Volume 27, Number 7—July 2021
Research
Plasmodium falciparum kelch 13 Mutations, 9 Countries in Africa, 2014–2018
Cite This Article
Citation for Media
Abstract
The spread of drug resistance to antimalarial treatments poses a serious public health risk globally. To combat this risk, molecular surveillance of drug resistance is imperative. We report the prevalence of mutations in the Plasmodium falciparum kelch 13 propeller domain associated with partial artemisinin resistance, which we determined by using Sanger sequencing samples from patients enrolled in therapeutic efficacy studies from 9 sub-Saharan countries during 2014–2018. Of the 2,865 samples successfully sequenced before treatment (day of enrollment) and on the day of treatment failure, 29 (1.0%) samples contained 11 unique nonsynonymous mutations and 83 (2.9%) samples contained 27 unique synonymous mutations. Two samples from Kenya contained the S522C mutation, which has been associated with delayed parasite clearance; however, no samples contained validated or candidate artemisinin-resistance mutations.
Malaria remains a serious global health concern, causing ≈405,000 deaths annually, mainly in young children in Africa (1). Although substantial progress has been made over the past decade to reduce the global burden of malaria, several factors threaten these gains, including the emergence and spread of antimalarial drug resistance (1). Artemisinin-based combination therapies (ACTs) are the first-line treatment for uncomplicated malaria caused by Plasmodium falciparum parasites, as recommended by the World Health Organization (WHO) (2). Unfortunately, resistance to ACTs (i.e., delayed parasite clearance and clinical treatment failures) has emerged in the Greater Mekong Subregion of Southeast Asia, posing a considerable risk to malaria control in the region (3). Even though clinical resistance to ACTs has not been reported in Africa (1), the threat of its emergence remains.
As part of antimalarial therapeutic efficacy activities, WHO recommends molecular surveillance of the P. falciparum kelch 13 gene (Pfk13) (with focus on the propeller domain region), a molecular marker associated with delayed clearance of parasitemia after therapy with artemisinin monotherapy or an ACT (3–7). Because specific single-nucleotide polymorphisms (SNPs) within the propeller domain region of Pfk13 continue to be discovered, WHO continues to update a list of these SNPs on the basis of association with delayed parasite clearance and reduced in vitro drug susceptibility (Table 1). Nine SNPs are currently considered validated by WHO to have delayed parasite clearance and in vitro data demonstrating partial resistance to artemisinin (3). WHO categorized 11 SNPs as candidate mutations, correlated with delayed parasite clearance but not validated with in vitro data (3). An additional 11 SNPs are listed by WHO as associated with delayed parasite clearance but without statistical significance because of limited data (3).
WHO recommends that malaria-endemic countries perform therapeutic efficacy studies (TESs) every 2 years to evaluate antimalarial treatments currently used in a particular region (8). Surveillance for molecular markers associated with antimalarial resistance is a recommended part of a TES to detect the presence of mutations associated with resistance (8). As part of the US President’s Malaria Initiative, the Centers for Disease Control and Prevention (CDC) and the US Agency for International Development provide support to countries in Africa to perform TESs, including molecular characterization of antimalarial-resistance markers, through the PMI-supported Antimalarial Resistance Monitoring in Africa (PARMA) network (9). Established in 2015, this endeavor involves laboratory trainees in Africa who bring TES samples from their home country to the CDC (Atlanta, Georgia, USA) to receive advanced laboratory training and perform molecular testing for antimalarial-resistance mutations (9). In this article, we report Pfk13 mutation data generated from samples analyzed and collected from TESs conducted in 9 countries in Africa during 2014–2018.
Samples, Ethics Statement, and TES Protocols
Before initiation, all work described in this article was approved by the respective institutional ethics review committee in each country and the Office of the Associate Director of Science of CDC’s Center for Global Health and assigned the following tracking numbers: 2014–233a and 2014–233b (Angola), 2017–141 (Benin), 2018–035 (DRC), 2016–046 (Guinea), 6696.0 (Kenya), 6029.0 (Malawi), 2016–012a (Mali), 2015–073a (Tanzania), and 2016–200 (Zambia). Dried blood spots were collected from TESs conducted in 9 countries in Africa (Angola, Benin, the Democratic Republic of the Congo [DRC], Guinea, Kenya, Malawi, Mali, Tanzania, and Zambia; Table 2) during 2014–2018. The samples included those obtained pretreatment (at day of enrollment) and at day of treatment failure. Day of treatment failure samples came from patients experiencing a recrudescence or new infection during the follow-up period of (usually ending at 28 or 42 days) after administration of an ACT. TES and antimalarial molecular marker results for some of the data analyzed have been previously published for Angola (10–12), Kenya (13), and Tanzania (14). Results might differ slightly from previously published works because those works might not have reported results from all samples, might not have reported mutations in mixed infections, or might not have reported synonymous mutation results. Our study was a reanalysis of all available sequences using the same sequence data analysis quality filters, cut-offs, and quality scores for all countries.
