Volume 27, Number 6—June 2021
Research Letter
Rapid Antigen Test for Postmortem Evaluation of SARS-CoV-2 Carriage
Cite This Article
Citation for Media
Abstract
Detecting severe acute respiratory syndrome coronavirus 2 in deceased patients is key when considering appropriate safety measures to prevent infection during postmortem examinations. A prospective cohort study comparing a rapid antigen test with quantitative reverse transcription PCR showed the rapid test’s usability as a tool to guide autopsy practice.
Rapid detection of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is essential to prevent viral dissemination. Rapid antigen tests (RATs) have recently been approved and are now widely used in the current coronavirus disease (COVID-19) pandemic (1). Although the performance of RATs has been evaluated extensively in clinics (2–4), data on postmortem testing are still lacking (5).
We performed a prospective cohort study in which we evaluated the performance of the Roche/SD Biosensor SARS-CoV-2 RAT (https://www.roche.com) in 30 consecutive deceased COVID-19 patients at the University Hospital, Medical University of Graz (Graz, Austria), during November 28–December 23, 2020. We tested each corpse with nasopharyngeal swabs for RAT (using the manufacturer’s kit) and eSwabs (https://www.copanusa.com) for quantitative reverse transcription PCR (qRT-PCR) targeted to the viral envelope (E) and nucleocapsid (N) genes of SARS-CoV-2. Furthermore, we used virus isolation from lung tissue swabs from an additional cohort of deceased COVID-19 patients (n = 11) to compare molecular detection and virus cultivability (Appendix).
All patients were Caucasian, median age was 78 years (range 62–93 years), and 51.2% were female. The median disease duration (interval between the first positive SARS-CoV-2 PCR and death) was 11 days (range 1–43 days). The median postmortem interval (time between death and specimen sampling) was 23 hours (range 8–124 hours; Table; Appendix).
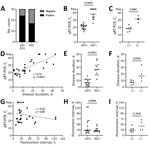
PCR is the current standard for SARS-CoV-2 detection (1,2). In our cohort, qRT-PCR targeted to the E gene showed a higher sensitivity than qRT-PCR for the N gene (Appendix Figure 1). Consequently, we used E gene qRT-PCR as the reference in subsequent evaluations. Results showed that 80% (24/30) of cases were qRT-PCR positive, whereas 56.7% (17/30) were RAT positive (Figure, panel A). RAT had an overall specificity of 100% (95% CI 61%–100%) and an overall sensitivity of 70.8% (95% CI 50.8%–85.1%) when using E gene qRT-PCR as the reference. RAT negative cases showed significantly higher Ct values in qRT-PCR compared with RAT positive cases (mean 38.24 [SD 7.01] vs 20.74 [SD 3.46]; Figure, panel B). Correspondingly, RAT sensitivity increased when cases were stratified according to Ct values (Ct <35, sensitivity 73.9% [95% CI 53.5%–87.5%]; Ct <30, sensitivity 94.4% [95% CI 74.2%–99.7%]; Ct <25, sensitivity 100% [95% CI 80.6%–100%]) (Table; Appendix Table 1). Furthermore, when we compared qRT-PCR results from nasopharyngeal swabs of patients in which viral culture was performed (from corresponding lung tissue swabs of an additional cohort), cultivability was restricted to cases with Ct values <23.7, which is below the threshold of false-negative RAT cases (Ct values >25.8; Figure, panels B, C). These results are in line with most clinical RAT studies that also used virus culture (2–4,6), in which cultivability is exceedingly rare in cases with low viral loads determined with qRT-PCR. We used cultivation from lung tissue swab specimens for this analysis because the lung often shows increased SARS-CoV-2 loads in deceased patients (7; Appendix Table 2) and therefore represents a major infection source during autopsy.
Furthermore, we determined parameters that influenced test performance. We noted a significant positive correlation between disease duration and Ct values (Figure, panel D). Such correlation was also evident in RATs; all cases with disease courses >17 days were RAT negative (Figure, panel E). Postmortem intervals did not correlate with Ct values or RAT results (Figure, panels G, H). Thus, a long disease duration rather than a long postmortem interval seems to be the main factor for increased Ct values and negative RATs. RAT and cultivation results closely mirrored each other with respect to viral load (Figure, panels B, C), disease duration (Figure, panels E, F), and postmortem interval (Figure, panels H, I).
