Volume 3, Number 4—December 1997
THEME ISSUE
Foodborne
Controlling Emerging Foodborne Microbial Hazards
Epidemiology and Detection as Options for Control of Viral and Parasitic Foodborne Disease
Cite This Article
Citation for Media
Abstract
Human enteric viruses and protozoal parasites are important causes of emerging food and waterborne disease. Epidemiologic investigation and detection of the agents in clinical, food, and water specimens, which are traditionally used to establish the cause of disease outbreaks, are either cumbersome, expensive, and frequently unavailable or unattempted for the important food and waterborne enteric viruses and protozoa. However, the recent introduction of regulatory testing mandates, alternative testing strategies, and increased epidemiologic surveillance for food and waterborne disease should significantly improve the ability to detect and control these agents. We discuss new methods of investigating foodborne viral and parasitic disease and the future of these methods in recognizing, identifying, and controlling disease agents.
Emerging infectious diseases can be defined as infections that have newly appeared in a population or have existed but are rapidly increasing in incidence or geographic range (1). Agents for which a particular route of transmission is newly recognized and agents (previously unidentifiable) that are now known because of advances in detection methods should also be included in this definition. Advances in epidemiologic and detection methods during the last 10 to 20 years have placed food and waterborne human enteric viruses and protozoal parasites within this category.
Human enteric viruses and protozoa are parasitic agents that replicate in the intestines of infected hosts and are excreted in the feces. In general, the viruses are limited to human hosts, while the parasitic agents (in the form of cysts or oocytes) have a variety of human and nonhuman animal hosts. Both are transmitted primarily by the fecal-oral route, and as a result, the major source of contamination for foods and water is through contact with human and animal fecal pollution. This contamination may occur directly, through contaminated meat carcasses or poor personal hygiene practices of infected food handlers, or indirectly, through contact with fecally contaminated water or other cross-contamination routes. Viruses and parasites differ from foodborne bacterial pathogens in important ways. Because they are environmentally inert, they do not replicate in food, water, or environmental samples. Additionally, unlike bacterial pathogens, human enteric viruses and protozoal parasites are environmentally stable (2), are resistant to many of the traditional methods used to control bacterial pathogens (2), and have notably low infectious doses (3,4). This allows virtually any food to serve as a vehicle for transmission and enables these agents to withstand a wide variety of commonly practiced food storage and processing conditions (2,5).
Human Enteric Viruses
Human enteric viruses are increasingly recognized as important causes of foodborne illness. A recent report issued by the Council for Agricultural Science and Technology ranked human enteric viruses as fifth and sixth among identified causes of foodborne disease in the United States (6). A review of U.S. national surveillance data for 1979 showed that 14 (44%) of 32 foodborne disease outbreaks in institutional settings were epidemiologically typical of viral gastroenteritis (7). Viral gastroenteritis was reported as the most common foodborne illness in Minnesota from 1984 to 1991, predominantly associated with poor personal hygiene of infected food handlers (8). Furthermore, recent data indicate that 10% of the 4,617 outbreaks of foodborne disease of unconfirmed etiology reported from 1973 to 1987 met at least two of the clinical criteria for outbreaks of acute viral gastroenteritis (8,9). The apparent failure to confirm a viral etiology in such outbreaks has been due largely to the lack of available tests and the reluctance of public health officials to use epidemiologic criteria in the classification of foodborne viral disease (9-11). The unavailability of food specimens and the failure to report outbreaks of mild gastrointestinal disease have also contributed to reporting difficulties. All of these factors have resulted in a drastic underestimate of the true scope and importance of foodborne viral infection (8).
The most common types of foodborne viral disease are infectious hepatitis due to hepatitis A virus and acute viral gastroenteritis associated with the Norwalk agent and other related small, round-structured gastrointestinal viruses (12). The human enteroviruses may also be transmitted by foodborne routes (13) and are the most commonly isolated agents in surveys of naturally occurring viral contamination in foods (14). Foodborne outbreaks due to small round viruses, parvoviruses, and astroviruses are occasionally reported (15). Rotaviruses, some adenoviruses, and hepatitis E virus are important causes of waterborne disease outbreaks, particularly in developing countries (12). Foodborne outbreaks associated with human enteric viruses are almost always due to the consumption of fecally contaminated raw or undercooked shellfish and ready-to-eat products contaminated by infected food handlers (12). Postrecovery and secondary transmission are a concern (16,17). Recent outbreaks are summarized in Table 1 (18-24).
Parasitic Protozoa
Since 1981, enteric protozoa have become the leading cause of waterborne disease outbreaks for which an etiologic agent could be determined (5). A recent study reported that 21% of drinking water-associated outbreaks between 1991 and 1992 were attributable to parasitic agents (25). Furthermore, these agents are frequent contaminants of potable water supplies (26,27). The potential for transmission of these agents by foodborne routes is increasingly recognized (28). For instance, from 1988 to 1992, seven food-associated outbreaks of giardiasis, comprising 184 cases, were reported in the United States (29). However, since foodborne transmission is only recently documented and more than half of all reported foodborne disease outbreaks have undetermined etiology, the true importance of foodborne transmission of parasitic protozoa is unknown.