Sequencing of Pfk13 Propeller Domain Region
We extracted DNA from dried blood spots using the QIAamp Blood DNA Kit (QIAGEN, https://www.qiagen.com) according to the manufacturer’s instructions. We amplified the propeller domain region from codon positions 389–649 by PCR and Sanger sequenced according to methods previously described (15).
Data Analysis
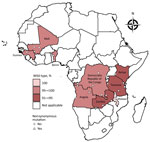
We analyzed sequence data by using Geneious Prime (Biomatters, https://www.geneious.com). We trimmed and quality filtered forward and reverse sequence reads for each sample (error probability limit 0.05, maximum low-quality bases 30) from the 3′ and 5′ ends to remove low-quality bases. We aligned trimmed sequences to the Pfk13 National Center for Biotechnology Information gene reference no. PF3D7_1343700 (https://www.ncbi.nlm.nih.gov/gene/814205) and assessed for SNPs. We only considered SNPs if they had a Phred quality score of >30 and were present in both forward and reverse strands. Mixed infections were detected by using the heterozygous caller plug-in tool in Geneious with a threshold of >30%. A second analyst confirmed all SNP and heterozygous calls by manual technical review. We submitted all Pfk13 sequences with SNPs reported in this study to GenBank (accession nos. MN072940–3042). We used R software version 4.0.1 (R Foundation for Statistical Computing, https://www.r-project.org) to generate a map showing the distribution of mutations in the 9 countries (Figure).
We attempted Pfk13 sequencing on 3,248 samples (2,579 pretreatment and 669 day of failure samples) from the 9 countries (Table 2); 2,865 were successfully sequenced (Table 3). Of those, 2,753 samples were wild-type. A total of 11 unique nonsynonymous mutations and 27 unique synonymous mutations were detected in 2,865 successfully sequenced pretreatment and day of failure samples from Angola, Benin, DRC, Guinea, Kenya, Malawi, Mali, Tanzania, and Zambia collected during 2014–2018 (Figure, Table 4; Appendix 1).
Of the 2,303 sequenced pretreatment samples, 2,213 were wild-type and 90 (3.9%) contained mutations (Table 3). Of the 90 pretreatment samples with mutations, 10 unique nonsynonymous mutations were present in 25 samples from 8 of the 9 countries assessed (Table 4) and 25 unique synonymous mutations were present in 65 samples from 8 of the 9 countries assessed (Appendix 1 Table 1). Two samples from Kenya contained the S522C mutation, reported by WHO as a less-frequent mutation associated with delayed parasite clearance but without statistical significance because of limited data (3). Both of these patients cleared their initial infection. A578S, the most commonly found mutation in Africa (not associated with resistance) (3), was the most common nonsynonymous mutation we identified. The mutation was found in 14 pretreatment isolates: 4 in Angola, 1 in DRC, 1 in Mali, 6 in Kenya, 1 in Tanzania, and 1 in Zambia (Table 4). No mutations were identified in the samples from Malawi. Eight of the 10 unique nonsynonymous mutations in the pretreatment samples have been reported previously in other countries, whereas 2 mutations, P419S (Guinea) and Q613R (Angola), were newly identified in our study. No WHO-validated or candidate Pfk13 mutations were identified.
Of the 669 day of failure samples, 562 were successfully sequenced; 107 (16.0%) samples failed to amplify, produced poor-quality sequences, or both (Table 3). A total of 540 samples were wild-type. Two nonsynonymous mutations were found in 4 day of failure samples (Table 4) and 10 synonymous mutations (Appendix 1 Table 2) were identified in 18 day of failure samples from 4 countries. Of the nonsynonymous mutations in day of failure samples, 2 samples from Kenya and 1 sample from DRC contained the A578S mutation, and 1 sample from DRC contained the S477Y mutation (Table 4). We compiled the complete results of the sequence data reanalysis (Appendix 2).
