patients (Appendix). We subsequently used the infector–infectee pairs constructed to estimate the serial and generation interval distribution.
Assuming the same incubation period with mean (+ SD) of 5.2 (+ 2.8) days, we replicated the Bayesian Markov chain Monte Carlo procedure detailed in Ganyani et al. (5) to estimate the mean (SD) of the Tg (Appendix). With the estimated parameters, we constructed the distribution of R0 and subsequently p by simulating infections and computing the proportion of presymptomatic transmissions. We conducted subgroup analyses for case-patients with multiple, family, or no contacts, and for clusters with no missing DOO. We conducted sensitivity analyses estimating the distribution of R0 by using resampled values from a 95% CI of the epidemic growth rate and group-specific rates. For each distribution, we reported the median and 95% CrI. All analyses were conducted by using RStudio 1.2.5033 (https://rstudio.com).
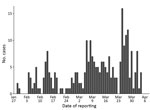
A total of 1,375 confirmed cases had been reported as of April 6, 2020, and we applied our exclusion criteria to obtain a final sample size of 257 cases (Figure 1). We have summarized sample characteristics (Table 1) and the spread of cases over time (Figure 2). Because 48 index case-patients had no known infector, a maximum of 209 infector–infectee pairs were constructed for analysis.
Analyzing the 209 pairs, we estimated the mean Tg to be 3.44 (95% CrI 2.79–4.11) days, with an SD of 2.39 (95% CrI 1.27–3.45) days (Table 2). This estimate corresponded to an R0 of 1.09 (95% CrI 1.08–1.11) and p of 0.72 (95% CrI 0.64–0.80). We estimated the serial interval distribution (Appendix Table 1) and convergence plots for all analyses (Appendix Figure 3).
Examining the 93 pairs with only 1 known contact, the estimates for mean Tg, SD Tg, and R0 increased, whereas p decreased (Table 2). The 116 pairs that required identification of potential infectors had a shorter mean Tg and a higher p in comparison (Table 2). Subgroup analyses are summarized in Appendix Table 2. However, the chains for pairs with family or no known contact exhibited poor convergence, and estimates were not reported. Sensitivity analyses using resampled growth rates and group-specific rates did not yield estimates differing from those of the main analyses (Appendix Table 3).
The mean generation interval of the COVID-19 outbreak in Singapore was estimated at 3.44 days, suggesting that an infected person would be expected to pass on an infection to another person in 3 days, within the range of 3.3–5.0 days reported by other studies (4,5; Li et al., unpub. data). Pairs with only 1 known contact yielded a larger estimate of 3.93 days, whereas pairs for whom infectors were identified had a shorter mean generation interval of 3.03 days. These results suggest that we might best report the upper bound of estimates, accounting for the presence of unclear transmission links within the clusters.
The estimated was slightly >1, higher than other estimates reported as of March 31, 2020 (8). We observed a high p, potentially a result of prompt isolation of symptomatic case-patients (M. Casey et al., unpub. data, https://doi.org/10.1101/2020.05.08.20094870). This higher proportion might also be attributable to our allowance of infector DOOs to be up to 3 days after their infectees’ DOOs, establishing the plausibility of presymptomatic transmission. We acknowledge that this cutoff would have an influence on our eventual estimates. Although negative serial intervals >3 days have occurred in other studies (5; Z. Du et al., unpub. data, ), we chose a conservative cutoff of 3 days consistent with He et al. (6), where 9% of transmissions would occur before 3 days before DOO.
Nonetheless, the high prevalence of presymptomatic transmission in the community requires public health strategies to be responsive to this characteristic to remain effective. Universal wearing of masks in the community might reduce the likelihood of transmission through saliva and respiratory droplets (9). In place of testing when symptoms are observed, universal testing of persons living in or working with confined populations should be prioritized to mitigate the risk for transmission of the infection into these populations (10). Contact tracing should be modified to include the period before symptom onset (6,7) and should adopt a digital approach to be more comprehensive and less labor intensive (4).
Our study generated estimates that accounted for the uncertainty arising from multiple potential infectors and a small sample size, which contributes to the scarce information about disease characteristics. Because we dropped cases without a reported DOO, and DOO data and contact information were self-reported, our estimates might be subject to selection, self-report, and recall biases. Our estimation approach assumed equal probability of infecting among potential infectors, although a higher likelihood of transmission among household contacts has been suggested (11). We also did not account for the potential formation of cyclical infector networks, although their effects on the estimates have been demonstrated to be limited (12). Nevertheless, our estimates contribute to knowledge about the transmission dynamics of COVID-19 and have implications for control measures.
