Volume 32, Number 4—April 2026
Dispatch
Cardiomyopathy Caused by Coxsackievirus Strain A9 in Previously Healthy Child, Northeastern France, 2024
Cite This Article
Citation for Media
Abstract
We characterized a recombinant mosaic coxsackievirus A9 strain responsible for severe inflammatory cardiomyopathy in a previously healthy child in northeastern France in 2024 by using whole-genome sequencing. This case highlights that enterovirus species other than coxsackievirus strain B3 can cause cardiomyopathy in otherwise healthy pediatric patients.
Coxsackievirus strain A9 (CVA9) is a human pathogen classified within the species Enterovirus betacoxsackievirus (species B enterovirus [EV-B]). Infections linked to CVA9 show up in children, typically hand, foot, and mouth disease or episodes of aseptic meningitis (1). Epidemiologic tracking points to CVA9 as a frequent nonpolio enterovirus found in patient samples worldwide (1). During a 5-year survey in Europe from 2018–2023, this virus appeared among the 10 most documented enterovirus strains in clinical samples (2). Recent findings document CVA9 involvement in bronchiolitis, pneumonia, meningitis, and encephalitis; severe complications are rare but can be fatal (1,3,4). The severe neurologic or respiratory complications induced by CVA9 highlight its emerging clinical significance beyond its association with benign illnesses (1). Molecular epidemiology studies show frequent recombination in CVA9 coding and noncoding regions, potentially enhancing virulence and broadening the clinical spectrum of recombinant strains (5,6). We describe the clinical and virologic characterization of a recombinant mosaic CVA9 strain identified by whole-genome sequencing and involved in a severe inflammatory cardiomyopathy in a previously healthy child in France in 2024.
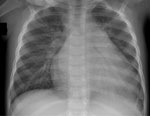
A <24-month-old girl was admitted to the pediatric emergency unit of the Centre Hospitalier Universitaire de Reims in Reims, France, in mid-July 2024 with a 6-week history of cough, parentally reported fever (temperature of 38°C), and suspected cardiomegaly on a chest radiograph obtained the day before (Figure 1). Her general practitioner had prescribed treatment for bronchiolitis (without viral documentation) and amoxicillin, followed by azithromycin and a chest radiograph. The child was born at term and had unremarkable growth and development, no history of recurrent infections, and unremarkable baseline immune globulin levels. Clinical examination revealed tympanic temperature of 38.2°C, blood pressure 112/65 mm Hg, sinus tachycardia (150 beats/min), gallop rhythm, and liver edge 3 cm below the costal border. Transthoracic echocardiography revealed severe left ventricular dilation and markedly reduced left ventricular ejection fraction (≈27.5%); cardiac structures and coronary arteries appeared unremarkable. Cardiac troponin T was 57 ng/L (reference range <14 ng/L), and N-terminal prohormone of brain natriuretic peptide (NT-ProBNP) was 32,486 pg/mL (reference range <300 pg/mL). We conducted a nasopharyngeal swab specimen multiplex real-time PCR on July 11 (R-GENE Respiratory PCR Kit; bioMérieux, https://www.biomerieux.com) that was negative for adenovirus, respiratory syncytial virus, metapneumovirus, influenza, and parainfluenza but detected enterovirus RNA. Results of blood real-time PCRs for human herpes virus 1–6, adenovirus, and parvovirus B19 were negative, whereas enterovirus viremia was detected and quantified in peripheral EDTA blood samples (1.32 × 104 copies/mL). Throat and blood cultures showed no pathogenic bacterial growth (7).
Besides congestive heart failure therapy, the patient received corticosteroids for 2 days and 1 course of intravenous immunoglobulin (IVIg) on day 2. Two supplemental courses of IVIg were prescribed on day 7 and 16 (Table). Cardiac magnetic resonance imaging done on day 19 revealed left ventricle dilation without enhancement after gadolinium injection. Outcome was ultimately favorable, with normalization of troponin T and NT-ProBNP levels within 3 months and of left ventricular ejection fraction within 6 months (Table). Results of genetic screening for inherited cardiomyopathies were negative (8). Although a cardiac biopsy was not performed because of an unfavorable benefit to risk ratio in this infant, stage C myocarditis was diagnosed according to recent American College of Cardiology guidelines (9). Written informed consent was obtained from a parent for research use of clinical and biologic data.