The most common human enteric parasitic infections in the United States are caused by Cryptosporidium parvum and Giardia lamblia. Cyclospora is also an emerging enteric protozoon that has recently been associated with the consumption of contaminated fruit (30). Large communitywide waterborne outbreaks of parasitic protozoa are usually associated with surface water supplies that are either unfiltered or subjected to inadequate flocculation and filtration (5). Two large waterborne outbreaks have occurred in the United States within the last 10 years (31,32); one of these was the largest recorded waterborne disease outbreak in U.S. history (32).
Limitations
Most of the information about viral and parasitic food and waterborne disease comes from outbreak investigations by state and local health departments and surveillance programs directed by the Centers for Disease Control and Prevention (CDC). However, since many of these diseases are not reportable and surveillance is based on voluntary reporting by state health departments, the magnitude of this disease problem is underestimated. This is exacerbated by a reluctance to use epidemiologic criteria in the classification of foodborne viral disease and the failure to report mild outbreaks of gastrointestinal disease. Furthermore, since investigation generally follows an outbreak, important information and samples may have been destroyed, consumed, or lost to inaccurate recall. Since many of these outbreaks are small and confined, epidemiologic investigation may be limited by the resources available to state and local health departments. Difficulties in investigating and reporting are further complicated by the fact that the enteric protozoa cause common opportunistic infections in the immunocompromised, and the role of foods in these diseases has not been studied. Likewise, the role of the foodborne transmission route in sporadic disease and the importance of carrier states and secondary illness after a primary foodborne disease outbreak are poorly characterized.
Clinical Samples
Illness caused by human enteric viruses can be suspected epidemiologically by considering incubation period and illness duration analysis, classic viral gastroenteritis symptoms, and the absence of bacterial or parasitic pathogens in stool samples (10,11). Laboratory confirmation of human enteric viral infection has been based on a rise in specific antibody to the virus, or alternatively, the demonstration of virus particles, antigen, or nucleic acid in stools. The detection methods most often applied to clinical samples have included immune electron microscopy, radioimmunoassay, and enzyme immunoassay (33). The usefulness of these assays has been reduced by low detection limits (>104-105 particles/ml) and the inability to cultivate Norwalk-like viruses in vitro, which has limited the supply of viral antigen available for developing reagents (8). In addition, the Norwalk agent is only one of several small round-structured gastrointestinal viruses that cause outbreaks with similar clinical and epidemiologic features (8).
Before 1981, parasitic disease in humans was diagnosed histologically by identifying the life cycle stages of parasitic agents in the intestinal mucosa (34). More recently, clinical diagnosis has involved methods to concentrate parasitic agents from stool specimens followed by a variety of fluorescent or immunofluorescent staining techniques and subsequent microscopic examination (34). Serodiagnostic methods have been developed (35,36), but additional evaluations are needed to confirm the diagnostic utility of these methods.
Environmental, Food, and Water Samples
Failure to confirm a viral and parasitic etiology in foodborne outbreaks has also been due to the lack of adequate methods to detect the causative agents in environmental samples. Like bacterial pathogens in food and water, viruses and parasites are frequently present in small numbers. However, unlike traditional food microbiologic techniques, which have relied on cultural enrichment and selective plating to increase cell numbers and differentiate pathogens in background microflora, techniques to detect human enteric viruses and parasitic protozoa require live mammalian cells for growth. For this reason, standard methods to detect enteric bacteria in foods cannot be used; instead, detection requires an initial concentration step, often from large volumes of food or water, followed by mammalian cell culture infectivity assay or immunofluorescent staining. Concentration methods are usually cumbersome, and yields are less than optimal. Both cell culture infectivity and immunofluorescent staining are expensive and slow and require highly trained personnel. Furthermore, mammalian cell culture lines are largely unavailable for the epidemiologically important foodborne viruses. Alternative detection methods based on immunologic and molecular methods have been reported; recent methodologic developments have focused on overcoming barriers to detection, such as improving recovery efficiencies and detection limits and preventing inhibitions due to food-related compounds.
Concentration
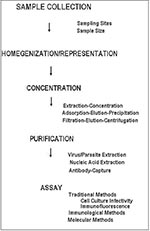
Two general schemes for the concentration of human enteric viruses from foods have been reported: extraction-concentration methods and adsorption-elution-concentration methods (Figure). The general purpose of concentration is to provide a high recovery of infectious virus in a low-volume aqueous solution free of cytotoxic materials. Both schemes employ conditions favoring the separation of viruses from shellfish tissues (most of the developmental work has used shellfish as model food commodities), primarily through the use of filtration, precipitation, polyelectrolyte flocculation, and solvent extraction. While either method can be used, adsorption-elution-precipitation methods have been favored in recent years (2,37). Virus yields after concentrations are 10% to 90% (38).
