Volume 32, Number 4—April 2026
Research
Geographically Distinct Circulation of Genotype II and III St. Louis Encephalitis Virus, Texas, USA, 2009–2024
Figure 6
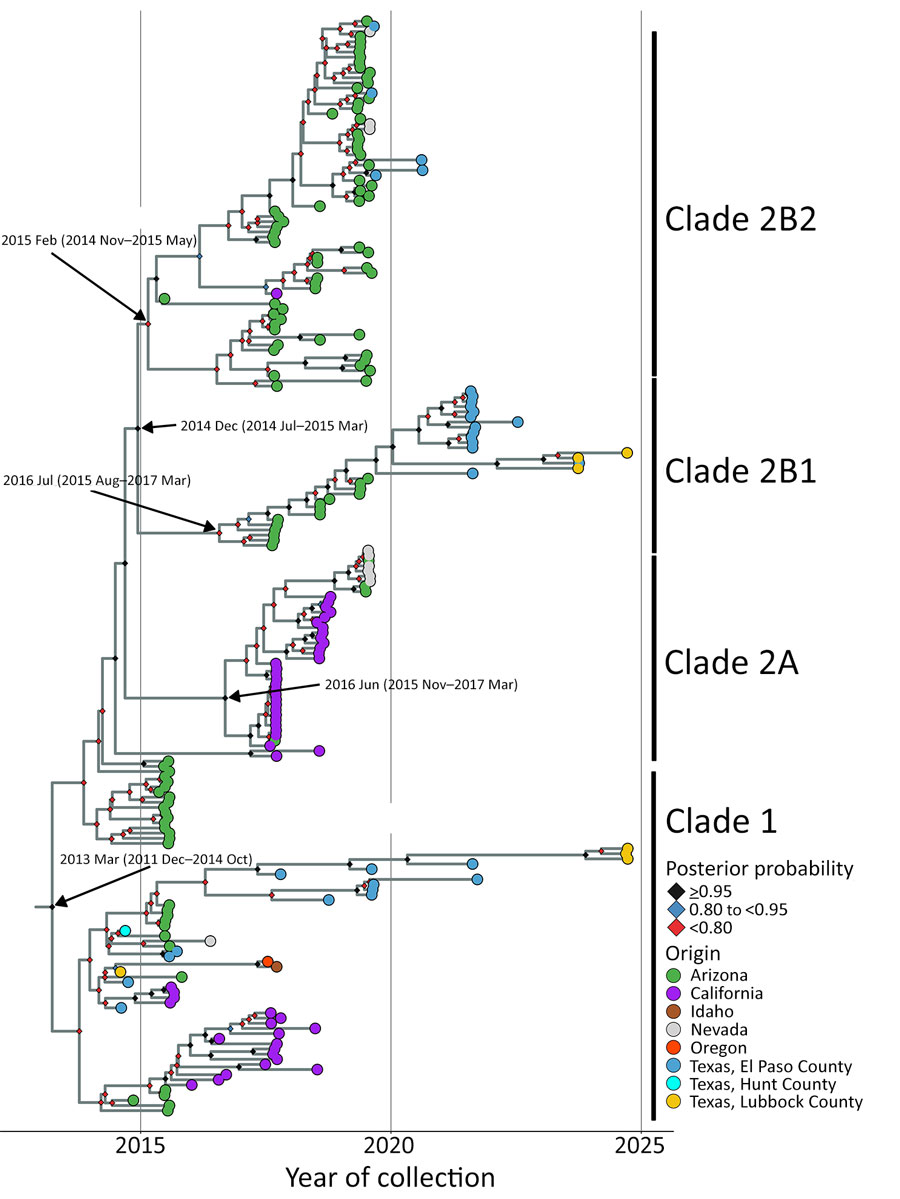
Figure 6. Bayesian phylogenetic analysis of SLEV genotype III from Texas and other states from study of circulation of genotype II and III St. Louis encephalitis virus, Texas, USA. Maximum clade credibility time tree of 216 whole genomes inferred by using BEAST (34). Tip colors reflect sample’s place of origin. Texas samples are further subdivided by county. Clade annotations are indicated to right of tips. Posterior probability for each node is indicated at node with a colored diamond. The y-axis scales in years are based on date of sample collection. Arrow labels indicate time to most recent common ancestor and 95% highest posterior density for each clade.