Ms. Ng is a research analyst with the National Healthcare Group in Singapore. Her research interests include the application of biostatistics in epidemiology and health services research.
Acknowledgments
We thank the Singapore Ministry of Health for their tireless efforts in outbreak control and publication of the data for this manuscript.
N.H. acknowledges funding provided by the EpiPose project from the European Union’s SC1-PHE-CORONAVIRUS-2020 programme (project no. 101003688) and by the European’s Horizon 2020 research and innovation programme (grant agreement no. 682540–TransMID).
References
- Park M, Cook AR, Lim JT, Sun Y, Dickens BL. A systematic review of COVID-19 epidemiology based on current evidence. J Clin Med. 2020;9:967. DOIPubMedGoogle Scholar
- Zhao S, Cao P, Chong MKC, Gao D, Lou Y, Ran J, et al. The time-varying serial interval of the coronavirus disease (COVID-19) and its gender-specific difference: a data-driven analysis using public surveillance data in Hong Kong and Shenzhen, China from January 10 to February 15, 2020. Infect Control Hosp Epidemiol. 2020;41:750–1. DOIPubMedGoogle Scholar
- Li Q, Guan X, Wu P, Wang X, Zhou L, Tong Y, et al. Early transmission dynamics in Wuhan, China, of novel coronavirus–infected pneumonia. N Engl J Med. 2020;382:1199–207. DOIPubMedGoogle Scholar
- Ferretti L, Wymant C, Kendall M, Zhao L, Nurtay A, Abeler-Dörner L, et al. Quantifying SARS-CoV-2 transmission suggests epidemic control with digital contact tracing. Science. 2020;368:
eabb6936 ; Epub ahead of print. DOIPubMedGoogle Scholar - Ganyani T, Kremer C, Chen D, Torneri A, Faes C, Wallinga J, et al. Estimating the generation interval for coronavirus disease (COVID-19) based on symptom onset data, March 2020. Euro Surveill. 2020;25:
2000257 . DOIPubMedGoogle Scholar - He X, Lau EHY, Wu P, Deng X, Wang J, Hao X, et al. Author Correction: Temporal dynamics in viral shedding and transmissibility of COVID-19. Nat Med. 2020;26:1491–3. DOIPubMedGoogle Scholar
- Wei WE, Li Z, Chiew CJ, Yong SE, Toh MP, Lee VJ. Presymptomatic Transmission of SARS-CoV-2 - Singapore, January 23-March 16, 2020. MMWR Morb Mortal Wkly Rep. 2020;69:411–5. DOIPubMedGoogle Scholar
- Tariq A, Lee Y, Roosa K, Blumberg S, Yan P, Ma S, et al. Real-time monitoring the transmission potential of COVID-19 in Singapore, March 2020. BMC Med. 2020;18:166. DOIPubMedGoogle Scholar
- Cheng VCC, Wong SC, Chuang VWM, So SYC, Chen JHK, Sridhar S, et al. The role of community-wide wearing of face mask for control of coronavirus disease 2019 (COVID-19) epidemic due to SARS-CoV-2. J Infect. 2020;81:107–14. DOIPubMedGoogle Scholar
- Gandhi M, Yokoe DS, Havlir DV. Asymptomatic transmission, the Achilles’ Heel of current strategies to control Covid-19. N Engl J Med. 2020;382:2158–60. DOIPubMedGoogle Scholar
- Bi Q, Wu Y, Mei S, Ye C, Zou X, Zhang Z, et al. Epidemiology and transmission of COVID-19 in 391 cases and 1286 of their close contacts in Shenzhen, China: a retrospective cohort study. Lancet Infect Dis. 2020;20:911–9. DOIPubMedGoogle Scholar
- Kremer C, Ganyani T, Chen D, Torneri A, Faes C, Wallinga J, et al. Authors’ response: Estimating the generation interval for COVID-19 based on symptom onset data. Euro Surveill. 2020;25:
2001269 . DOIPubMedGoogle Scholar
Figures
Tables
Cite This ArticleOriginal Publication Date: January 12, 2021
Table of Contents – Volume 27, Number 2—February 2021
| EID Search Options |
|---|
|
|
|
|
|
|

Please use the form below to submit correspondence to the authors or contact them at the following address:
Sheryl Hui-Xian Ng, National Healthcare Group, Health Services and Outcomes Research, 3 Fusionopolis Link, 03-08, Singapore 138543
Top