Detection
Traditional methods to directly detect viruses in foods after concentration have been based on the ability of enteric viruses to infect live mammalian cells in culture. Quantal and enumerative methods using a variety of mammalian cell culture lines, generally from primate kidneys, have been reported. Such approaches have been limited because levels of contaminating virus generally are low (1-200 infectious units per 100 grams of shellfish) (39), residual food components interfere with assays (38), and the epidemiologically important viruses do not replicate (small round-structured gastrointestinal viruses) or replicate poorly (hepatitis A virus) in mammalian cell culture (2). Alternative methods such as enzyme-linked immunosorbent assay (ELISA) and DNA/RNA probes have been reported but are limited by high detection limits (>103 infectious units), unavailability of reagents, and poor sample quality (2). These difficulties are illustrated by the confirmation of viral contamination in a food in only two reported instances (18,19).
The in vitro enzymatic amplification method of polymerase chain reaction (PCR) offers an opportunity to enrich a single specific nucleic acid sequence up to a millionfold and hence provides a sensitive and specific method with a theoretical detection limit of one virus unit. This method is readily adaptable to the detection of RNA viruses by preceding the PCR with a brief reverse transcription (RT) step, hence the designation RT-PCR. The recent cloning of the Norwalk agent and related small round-structured gastrointestinal viruses has provided an opportunity to develop effective molecular detection methods for these previously nondetectable agents (40,41).
The application of PCR methods to the detection of human enteric viruses in foods is an area of active research. However, the development of such methods is complicated by low levels of contamination, high sample volumes, and the presence of food components, which may interfere with enzymatic amplification reactions. To address these issues, three alternative approaches have been used to simultaneously reduce sample volumes and the level of interfering compounds. The most frequently applied approach involves isolating and purifying nucleic acids (RNA) from the food sample before RT-PCR (42-47). A second approach combines capture of the virus with specific antibody followed by nucleic acid amplification by using RT-PCR (19,48). In the third approach, the intact virus particle is concentrated and purified from the complex food matrix resulting in sample volume reduction and removal of inhibitors, followed by subsequent heat release of viral nucleic acid from the virion capsid and RT-PCR (49-51). All three methods have been applied to various shellfish species, and in some cases, to other food commodities and naturally contaminated field shellfish specimens. The methods are summarized in Table 2, along with optimized virus detection levels (19,42-51). A combined approach was reported by Chung et al. (51), who successfully detected human enteroviruses and hepatitis A virus in naturally contaminated oyster samples after viral amplification in mammalian cell cultures. Despite enormous strides in the ability to detect human enteric viruses with PCR, the technique is still limited by the absence of effective concentration methods, the presence of enzymatic inhibitors, and the inability to distinguish between infectious and noninfectious virions.
Concentration
Like viruses, parasitic protozoa are usually present in low concentrations in contaminated water and hence must be concentrated from large volumes of water before detection (Figure). Giardia cysts and Cryptosporidium oocysts are concentrated from 100 to 1,000 liters of water by filtration through yarn-wound filters. Retained particulates are eluted from the filters and reconcentrated by centrifugation. The pelleted cysts and oocysts are then separated from particulate debris by flotation on Percoll-sucrose gradients, followed by subsequent detection. With this method, a concentrate of less than 5 ml can be provided for final detection (52). While recovery efficiencies as high as 100% have been reported (53), recovery is generally poor and greatly affected by water quality and particulate matter (53,54). Alternative methods using calcium carbonate precipitation can concentrate Cryptosporidium oocysts from water with concentration efficiencies as high as 63% (55).
Detection
Traditional methods to detect parasitic agents from water sample concentrates have been based on immunofluorescent staining of filtered sample concentrates (52). The sample concentrate is filtered through cellulose acetate filters and commercial kits that use fluorescein isothiocyante-labeled monoclonal antibodies applied for immunofluorescent staining. The stained filters are examined under an ultraviolet microscope, and cysts and oocysts are classified according to immunofluorescence, size, shape, and internal morphologic characteristics. The results are reported as presumptive and confirmed cysts and oocysts per 100 liters of water (52). Confirmation is based on the ability to visualize organelles under light microscopy. This method is extremely limited because it is time-consuming and expensive, requires highly skilled personnel, and does not indicate viability of the cysts or oocysts; in addition, cross-reactions of monoclonal antibodies with algal cells and debris interfere with the interpretation of results (56).