This work provides an update on Pfk13 genetic markers in 9 countries in Africa with endemic malaria. Although clinical resistance to ACTs has yet to be confirmed in Africa (1), the early detection of Pfk13 mutations through surveillance allows for swift action before resistance spreads widely. To date, all WHO-validated SNPs detected in Africa have been the result of independent emergence as opposed to spreading through imported cases from Southeast Asia (21). More than 200 Pfk13 mutations have been identified in global samples (3,18,21), and >74 Pfk13 nonsynonymous mutations have been reported in Africa (22,23). In this study, we report the presence of S522C in Kenya, a less frequent mutation that has been previously reported to be associated with delayed parasite clearance but lacking sufficient evidence to be considered a WHO-validated or candidate mutation (3).
As more molecular surveillance data are collected, previous results should be reinterpreted to determine the presence of WHO-reportable mutations because the importance of these mutations in drug resistance might change based on new data (3,24). Although we report only 1 mutation identified by WHO to possibly play a role in resistance, other detected mutations, such as the other nonsynonymous mutations with unknown resistance status reported in this study, might be deemed important in the future as more data are collected and validated. In 2017, WHO categorized only 5 mutations as validated (N458Y, Y493H, R539T, I543T, and 580Y) (24), but in 2018 the validated list was updated to include an additional 4 mutations, including F446I, P553L, and R561H (formerly candidate markers) and M476I (formerly reported as a less frequent variant associated with in vivo or in vitro test results) (3). In addition, the Worldwide Antimalarial Resistance Network tracks Pfk13 mutations worldwide and strives to detect new associations of mutations with delayed parasite clearance, which might inform WHO classifications (18).
We report the presence of 11 unique nonsynonymous mutations in Angola, Benin, Guinea, DRC, Kenya, Mali, Tanzania, and Zambia; all were previously reported in the literature (Table 4) except P419S and Q613R. The most common nonsynonymous mutation observed in our study was A578S, a nonsynonymous mutation frequently described in Africa (3) and, to a lesser extent, Asia (e.g., Thailand [19] and Bangladesh [25]). WHO has reported that A578S is not associated with partial artemisinin resistance (3). Most mutations detected were synonymous mutations consistent with previous reports (21). Because synonymous mutations do not result in an amino acid change, they are not associated with resistance. Parasites from Africa have been shown to have a higher prevalence of synonymous mutations, which is not surprising given that P. falciparum originated in Africa and continues to have a high level of transmission in this region (19).
The results described in this article represent the collaborative output of the PARMA network, which originated in 2015 with the objectives of assisting countries in Africa in testing malaria samples from TESs for genetic markers associated with antimalarial resistance and supporting training and capacity building of collaborators in Africa (9). In 8 of the 9 countries included in this report (all but Angola), the Pfk13 results were generated during a 6–8-week visit to CDC by trainees from a laboratory in the country where the TES was performed. Results were subsequently shared by the trainee’s laboratory with their national malaria control program and other local stakeholders to make decisions related to antimalarial use. Although the Pfk13 results we have described would not be cause for alarm or policy change, recent findings in Rwanda suggests a substantial presence of the Pfk13 R561H mutation (26) that has evolved locally, highlighting the importance of molecular surveillance for early detection of emerging patterns of resistance. In this context, PARMA training visits generate a vast amount of data from TES samples, ranging from efficacy results to prevalence of other molecular markers (e.g., P. falciparum multidrug-resistant protein 1 and P. falciparum chloroquine-resistance transporter) to the presence of P. falciparum histidine-rich protein 2 and 3 deletions (which might affect rapid diagnostic test performance). Generating phenotypic (i.e., efficacy) and genotypic data on the same sample provides an opportunity to identify novel mutations associated with resistance and enables detection of known mutations in samples with well-characterized efficacy outcomes. Because the PARMA network encourages standardization of laboratory methods and data reporting, such explorations might detect trends over time in a single country or produce insightful observations by using data from multiple countries. With the increased use of next-generation sequencing, the PARMA network has embarked on applying these principles of data generation, capacity building, networking, and standardization to this emerging technology (27). The ultimate goal is laboratories in Africa independently analyzing their own malaria samples.
Dr. Schmedes is lead bioinformatician at the Florida Department of Health; her primary interests include developing and implementing bioinformatics methods for studying bacterial and viral pathogens of public health significance in the state of Florida. Dr. Talundzic is an informatics health scientist in the Center for Global Health at CDC; his primary interests include developing and implementing next generation sequencing and bioinformatics methods for studying Plasmodium parasites.
