Volume 32, Number 3—March 2026
Research Letter
lsaC and Tandem lsaE-lnuB Resistance Genes in Invasive Group A Streptococcus
Figure 2
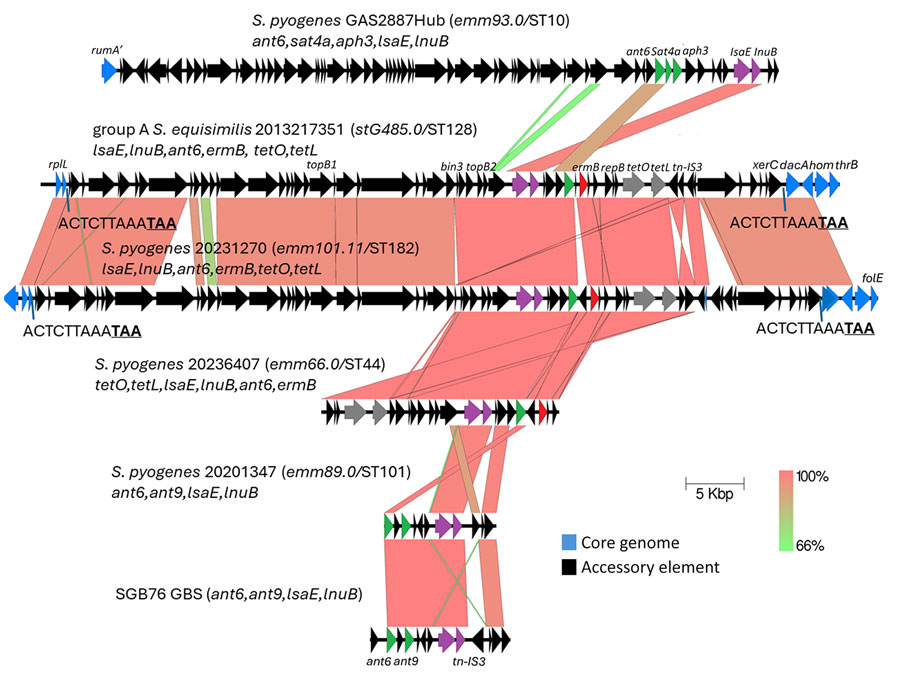
Figure 2. Alignment of complete lsaE-carrying elements from study of repeated acquisitions of lsaC and tandem lsaE-lnuB resistance genes by group A Streptococcus. Alignments shown are from group A ABCs strains 2013217351 and 20231270 with partial elements from strains 20236407 and 20201347. Also included are complete elements from GAS2887Hub (8) and GBS strain SGB76 (GenBank accession no. KF772204). Antimicrobial resistance genes include 3 aminoglycoside 6-adenyltransferase genes (ant6, aph3, and ant9), and the streptothricin acetyltransferase gene aph3. Prokka annotations include topB (DNA topoisomerase genes), bin3 (DNA invertase gene), repB (DNA replication gene), tn-IS3 (IS3 family transposase gene), and xerC (tyrosine recombinase gene). Underlined bold text indicates the stop codon of the rplL gene in 2 strains. Scale bar indicates 5,000 base pairs. ABCs, Active Bacterial Core surveillance; sero, serotype; ST, sequence type.
References
- Khan ZZ. Group A streptococcal (GAS) infections treatment & management, 2024 [cited 2026 Feb 1]. https://emedicine.medscape.com/article/228936-treatment
- Li Y, Rivers J, Mathis S, Li Z, McGee L, Chochua S, et al. Continued increase of erythromycin nonsusceptibility and clindamycin nonsusceptibility among invasive group A streptococci driven by genomic clusters, United States, 2018–2019. Clin Infect Dis. 2023;76:e1266–9. DOIGoogle Scholar
- Schwarz S, Shen J, Kadlec K, Wang Y, Brenner Michael G, Feßler AT, et al. Lincosamides, streptogramins, phenicols, and pleuromutilins: mode of action and mechanisms of resistance. Cold Spring Harb Perspect Med. 2016;6:
a027037 . DOIGoogle Scholar - File TM Jr, Goldberg L, Das A, Sweeney C, Saviski J, Gelone SP, et al. Efficacy and safety of intravenous-to-oral lefamulin, a pleuromutilin antibiotic, for the treatment of community-acquired bacterial pneumonia: the phase III Lefamulin Evaluation Against Pneumonia (LEAP 1) trial. Clin Infect Dis. 2019;69:1856–67. DOIGoogle Scholar
- Paukner S, Sader HS, Ivezic-Schoenfeld Z, Jones RN. Antimicrobial activity of the pleuromutilin antibiotic BC-3781 against bacterial pathogens isolated in the SENTRY antimicrobial surveillance program in 2010. Antimicrob Agents Chemother. 2013;57:4489–95. DOIGoogle Scholar
- Malbruny B, Werno AM, Murdoch DR, Leclercq R, Cattoir V. Cross-resistance to lincosamides, streptogramins A, and pleuromutilins due to the lsa(C) gene in Streptococcus agalactiae UCN70. Antimicrob Agents Chemother. 2011;55:1470–4. DOIGoogle Scholar
- Douarre PE, Sauvage E, Poyart C, Glaser P. Host specificity in the diversity and transfer of lsa resistance genes in group B Streptococcus. J Antimicrob Chemother. 2015;70:3205–13.
- Berbel D, Càmara J, García E, Tubau F, Guérin F, Giard JC, et al. A novel genomic island harbouring lsa(E) and lnu(B) genes and a defective prophage in a Streptococcus pyogenes isolate resistant to lincosamide, streptogramin A and pleuromutilin antibiotics. Int J Antimicrob Agents. 2019;54:647–51. DOIGoogle Scholar
- Beall B, Lin W, Li Z, Tran T, Metcalf BJ, Anderson BJ, et al. Two independent acquisitions of multidrug resistance gene lsaC in Streptococcus pneumoniae serotype 20 multilocus sequence type 1257. Emerg Infect Dis. 2025;31:2098–108. DOIGoogle Scholar
- Chochua S, Rivers J, Mathis S, Li Z, Velusamy S, McGee L, et al. Emergent invasive group A Streptococcus dysgalactiae subsp. equisimilis, United States, 2015–2018. Emerg Infect Dis. 2019;25:1543–7. DOIGoogle Scholar