Because of the limitations of immunofluorescence assays, alternative strategies for the detection of protozoal parasites in environmental and water samples are being sought. Most of the approaches use the traditional methods of concentration, in conjunction with alternative detection methods. In many cases, the inclusion/exclusion of fluorogenic dyes is used to enhance morphologic examination; in particular, 4,6-diamidino-2-phenyl indole and propidium iodide have been used to facilitate detection and also assess viability (57). Two recent methodologic developments include cell sorting/particle counting approaches and molecular approaches. In many cases, a combination of approaches is used. Particle-counting approaches include the Fluorescence-Activated Cell Sorting system, a laser-based particle counter that is able to simultaneously sort particles, sense fluorescence, and determine size (58). This system is being used to detect Giardia and Cryptosporidium in water samples in England and Australia (55). Differentially stained cysts and oocysts may be visualized microscopically with cooled charge couple devices (58,59). Molecular approaches that apply DNA hybridization to the detection of Giardia species have been reported (60). A method that combines fluorescence and in situ hybridization with confocal microscopy has been applied to both detect and speciate Giardia cysts (61). PCR methods have been developed to detect Giardia and Cryptosporidium (62-64). While PCR methods have the potential to detect one single infectious unit and may be applied to discriminate pathogenic from nonpathogenic species, they remain limited because of enzymatic inhibition, the inability to discriminate between viable and nonviable organisms, and the current absence of quantitative assay.
Several methods in development are combinations of multiple detection approaches. A combined method, designated the electrorotation assay, couples filtration and subsequent elution with affinity immunocapture by using paramagnetic beads. By inducing an electric field on a microscope with a special stage attachment, the organism-bead complexes rotate in a characteristic pattern that enables detection of parasitic protozoa (52). Several investigators are also working on methods that couple cell culture infectivity with immunostaining, thereby providing detection with simultaneous indication of viability (M. Sobsey, pers. comm.). Clinical ELISA kits have been evaluated for use in environmental water samples with reported detection limits of fewer than 10 cysts or oocyts (65); however, cross-reaction with algae continues to be a problem. Emerging detection approaches are summarized in Table 3 (58-64). Methods to concentrate and detect parasitic protozoa specifically from foods are under development.
Considerations
Although prototype alternative and rapid methods to detect human enteric viruses and parasitic protozoa in foods and water have been reported, multiple barriers must be overcome before these methods are applicable to routine monitoring. To obtain sample representation and detection sensitivity adequate for the low levels of contamination found with these enteric pathogens in naturally contaminated environmental specimens, large sample volumes of food and water need to be processed. While some of the detection methods begin with large samples, many do not consider sample size, which limits the sensitivity of the assay procedure from the beginning. The approaches then applied to concentrate and purify the pathogens from the samples do not only need to be reasonably efficient but also need to produce a concentrate low in volume and free of inhibitory compounds. To complicate matters further, different food commodities have differing types and levels of inhibitors, many of which have remained recalcitrant to almost all removal processes.
The relationship between detection by molecular and immunologic approaches and the subsequent viability or infectivity of these enteric pathogens remains a concern. While investigators have addressed these issues using in vitro excystation, inclusion and exclusion of vital dyes, animal infectivity (parasitic protozoa) (57,66,67), and assays combining mammalian cell culture infectivity with alternate detection strategies (viruses) (51; M. Sobsey, pers. comm.), these methods are not useful for the nonculturable human enteric viruses; they are expensive, time-consuming, and not readily amenable to routine diagnostic work. Establishing quantitative detection methods also needs further research.
The development of detection methods is also limited by the current state of knowledge. For instance, while recent sequencing evidence indicates that the Norwalk-like small round-structured gastrointestinal virus group consists of multiple members having the same physical and genomic characteristics as other viruses in the family Caliciviridae (40,41), considerable sequence and antigenic diversity remain among the members of this group (68-70). While PCR primers for these genetically diverse agents have been reported (46,71), development of universal detection methods is clearly limited until more complete information about this group of human enteric viruses is available.
Research is needed to develop and refine the prototype protocols into collaboratively tested methods that could be routinely and expeditiously used to evaluate the microbiologic safety of food products. In general, future research needs for the routine application of alternative methods to detect enteric viral and parasitic protozoal contamination in foods requires development of the following: 1) simple, rapid, and cost-effective extraction and concentration procedures; 2) simple and reliable methods for the removal of inhibitors; 3) methods that are not restricted by food product; and 4) quantitative approaches for assessing the relative levels of contamination.
Improved epidemiologic surveillance, predominantly through the creation of population-based centers that will focus on the epidemiology and prevention of food and waterborne infectious disease (72), should improve knowledge regarding viral etiology in foodborne disease outbreaks and the clinical and economic importance of viral and parasitic disease agents. Data obtained from these studies may elucidate the role of foods in sporadic disease as well as in secondary spread.