Acknowledgment
This work was made possible with funding provided by the US President’s Malaria Initiative (PMI) through PARMA. We also acknowledge partial support from the Advanced Molecular Detection Initiative at the CDC and partial support by the Bioinformatics Fellowship Program administered by the Association of Public Health Laboratories and funded by CDC. S.E.S. was supported by the Bioinformatics Fellowship Program. J.K. was supported in part by the CDC Foundation. D.P. was employed by Williams Consulting LLC, which provided support in the form of salary for D.P. D.S.I. was partly supported by the Developing Excellence in Leadership and Genetics Training for Malaria Elimination (DELGEME) in Sub-Saharan Africa program through the Developing Excellence in Leadership, Training and Science Africa Initiative (DELGEME grant no. 107740/Z/15/Z). The Developing Excellence in Leadership, Training and Science Africa Initiative is an independent funding scheme of the African Academy of Sciences’ Alliance for Accelerating Excellence in Science in Africa and is supported by the New Partnership for Africa’s Development Planning and Coordinating Agency with funding from the Wellcome Trust (DELGEME grant no. 107740/Z/15/Z) and government of the United Kingdom. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- World Health Organization. World malaria report 2019 [cited 2020 May 15]. https://www.who.int/publications/i/item/9789241565721
- World Health Organization. Guidelines for the treatment of malaria. 3rd edition. 2015 [cited 2020 May 15]. https://apps.who.int/iris/handle/10665/162441
- World Health Organization. Artemisinin resistance and artemisinin-based combination therapy efficacy: status report. 2018 [cited 2020 May 15]. https://apps.who.int/iris/handle/10665/274362
- Cheeseman IH, Miller BA, Nair S, Nkhoma S, Tan A, Tan JC, et al. A major genome region underlying artemisinin resistance in malaria. Science. 2012;336:79–82. DOIPubMedGoogle Scholar
- Takala-Harrison S, Clark TG, Jacob CG, Cummings MP, Miotto O, Dondorp AM, et al. Genetic loci associated with delayed clearance of Plasmodium falciparum following artemisinin treatment in Southeast Asia. Proc Natl Acad Sci U S A. 2013;110:240–5. DOIPubMedGoogle Scholar
- Ariey F, Witkowski B, Amaratunga C, Beghain J, Langlois A-C, Khim N, et al. A molecular marker of artemisinin-resistant Plasmodium falciparum malaria. Nature. 2014;505:50–5. DOIPubMedGoogle Scholar
- Ashley EA, Dhorda M, Fairhurst RM, Amaratunga C, Lim P, Suon S, et al.; Tracking Resistance to Artemisinin Collaboration (TRAC). Spread of artemisinin resistance in Plasmodium falciparum malaria. N Engl J Med. 2014;371:411–23. DOIPubMedGoogle Scholar
- World Health Organization. Methods for surveillance of antimalarial drug efficacy. 2009 [cited 2020 May 15]. https://www.who.int/malaria/publications/atoz/9789241597531/en
- Halsey ES, Venkatesan M, Plucinski MM, Talundzic E, Lucchi NW, Zhou Z, et al. Capacity Development through the US President’s Malaria Initiative-Supported Antimalarial Resistance Monitoring in Africa Network. Emerg Infect Dis. 2017;23:S53–6. DOIPubMedGoogle Scholar
- Plucinski MM, Dimbu PR, Macaia AP, Ferreira CM, Samutondo C, Quivinja J, et al. Efficacy of artemether-lumefantrine, artesunate-amodiaquine, and dihydroartemisinin-piperaquine for treatment of uncomplicated Plasmodium falciparum malaria in Angola, 2015. Malar J. 2017;16:62. DOIPubMedGoogle Scholar
- Ljolje D, Dimbu PR, Kelley J, Goldman I, Nace D, Macaia A, et al. Prevalence of molecular markers of artemisinin and lumefantrine resistance among patients with uncomplicated Plasmodium falciparum malaria in three provinces in Angola, 2015. Malar J. 2018;17:84. DOIPubMedGoogle Scholar
- Davlantes E, Dimbu PR, Ferreira CM, Florinda Joao M, Pode D, Félix J, et al. Efficacy and safety of artemether-lumefantrine, artesunate-amodiaquine, and dihydroartemisinin-piperaquine for the treatment of uncomplicated Plasmodium falciparum malaria in three provinces in Angola, 2017. Malar J. 2018;17:144. DOIPubMedGoogle Scholar