References
- Diaz A, Coffey LL, Burkett-Cadena N, Day JF. Reemergence of St. Louis Encephalitis Virus in the Americas. Emerg Infect Dis. 2018;24:2150–7. DOIPubMedGoogle Scholar
- Reisen WK, Fang Y, Martinez VM. Avian host and mosquito (Diptera: Culicidae) vector competence determine the efficiency of West Nile and St. Louis encephalitis virus transmission. J Med Entomol. 2005;42:367–75. DOIPubMedGoogle Scholar
- Reisen WK, Chiles RE, Martinez VM, Fang Y, Green EN. Experimental infection of California birds with western equine encephalomyelitis and St. Louis encephalitis viruses. J Med Entomol. 2003;40:968–82. DOIPubMedGoogle Scholar
- Gruwell JA, Fogarty CL, Bennett SG, Challet GL, Vanderpool KS, Jozan M, et al. Role of peridomestic birds in the transmission of St. Louis encephalitis virus in southern California. J Wildl Dis. 2000;36:13–34. DOIPubMedGoogle Scholar
- McLean RG, Mullenix J, Kerschner J, Hamm J. The house sparrow (Passer domesticus) as a sentinel for St. Louis encephalitis virus. Am J Trop Med Hyg. 1983;32:1120–9. DOIPubMedGoogle Scholar
- Leake J, Musson E, Chope H. Epidemiology of epidemic encephalitis, St. Louis type. JAMA. 1934;103:728–31. DOIGoogle Scholar
- Curren EJ, Lindsey NP, Fischer M, Hills SLSS. Louis encephalitis virus disease in the United States, 2003–2017. Am J Trop Med Hyg. 2018;99:1074–9. DOIPubMedGoogle Scholar
- Danforth ME, Snyder RE, Feiszli T, Bullick T, Messenger S, Hanson C, et al. Epidemiologic and environmental characterization of the re-emergence of St. Louis encephalitis virus in California, 2015–2020. PLoS Negl Trop Dis. 2022;16:
e0010664 . DOIPubMedGoogle Scholar - Venkat H, Krow-Lucal E, Kretschmer M, Sylvester T, Levy C, Adams L, et al. Comparison of characteristics of patients with West Nile Virus or St. Louis encephalitis virus neuroinvasive disease during concurrent outbreaks, Maricopa County, Arizona, 2015. Vector Borne Zoonotic Dis. 2020;20:624–9. DOIPubMedGoogle Scholar
- Centers for Disease Control and Prevention (CDC). Outbreak of West Nile-like viral encephalitis—New York, 1999. MMWR Morb Mortal Wkly Rep. 1999;48:845–9.PubMedGoogle Scholar
- Reisen WK, Lothrop HD, Wheeler SS, Kennsington M, Gutierrez A, Fang Y, et al. Persistent West Nile virus transmission and the apparent displacement St. Louis encephalitis virus in southeastern California, 2003-2006. J Med Entomol. 2008;45:494–508.PubMedGoogle Scholar
- Reimann CA, Hayes EB, DiGuiseppi C, Hoffman R, Lehman JA, Lindsey NP, et al. Epidemiology of neuroinvasive arboviral disease in the United States, 1999-2007. Am J Trop Med Hyg. 2008;79:974–9. DOIPubMedGoogle Scholar
- Mackenzie JS, Barrett AD, Deubel V. The Japanese encephalitis serological group of flaviviruses: a brief introduction to the group. Curr Top Microbiol Immunol. 2002;267:1–10. DOIPubMedGoogle Scholar
- Fang Y, Reisen WK. Previous infection with West Nile or St. Louis encephalitis viruses provides cross protection during reinfection in house finches. Am J Trop Med Hyg. 2006;75:480–5. DOIPubMedGoogle Scholar