Current research programs under way at CDC, U.S. Department of Agriculture, and Food and Drug Administration, as well as extramural funding through the National Institutes of Health and U.S. Department of Agriculture programs, should promote the development, testing, and dissemination of rapid and accurate detection methods for viral and parasitic foodborne disease agents. Such research will continue to be important as testing regulations, such as the landmark Environmental Protection Agency Information Collections Requirement Rule, and the new U.S. Department of Agriculture Pathogen Reduction: Hazard Analysis and Critical Control Points (HACCP) Systems Rule, emerge in the future. From a clinical standpoint, the development of inexpensive and widely available reagents has been improved through recent developments in molecular biology (33). This will increase the number of facilities able to perform diagnostic testing, which will ultimately facilitate epidemiologic investigation. From a food safety standpoint, improved detection methods should eventually provide a regulatory option for monitoring food safety, particularly in the economically important shellfish species and in detecting viral contamination due to human handling, or parasitic protozoal contamination from animal wastes. The development of rapid detection methods will also aid in the evaluation of control strategies for viral and protozoal contamination of foods. For instance, depuration, thermal processing, and irradiation for the control of foodborne viral contamination in shellfish need to be further evaluated. In the case of contamination by infected food handlers and animal wastes, the availability of rapid methods will increase our understanding of this transmission route and help determine critical control points for HACCP approaches to control the transmission of these foodborne disease agents. This will allow the integration of a true farm-to-table food safety approach for the control of viral and parasitic food and waterborne disease.
In the United States, there is currently a clear regulatory mandate to evaluate food safety risks within a systematic conceptual framework in the form of quantitative microbial risk assessment. The application of quantitative risk assessment to food safety issues has been hampered by the lack of available data regarding prevalence, transmission, infectious dose, and behavior of microorganisms in foods; this has been particularly true for emerging pathogens. Improved epidemiologic and detection methods should dramatically affect the ability to evaluate food safety risks through risk assessment strategies. For instance, by using emerging epidemiologic data in conjunction with epidemiologic modeling, statistical overview analysis (meta-analysis), and geographic information systems, scientists should be able to improve hazard analysis and exposure assessment and provide a clearer picture of disease transmission patterns. More rapid, accurate, and readily available detection methods should allow determination of prevalence of contamination and disease, and when quantitative, should provide a means of assessing dose-response relationships. Together these will improve subsequent risk characterization, allowing regulators to quantitate the magnitude of the problem, evaluate risk reduction strategies, and prioritize competing risks.
The combination of increased surveillance, improved detection methods, and testing requirements should result in a marked improvement in the ability to detect, investigate, and control food and waterborne enteric viral and parasitic protozoal agents. Taken together, these approaches promise to provide increased information necessary to assess risks, control disease, and ultimately improve public health in the next century.
References
- Morse SS. Factors in the emergence of infectious disease. Emerg Infect Dis. 1995;1:5–10. DOIGoogle Scholar
- Jaykus LA, Hemard MT, Sobsey MD. Human enteric pathogenic viruses. In: Hackney CR, Peirson MD, editors. Environmental indicators and shellfish safety. New York: Chapman and Hall; 1994.
- DuPont HL, Chappell CL, Sterling CR, Okhuysen PC, Rose JB, Jakubowski W. The infectivity of Cryptosporidium parvum in healthy volunteers. N Engl J Med. 1995;332:855–9. DOIPubMedGoogle Scholar
- Haas CN. Estimation of risk due to low doses of microorganisms: a comparison of alternative methodologies. Am J Epidemiol. 1983;118:573–82.PubMedGoogle Scholar
- Moe CL. Waterborne transmission of infectious agents. In: Hurst CJ, Knudsen GR, McInerney MJ, Stenzenbach LD, Walter MV, editors. Manual of environmental microbiology. Washington (DC): American Society for Microbiology; 1996.
- Council for Agricultural Science and Technology. Foodborne pathogens: risks and consequences. Washington (DC): Library of Congress No. 122; 1994.
- Jenkins S, Horman JT, Isreal E, Cukor G, Blacklow NR. An outbreak of Norwalk-related gastroenteritis in a boys' camp. Am J Dis Child. 1985;139:787–9.PubMedGoogle Scholar
- Hedberg CW, Osterholm MT. Outbreaks of foodborne and waterborne viral gastroenteritis. Clin Microbiol Rev. 1993;6:199–210.PubMedGoogle Scholar
- Bean NH, Griffin PM. Foodborne disease outbreaks in the United States, 1973-1987: pathogens, vehicles, and trends. J Food Prot. 1990;53:804–17.
- Kaplan JE, Feldman R, Campbell DS, Lookabaugh C, Gary W. The frequency of a Norwalk-like pattern of illness in outbreaks of acute gastroenteritis. Am J Public Health. 1982;72:1329–32. DOIPubMedGoogle Scholar
- Kaplan JE, Gary GW, Baron RC, Singh N, Schonberger LB, Feldman R, Epidemiology of Norwalk gastroenteritis and the role of Norwalk virus in outbreaks of acute nonbacterial gastroenteritis. Ann Intern Med. 1982;96:756–76.PubMedGoogle Scholar
- Cliver DO, Ellender RD, Fout GS, Shields PA, Sobsey MD. Foodborne viruses. In: Vanderzant C, Splittstoesser DF, editors. Compendium of methods for the microbiological examination of foods. Washington (DC): American Public Health Association; 1992.
- Centers for Disease Control. Foodborne and waterborne disease outbreaks annual summary 1976. Washington (DC): U.S. Department of Health, Education, and Welfare Publication No. CDC 78-8185; 1978.
