Genetically Similar High-Risk Strains of Carbapenemase-Producing Enterobacterales in Humans and Companion Animals, United States
Lingzi Xiaoli, Allison E. James

, Anna L. Stahl, Maho Okumura, Stephen D. Cole, Jaclyn M. Dietrich, Molly M. Leeper, Jordan K. Putney, Maroya Spalding Walters, and Richard A. Stanton
Author affiliation: Centers for Disease Control and Prevention, Atlanta, Georgia, USA (L. Xiaoli, A.E. James, A.L. Stahl, M.M. Leeper, J.K. Putney, M.S. Walters, R.A. Stanton); University of Pennsylvania School of Veterinary Medicine, Philadelphia, Pennsylvania, USA (M. Okumura, S.D. Cole, J.M. Dietrich); Applied Science Research and Technology, Inc., Smyrna, Georgia, USA (J.K. Putney); US Public Health Service, Rockville, Maryland, USA (M.S. Walters)
Main Article
Figure 2
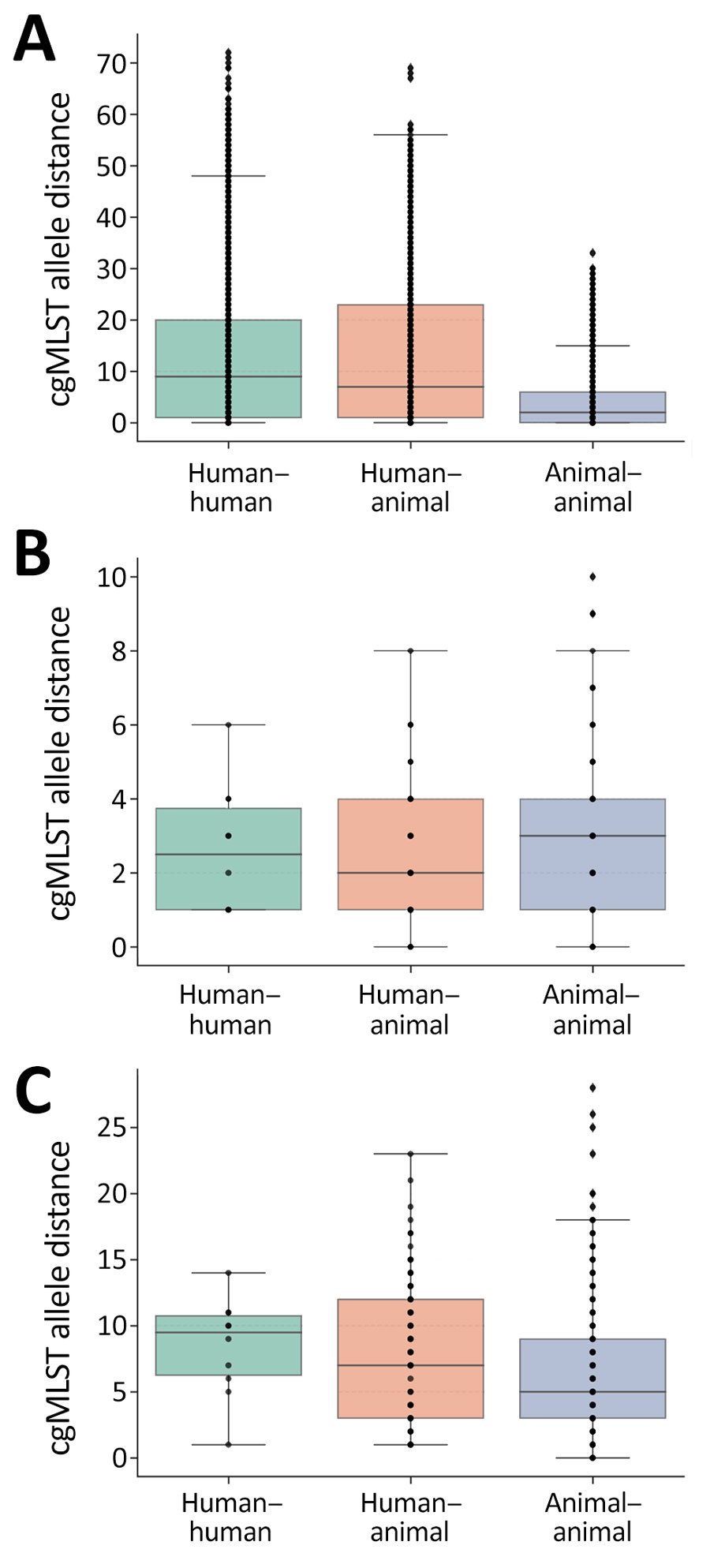
Figure 2. Frequency boxplots of pairwise within-cluster cgMLST allele distances among carbapenemase-producing carbapenem-resistant Enterobacterales isolates collected from humans and companion animals in study of genetically similar high-risk strains of carbapenemase-producing Enterobacterales in humans and companion animals, United States. Pairwise cgMLST allele distances were calculated between pairs within individual clusters and depicted by bacterial species with Escherichia coli (A), Klebsiella pneumoniae (B), and Enterobacter cloacae (C). Box top and bottom boundaries depict 25th and 75th quartiles, horizontal lines within boxes depict median values, dots represent individual data points, and whiskers represent datapoints within 1.5 times the interquartile range. cgMLST, core-genome multilocus sequence typing.
Main Article
Page created: February 07, 2026
Page updated: March 20, 2026
Page reviewed: March 20, 2026
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.