Although RAT had an overall lower sensitivity than qRT-PCR in this study, our data suggest that viral loads of false-negative RAT cases are probably below the threshold of cultivability. Because culture is regarded as a measure of virus viability and infectivity (8), these cases likely pose only minimal risks of SARS-CoV-2 transmission during postmortem examinations. However, each corpse having a postmortem evaluation must be treated as potentially infectious. Even a PCR-negative nasopharyngeal swab specimen does not exclude the presence of viable virus in other body sites, as shown in COVID-19 (7), thus emphasizing the general application of appropriate autopsy safety measures.
In conclusion, RAT should not be seen as a potential replacement for but rather as an addition to current postmortem testing strategies. Especially when qRT-PCR is not readily available, RAT might be useful in selecting the most hazardous corpses that should be examined under special conditions (e.g., Biosafety Level 3 [9]). RAT could therefore be a valuable adjunct tool in guiding autopsy practice.
Dr. Zacharias is a physician-scientist at the Diagnostic and Research Institute of Pathology, Medical University of Graz, Graz, Austria. His main research interests include pulmonary and infectious disease pathology.
References
- Centers for Disease Control and Prevention. Interim guidance for antigen testing for SARS-CoV-2 [cited 2021 Mar 27]. https://www.cdc.gov/coronavirus/2019-ncov/lab/resources/antigen-tests-guidelines.html
- Dinnes J, Deeks JJ, Berhane S, Taylor M, Adriano A, Davenport C, et al.; Cochrane COVID-19 Diagnostic Test Accuracy Group. Rapid, point-of-care antigen and molecular-based tests for diagnosis of SARS-CoV-2 infection. Cochrane Database Syst Rev. 2021;3:
CD013705 .PubMedGoogle Scholar - Albert E, Torres I, Bueno F, Huntley D, Molla E, Fernández-Fuentes MÁ, et al. Field evaluation of a rapid antigen test (Panbio™ COVID-19 Ag Rapid Test Device) for COVID-19 diagnosis in primary healthcare centres. Clin Microbiol Infect. 2021;27:472.e7–10. DOIPubMedGoogle Scholar
- Iglὁi Z, Velzing J, van Beek J, van de Vijver D, Aron G, Ensing R, et al. Clinical evaluation of Roche SD Biosensor rapid antigen test for SARS-CoV-2 in municipal health service testing site, the Netherlands. Emerg Infect Dis. 2021;27. DOIPubMedGoogle Scholar
- Centers for Disease Control and Prevention. Collection and submission of postmortem specimens from deceased persons with confirmed or suspected COVID-19: postmortem guidance [cited 2021 Mar 27]. https://www.cdc.gov/coronavirus/2019-ncov/hcp/guidance-postmortem-specimens.html
- Singanayagam A, Patel M, Charlett A, Lopez Bernal J, Saliba V, Ellis J, et al. Duration of infectiousness and correlation with RT-PCR cycle threshold values in cases of COVID-19, England, January to May 2020. Euro Surveill. 2020;25:
2001483 . DOIPubMedGoogle Scholar - Puelles VG, Lütgehetmann M, Lindenmeyer MT, Sperhake JP, Wong MN, Allweiss L, et al. Multiorgan and renal tropism of SARS-CoV-2. N Engl J Med. 2020;383:590–2. DOIPubMedGoogle Scholar
- Jefferson T, Spencer EA, Brassey J, Heneghan C. Viral cultures for COVID-19 infectious potential assessment—a systematic review. Clin Infect Dis. 2020 Dec 20 [Epub ahead of print]. DOIGoogle Scholar
- Loibner M, Langner C, Regitnig P, Gorkiewicz G, Zatloukal K. Biosafety requirements for autopsies of patients with COVID-19: example of a BSL-3 autopsy facility designed for highly pathogenic agents. Pathobiology. 2021;88:37–45. DOIPubMedGoogle Scholar
- Brown LD, Cai TT, DasGupta A. Interval estimation for a binomial proportion. Stat Sci. 2001;16:101–33. DOIGoogle Scholar
Figure
Table
Cite This ArticleOriginal Publication Date: April 13, 2021
Table of Contents – Volume 27, Number 6—June 2021
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Martin Zacharias, Diagnostic and Research Institute of Pathology, Medical University of Graz, Neue Stiftingtalstraße 6, 8010 Graz, Austria
Top