- Bosco-Lauth AM, Kooi K, Hawks SA, Duggal NK. Cross-protection between West Nile virus and emerging flaviviruses in wild birds. Am J Trop Med Hyg. 2024;112:657–62. DOIPubMedGoogle Scholar
- Kramer LD, Chandler LJ. Phylogenetic analysis of the envelope gene of St. Louis encephalitis virus. Arch Virol. 2001;146:2341–55. DOIPubMedGoogle Scholar
- Rodrigues SG, Nunes MR, Casseb SM, Prazeres AS, Rodrigues DS, Silva MO, et al. Molecular epidemiology of Saint Louis encephalitis virus in the Brazilian Amazon: genetic divergence and dispersal. J Gen Virol. 2010;91:2420–7. DOIPubMedGoogle Scholar
- Diaz LA, Ré V, Almirón WR, Farías A, Vázquez A, Sanchez-Seco MP, et al. Genotype III Saint Louis encephalitis virus outbreak, Argentina, 2005. Emerg Infect Dis. 2006;12:1752–4. DOIPubMedGoogle Scholar
- Venkat H, Krow-Lucal E, Hennessey M, Jones J, Adams L, Fischer M, et al. Concurrent outbreaks of St. Louis encephalitis virus and West Nile Virus disease—Arizona, 2015. MMWR Morb Mortal Wkly Rep. 2015;64:1349–50.PubMedGoogle Scholar
- White GS, Symmes K, Sun P, Fang Y, Garcia S, Steiner C, et al. Reemergence of St. Louis encephalitis Virus, California, 2015. Emerg Infect Dis. 2016;22:2185–8. DOIPubMedGoogle Scholar
- Ridenour CL, Cocking J, Poidmore S, Erickson D, Brock B, Valentine M, et al. St. Louis encephalitis virus in the southwestern United States: a phylogeographic case for a multi-variant introduction event. Front Genet. 2021;12:
667895 . DOIPubMedGoogle Scholar - Swetnam DM, Stuart JB, Young K, Maharaj PD, Fang Y, Garcia S, et al. Movement of St. Louis encephalitis virus in the Western United States, 2014- 2018. PLoS Negl Trop Dis. 2020;14:
e0008343 . DOIPubMedGoogle Scholar - Chandler LJ, Parsons R, Randle Y. Multiple genotypes of St. Louis encephalitis virus (Flaviviridae: Flavivirus) circulate in Harris County, Texas. Am J Trop Med Hyg. 2001;64:12–9. DOIPubMedGoogle Scholar
- Lillibridge KM, Parsons R, Randle Y, Travassos da Rosa AP, Guzman H, Siirin M, et al. The 2002 introduction of West Nile virus into Harris County, Texas, an area historically endemic for St. Louis encephalitis. Am J Trop Med Hyg. 2004;70:676–81. DOIPubMedGoogle Scholar
- Williams KH, Hollinger FB, Metzger WR, Hopkins CC, Chamberlain RW. The epidemiology of St. Louis encephalitis in Corpus Christi, Texas, 1966. Am J Epidemiol. 1975;102:16–24. DOIPubMedGoogle Scholar
- Luby JP, Miller G, Gardner P, Pigford CA, Henderson BE, Eddins D. The epidemiology of St. Louis encephalitis in Houston, Texas, 1964. Am J Epidemiol. 1967;86:584–97. DOIPubMedGoogle Scholar
- Kneubehl AR. Texas-SLEV-genome-surveillance. 2025 [cited 2025 Jan 1]. https://github.com/kneubehl/Texas-SLEV-Genome-Surveillance
- Lanciotti RS, Kerst AJ. Nucleic acid sequence-based amplification assays for rapid detection of West Nile and St. Louis encephalitis viruses. J Clin Microbiol. 2001;39:4506–13. DOIPubMedGoogle Scholar