- Gerba CP, Goyal SM. Detection and occurrence of enteric viruses in shellfish: a review. J Food Prot. 1978;41:743–54.
- Appleton H. Small round viruses: classification and role in food-borne infections. In: Ciba Foundation Symposium. Novel Diarrhoea Viruses. Chinchester, England: Wiley Publishers; 1987.
- Heun EM, Vogt RL, Hudson PJ, Parren S, Gary GW. Risk factors for secondary transmission in households after a common-source outbreak of Norwalk gastroenteritis. Am J Epidemiol. 1987;126:1181–6.PubMedGoogle Scholar
- White KE, Osterholm MT, Mariotti JA, Korlath JA, Lawrence DH, Ristinen TL, A foodborne outbreak of Norwalk virus gastroenteritis: evidence for post-recovery transmission. Am J Epidemiol. 1986;124:120–6.PubMedGoogle Scholar
- Xu ZY, Li ZH, Wang JX, Xiao ZP, Dong DH. Ecology and prevention of a shellfish-associated hepatitis A epidemic in Shanghai, China. Vaccine. 1992;Suppl 1:S67–8. DOIPubMedGoogle Scholar
- Desenclos JCA, Klontz KC, Wilder MH, Nainan OV, Margolis HS, Gunn RA. A multistate outbreak of hepatitis A caused by the consumption of raw oysters. Am J Public Health. 1991;81:1268–72. DOIPubMedGoogle Scholar
- Centers for Disease Control and Prevention. Multistate outbreak of viral gastroenteritis associated with consumption of oysters-Apalachicola Bay, Florida, December 1995-January 1995. MMWR Morb Mort Wkly Rpt. 1995;44:37–9.
- Centers for Disease Control and Prevention. Viral gastroenteritis associated with consumption of raw oysters-Florida, 1993. MMWR Morb Mort Wkly Rpt. 1994;43:446–9.
- Centers for Disease Control and Prevention. State outbreak of viral gastroenteritis related to consumption of oysters-Louisiana, Maryland, Mississippi, and North Carolina, 1993. MMWR Morb Mort Wkly Rpt. 1993;42:945–8.
- Cannon RO, Poliner RJ, Hirshhorn RB, Rodeheaver DC, Silverman PR, Brown EA, A multistate outbreak of Norwalk virus gastroenteritis associated with consumption of commercial ice. J Infect Dis. 1991;164:860–4.PubMedGoogle Scholar
- Warner RD, Carr RW, McClesky FK, Johnson PC, Goldy Elmer LM, Davison VE. A large nontypical outbreak of Norwalk virus: gastroenteritis associated with exposing celery to nonpotable water and with Citrobacter freundii. Arch Intern Med. 1991;151:2419–24. DOIPubMedGoogle Scholar
- Centers for Disease Control and Prevention. Surveillance for waterborne disease outbreaks-United States, 1991-1992. MMWR Morb Mort Wkly Rpt. 1993;42:1–22.
- LeChevallier MW, Norton WD, Lee RG. Giardia and Cryptosporidium spp. in filtered drinking water supples. Appl Environ Microbiol. 1991;57:2617–21.PubMedGoogle Scholar
- Rose JB, Gerba CP, Jakubowski W. Survey of potable water supplies for Cryptosporidium and Giardia. Environ Sci Technol. 1991;25:1393–400. DOIGoogle Scholar
- Bean NH, Griffin PM, Goulding JS, Ivey CB. Foodborne disease outbreaks, 5-year summary, 1983-1987. MMWR Morb Mort Wkly Rpt. 1990;39(SS-1):15–57.
- Bean NH, Goulding JS, Lao C, Angulo FJ. Surveillance for foodborne disease outbreaks-United States, 1988-1992. MMWR Morb Mort Wkly Rpt. 1996;45(SS-5):1–66.
- Ortega YR, Sterling CR, Gilman RH, Cama VA, Diaz F. Cyclospora speciesA new protozoan pathogen of humans. N Engl J Med. 1993;328:1308–12. DOIPubMedGoogle Scholar
- Hayes EB, Matte TD, O'Brien TR, McKinley TW, Logsdon GS, Rose JB, Large community outbreak of cryptosporidiosis due to contamination of a filtered public water supply. N Engl J Med. 1989;320:1372–6.PubMedGoogle Scholar
- MacKenzie WR, Hoxie NJ, Proctor ME, Gradus MS, Blair KA, Peterson DE, A massive outbreak in Milwaukee of Cryptosporidium infection transmitted through the public water supply. N Engl J Med. 1994;331:161–7. DOIPubMedGoogle Scholar
- Kapikian AJ, Estes MK, Chanock RM. Norwalk group of viruses. In: Fields BN, Knipe DM, Howley PM, editors. Fields Virology. Philadelphia: Lippencott-Raven Publishers, Inc.; 1996.
- Current WL, Garcia LS. Cryptosporidiosis. Clin Microbiol Rev. 1991;4:324–58.