- Chebore W, Zhou Z, Westercamp N, Otieno K, Shi YP, Sergent SB, et al. Assessment of molecular markers of anti-malarial drug resistance among children participating in a therapeutic efficacy study in western Kenya. Malar J. 2020;19:291. DOIPubMedGoogle Scholar
- Ishengoma DS, Mandara CI, Francis F, Talundzic E, Lucchi NW, Ngasala B, et al. Efficacy and safety of artemether-lumefantrine for the treatment of uncomplicated malaria and prevalence of Pfk13 and Pfmdr1 polymorphisms after a decade of using artemisinin-based combination therapy in mainland Tanzania. Malar J. 2019;18:88. DOIPubMedGoogle Scholar
- Talundzic E, Chenet SM, Goldman IF, Patel DS, Nelson JA, Plucinski MM, et al. Genetic analysis and species specific amplification of the artemisinin resistance-associated kelch propeller domain in P. falciparum and P. vivax. PLoS One. 2015;10:
e0136099 . DOIPubMedGoogle Scholar - Huang B, Deng C, Yang T, Xue L, Wang Q, Huang S, et al. Polymorphisms of the artemisinin resistant marker (K13) in Plasmodium falciparum parasite populations of Grande Comore Island 10 years after artemisinin combination therapy. Parasit Vectors. 2015;8:634. DOIPubMedGoogle Scholar
- Voumbo-Matoumona DF, Kouna LC, Madamet M, Maghendji-Nzondo S, Pradines B, Lekana-Douki JB. Prevalence of Plasmodium falciparum antimalarial drug resistance genes in Southeastern Gabon from 2011 to 2014. Infect Drug Resist. 2018;11:1329–38. DOIPubMedGoogle Scholar
- WWARN K13 Genotype-Phenotype Study Group. Association of mutations in the Plasmodium falciparum Kelch13 gene (Pf3D7_1343700) with parasite clearance rates after artemisinin-based treatments-a WWARN individual patient data meta-analysis. BMC Med. 2019;17:1. DOIPubMedGoogle Scholar
- Ménard D, Khim N, Beghain J, Adegnika AA, Shafiul-Alam M, Amodu O, et al.; KARMA Consortium. KARMA Consortium. A worldwide map of Plasmodium falciparum K13-propeller polymorphisms. N Engl J Med. 2016;374:2453–64. DOIPubMedGoogle Scholar
- Laminou I, Lamine M, Arzika I, Mahamadou B, Gora D, Dieye A. Detection of Plasmodium falciparum K13 propeller A569G mutation after artesunate-amodiaquine treatment failure in Niger. J Adv Biol Biotechnol. 2018;18:1–8. DOIGoogle Scholar
- Malaria GEN. Plasmodium falciparum Community Project. Genomic epidemiology of artemisinin resistant malaria. eLife. 2016;5:
e08714 . DOIGoogle Scholar - Ocan M, Akena D, Nsobya S, Kamya MR, Senono R, Kinengyere AA, et al. K13-propeller gene polymorphisms in Plasmodium falciparum parasite population in malaria affected countries: a systematic review of prevalence and risk factors. Malar J. 2019;18:60. DOIPubMedGoogle Scholar
- Conrad MD, Rosenthal PJ. Antimalarial drug resistance in Africa: the calm before the storm? Lancet Infect Dis. 2019;19:e338–51. DOIPubMedGoogle Scholar
- World Health Organization. Artemisinin and artemisinin-based combination therapy resistance: status report. 2017 [cited 2020 May 15]. https://apps.who.int/iris/handle/10665/255213
- Mohon AN, Alam MS, Bayih AG, Folefoc A, Shahinas D, Haque R, et al. Mutations in Plasmodium falciparum K13 propeller gene from Bangladesh (2009-2013). Malar J. 2014;13:431. DOIPubMedGoogle Scholar
- Uwimana A, Legrand E, Stokes BH, Ndikumana JM, Warsame M, Umulisa N, et al. Emergence and clonal expansion of in vitro artemisinin-resistant Plasmodium falciparum kelch13 R561H mutant parasites in Rwanda. Nat Med. 2020;26:1602–8. DOIPubMedGoogle Scholar
- Talundzic E, Ravishankar S, Kelley J, Patel D, Plucinski M, Schmedes S, et al. Next-generation sequencing and bioinformatics protocol for malaria drug resistance marker surveillance. Antimicrob Agents Chemother. 2018;62:e02474–17. DOIPubMedGoogle Scholar
Figure
Tables
Cite This ArticleOriginal Publication Date: June 07, 2021
1These authors contributed equally to this article.
Table of Contents – Volume 27, Number 7—July 2021
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Eldin Talundzic, Centers for Disease Control and Prevention, 1600 Clifton Rd NE, Mailstop H23-10, Atlanta, GA 30329-4018, USA
Top