- Wang MX, Lou EG, Sapoval N, Kim E, Kalvapalle P, Kille B, et al. Olivar: towards automated variant aware primer design for multiplex tiled amplicon sequencing of pathogens. Nat Commun. 2024;15:6306. DOIPubMedGoogle Scholar
- Katoh K, Standley DM. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol. 2013;30:772–80. DOIPubMedGoogle Scholar
- Hoang DT, Chernomor O, von Haeseler A, Minh BQ, Vinh LS. UFBoot2: Improving the ultrafast bootstrap approximation. Mol Biol Evol. 2018;35:518–22. DOIPubMedGoogle Scholar
- Minh BQ, Schmidt HA, Chernomor O, Schrempf D, Woodhams MD, von Haeseler A, et al. IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol Biol Evol. 2020;37:1530–4. DOIPubMedGoogle Scholar
- Rambaut A, Lam TT, Max Carvalho L, Pybus OG. Exploring the temporal structure of heterochronous sequences using TempEst (formerly Path-O-Gen). Virus Evol. 2016;2:
vew007 . DOIPubMedGoogle Scholar - Drummond AJ, Rambaut A. BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol Biol. 2007;7:214. DOIPubMedGoogle Scholar
- May FJ, Li L, Zhang S, Guzman H, Beasley DWC, Tesh RB, et al. Genetic variation of St. Louis encephalitis virus. J Gen Virol. 2008;89:1901–10. DOIPubMedGoogle Scholar
- Auguste AJ, Pybus OG, Carrington CV. Evolution and dispersal of St. Louis encephalitis virus in the Americas. Infect Genet Evol. 2009;9:709–15. DOIPubMedGoogle Scholar
- Lincoln FC. The waterfowl flyways of North America. Washington: US Department of Agriculture; 1935. p. 7–8.
- Tasi TF, Smith GC, Ndukwu M, Jakob WL, Happ CM, Kirk LJ, et al. Entomologic studies after a St. Louis encephalitis epidemic in Grand Junction, Colorado. Am J Epidemiol. 1988;128:285–97. DOIPubMedGoogle Scholar
- Hayes RO, LaMotte LC, Holden P. Ecology of arboviruses in Hale County, Texas, during 1965. Am J Trop Med Hyg. 1967;16:675–87.PubMedGoogle Scholar
- Gorris ME, Bartlow AW, Temple SD, Romero-Alvarez D, Shutt DP, Fair JM, et al. Updated distribution maps of predominant Culex mosquitoes across the Americas. Parasit Vectors. 2021;14:547. DOIPubMedGoogle Scholar
- Luby JP, Haley RW. Recurrent St. Louis encephalitis infection in residents of a flood plain of the Trinity River, Roosevelt Heights (Dallas, Texas). Am J Epidemiol. 1972;96:107–13. DOIPubMedGoogle Scholar
- Wiseman J, Grimes J, Eads RS. Louis encephalitis outbreak in Cameron County, Texas. Mosq News. 1959;19. DOIGoogle Scholar
- Erickson TA, Ronca SE, Gunter SM, Brown EL, Hasbun R, Murray KO. Zoonotic disease testing practices in pediatric patients with meningitis and encephalitis in a subtropical region. Pathogens. 2022;11:501. DOIPubMedGoogle Scholar
- Vanichanan J, Salazar L, Wootton SH, Aguilera E, Garcia MN, Murray KO, et al. Use of testing for West Nile virus and other arboviruses. Emerg Infect Dis. 2016;22:1587–93. DOIPubMedGoogle Scholar
Page created: March 04, 2026
Page updated: April 15, 2026
Page reviewed: April 15, 2026
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.