- Campbell PN, Current WL. Demonstration of serum antibodies to Cryptosporidium sp. in normal and immunodeficient humans with confirmed infections. J Clin Microbiol. 1983;18:165–9.PubMedGoogle Scholar
- Ungar BLP, Soave R, Fayer R, Nash TE. Enzyme immunoassay detection of immunoglobulin M and G antibodies to Cryptosporidium in immunocompetent and immunocompromised persons. J Infect Dis. 1986;153:570–8.PubMedGoogle Scholar
- DeLeon R, Jaykus L. Detection of bacteria and viruses in shellfish. In: Hurst CJ, Knudsen GR, McInerney MJ, Stenzenbach LD, Walter MV, editors. Manual of environmental microbiology. Washington (DC): American Society for Microbiology; 1996.
- Sobsey MD, Carrick RJ, Jensen HR. Improved methods for detecting enteric viruses in oysters. Appl Environ Microbiol. 1978;36:121–8.PubMedGoogle Scholar
- Cole MT, Kilgen MB, Reily LR, Hackney CR. Detection of enteroviruses, bacterial indicators, and pathogens in Louisiana oysters and their overlying waters. J Food Prot. 1986;49:596–601.
- Jiang X, Graham DY, Wang K, Estes MK. Norwalk virus genome cloning and characterization. Science. 1990;250:1580–3. DOIPubMedGoogle Scholar
- Lambden PR, Caul EO, Ashley CR, Clarke IN. Sequence and genome organization of a human small round-structured (Norwalk-like) virus. Science. 1993;259:516–9. DOIPubMedGoogle Scholar
- Atmar RL, Metcalf TG, Neill HF, Estes MK. Detection of enteric viruses in oysters by using the polymerase chain reaction. Appl Environ Microbiol. 1993;59:631–5.PubMedGoogle Scholar
- Atmar RL, Neill HF, Romalde JL, LeGuyader F, Woodley CM, Metcalf TG, Detection of Norwalk virus and hepatitis A virus in shellfish tissues with the PCR. Appl Environ Microbiol. 1995;61:3014–8.PubMedGoogle Scholar
- Goswami BB, Koch WH, Cebula TA. Detection of hepatitis A in Mercenaria mercenaria by coupled reverse transcription and polymerase chain reaction. Appl Environ Microbiol. 1993;59:2765–70.PubMedGoogle Scholar
- Lees DN, Henshilwood K, Dore WJ. Development of a method for detection of enteroviruses in shellfish by PCR with poliovirus as a model. Appl Environ Microbiol. 1994;60:2999–3005.PubMedGoogle Scholar
- Lees DN, Henshilwood K, Green J, Gallimore CI, Brown DWG. Detection of small round structured viruses in shellfish by reverse transcription-PCR. Appl Environ Microbiol. 1995;61:4418–24.PubMedGoogle Scholar
- Gouvea V, Santos N, Carmo Timenetsky M, Estes MK. Identification of Norwalk virus in artificially seeded shellfish and selected foods. J Virol Methods. 1994;48:177–87. DOIPubMedGoogle Scholar
- Deng MY, Day SP, Cliver DO. Detection of hepatitis A virus in environmental samples by antigen-capture PCR. Appl Environ Microbiol. 1994;60:1927–33.PubMedGoogle Scholar
- Jaykus L, DeLeon R, Sobsey MD. A virion concentration method for detection of human enteric viruses in oysters by PCR and oligoprobe hybridization. Appl Environ Microbiol. 1996;62:2074–80.PubMedGoogle Scholar
- Dix A. Development of methods to extract human enteric viruses from hard-shelled clams for detection by reverse transcriptase-polymerase chain reaction (RT-PCR) and oligoprobe hybridization (OP) [thesis]. Raleigh (NC): North Carolina State University; 1997.
- Chung H, Jaykus L, Sobsey MD. Improved detection of human enteric viruses in field oyster specimens by in vivo and in vitro amplification of nucleic acids. Appl Environ Microbiol. 1996;62:3772–8.PubMedGoogle Scholar
- Schaefer FW III. Detection of protozoan parasites in source and finished drinking waters. In: Hurst CJ, Knudsen GR, McInerney MJ, Stenzenbach LD, Walter MV, editors. Manual of Environmental Microbiology. Washington (DC): American Society for Microbiology; 1996.
- LeChevallier MW, Norton WD, Siegel JE, Abbaszadegan M. Evaluation of the immunofluorescence procedure for detection of Giardia cysts and Cryptosporidium oocyts in water. Appl Environ Microbiol. 1995;61:690–7.PubMedGoogle Scholar
- Nieminski EC, Schaefer FW III, Ongerth JE. Comparison of two methods for the detection of Giardia cysts and Cryptosporidium oocytes in water. Appl Environ Microbiol. 1995;61:1714–9.PubMedGoogle Scholar
- Versey G, Slade JS, Byrne M, Shepard K, Fricker CR. A new method for the concentration of Cryptosporidium oocysts from water. J Appl Bacteriol. 1993;75:82–6.PubMedGoogle Scholar
- Rodgers MR, Flanigan DJ, Jakubowski W. Identification of algae which interfere with the detection of Giardia cysts and Cryptosporidium oocysts and a method for alleviating this interference. Appl Environ Microbiol. 1995;61:3759–63.PubMedGoogle Scholar
- Campbell AT, Robertson LJ, Smith HV. Viability of Cryptosporidium parvum oocytes: correlation of in vitro excystation with inclusion/exclusion of fluorogenic vital dyes. Appl Environ Microbiol. 1992;58:3488–93.PubMedGoogle Scholar
- Campbell AT, Robertson LJ, Smith HV. Novel methodology in the detection of Cryptosporidium parvum: a comparison of cooled charge coupled device (CCD) and flow cytometry. Water Sci Technol. 1993;27:89–92.
- Campbell AT, Haggart R, Robertson LJ, Smith HV. Fluorescent imaging of Cryptosporidium using a cooled charge couple device (CCD). J Microbiol Methods. 1992;16:169–74. DOIGoogle Scholar
- Abbaszadegan M, Gerba CP, Rose JB. Detection of Giardia cysts with a cDNA probe and applications to water samples. Appl Environ Microbiol. 1991;57:927–31.PubMedGoogle Scholar
- Erlandsen SL, VanKeulen H, Gurien A, Jakubowski W, Schafer FW III, Wallis P, Molecular approach to speciation and detection of Giardia: fluorochrome rDNA probes for identification of Giardia lamblia, Giardia muris, and Giardia ardeae in laboratory and environmental samples by in situ hybridization. In: Thompson RCA, Reynoldson JA, Lymbery AJ, editors. Giardia: from molecules to disease. Oxford: CAB International; 1994.
- Mahbubani MH, Bej AK, Perlin M, Schaefer FW III, Jakubowski W, Atlas RM. Detection of Giardia cysts by using polymerase chain reaction and distinguishing live from dead cysts. Appl Environ Microbiol. 1991;57:3456–61.PubMedGoogle Scholar
- Mahbubani MH, Bej AK, Perlin M, Schaefer FW III, Jakubowski W, Atlas RM. Differentiation of Giardia duodenalis and other Giardia spp. by using polymerase chain reaction and gene probes. J Clin Microbiol. 1992;30:74–8.PubMedGoogle Scholar
- Webster KA, Pow JDE, Giles M, Catchpole J, Woodward MJ. Detection of Cryptosporidium parvum using a specific polymerase chain reaction. Vet Parasitol. 1993;50:35–44. DOIPubMedGoogle Scholar
- de la Cruz AA, Sivaganesan M. Detection of Giardia and Cryptosporidium spp. in source water samples by commercial enzyme-immunosorbant assay. In: Proceedings 1994 Water Technology Conference; 1994 Nov 6-10; San Francisco, California. Denver (CO): American Water Works Association, 1995.
- Sauch JF, Flanigan D, Galvin ML, Berman D, Jakubowski W. Propidium iodide as an indicator of Giardia cyst viability. Appl Environ Microbiol. 1991;57:3243–7.PubMedGoogle Scholar
- Schupp DG, Erlandsen SL. A new method to determine Giardia cyst viability: correlation of florescein diacetate and propidium iodide staining with animal infectivity. Appl Environ Microbiol. 1987;53:704–7.PubMedGoogle Scholar
- Moe CL, Gentsch J, Ando T, Grohmann G, Monroe SS, Jiang X, Application of PCR to detect Norwalk virus in fecal specimens from outbreaks of gastroenteritis. J Clin Microbiol. 1994;32:642–8.PubMedGoogle Scholar
- Green J, Norcott JP, Lewis D, Arnold C, Brown DWG. Norwalk-like viruses: demonstration of genomic diversity by polymerase chain reaction. J Clin Microbiol. 1993;31:3007–12.PubMedGoogle Scholar
- Wang J, Jiang X, Madore HP, Gray J, Desselberge U, Ando T, Sequence diversity of small, round-structured viruses in the Norwalk virus group. J Virol. 1994;68:5982–90.PubMedGoogle Scholar
- Ando T, Monroe SS, Gentsch JR, Jin Q, Lewis DC, Glass RL. Detection and differentiation of antigenically distinct small round-structured viruses (Norwalk-like viruses) by reverse transcription-PCR and southern hybridization. J Clin Microbiol. 1995;33:64–71.PubMedGoogle Scholar
- Centers for Disease Control and Prevention. Addressing emerging infectious disease threats: a prevention strategy for the United States. Atlanta (GA): U.S. Department of Health and Human Services, U.S. Public Health Service; 1996.
Figure
Tables
Cite This ArticleTable of Contents – Volume 3, Number 4—December 1997
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Lee-Ann Jaykus, Department of Food Science, North Carolina State University, Box 7624, Raleigh, NC 27695 USA; fax: 919-515-7124
Top