Published studies on cardiac tissue and whole blood from myocarditis patients identified EV-B populations with 5′-terminal RNA deletions. We assessed those forms in our patient’s blood specimen by using validated rapid amplification of cDNA ends PCR (7,10). Our findings indicate that the major CVA9 genome exhibited a 50-nt 5′-terminal deletion, previously reported in a coxsackievirus B (CVB) 3 strain as a cardiotropic, replication-competent form (11) (Appendix Figure 1).
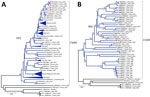
Reverse transcription PCR targeting the complete viral capsid protein viral protein (VP) 1 genomic region with Sanger sequencing or the complete genome with Illumina sequencing (Illumina, https://www.illumina.com) enabled genotypic identification of 2 original CVA9 clinical strains from our study patient (GenBank accession nos. PV939324 and PX441804) (12). Those strains exhibited 100% nucleotide and amino acid identity in VP1. Phylogenetic analysis of complete CVA9 VP1 genomic region sequences indicated strong homology with another CVA9 isolated in 2024 in Versailles, France (GenBank accession no. PQ612466), from the peripheral blood of a 28-day-old infant with acute respiratory tract infection. The strain from our patient also closely clustered with another CVA9 isolated in 2024 in Versailles, France, and other CVA9 strains isolated in China during 2012–2021 (Figure 2, panel A).
Whole-genome sequencing of CVA9 strains by using Illumina protocols (12) provided interpretable data only for our study respiratory sample. Phylogenetic analysis revealed elongated branches and close clustering of genetically distinct sequences within the first polyprotein precursor region region (e.g., E6/E3 and E11) (Figure 2, panel B), suggesting recombination hotspots in structural or nonstructural protein-coding regions, which likely drive CVA9 diversity and evolution (5).
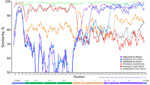
To investigate CVA9 genomic recombination, we conducted SimPlot analyses (Figure 3). The 2 CVA9 isolates from 2024 (a cardiotropic reference and the Versailles strain) were colinear across the genome, indicating no evidence of recombination between them. Sequence comparison between our reference cardiotropic strain, a CVA9 isolated in 2025 in France and the virus isolated in China in 2019 revealed high similarity (≈95%) within the first polyprotein precursor region and 5′ regions, but a distinct breakpoint within the 2A gene, suggesting recombination in our sequence. Of note, between positions 3,600 and 3,900, sequence similarity increased with the CVA9 2022 strain from Russia, consistent with recombination events involving viruses isolated in Russia and Kazakhstan. An additional recombination event likely involved a strain related to E3 France 2024. Furthermore, the E3 strain itself is recombinant, sharing a genomic segment with E6 from position 4,700 to the genome terminus (Appendix Figure 2). This terminal region, which encodes RNA polymerase (positions 5,960–7,645), appears to have been further acquired by recombination with an E11 strain associated with epidemics in Europe and severe neonatal infections (12). Phylogenetic analyses of partial protein 2A–2B and 3D polymerase (3Dpol) gene sequences confirmed the genetic homology between our CVA9 strain and the strain isolated in Versailles in 2024. Those analyses also clarified the geographic origins of the recombinant segments: the mid-part of P2A was derived from a CVA9 strain previously detected in Russia, whereas 3Dpol originated from E11 strains from Europe associated with severe neonatal infections (Appendix Figure 2) (12). P2A and 3Dpol genes are key determinants of enterovirus caused cardiac infection because they dampen innate type I interferon responses and drive viral RNA replication, including production of 5′ terminally deleted RNA forms (11,13).
Unlike well-established cardiotropic species, B enterovirus strains such as CVB3 and CVB5, other species B enteroviruses such as CVA9, and some species A enteroviruses such as CVA6 have only sporadically been linked to myocarditis and cardiac histologic lesions, with rare fatal outcomes (14,15). Although the cardiotropic potential of those strains is not as well documented as for CVB viruses, recent molecular and clinical evidence supports the capacity of CVA9 to induce cardiac inflammation, particularly in neonates and children (1). Our study case not only confirms the myocardial tropism of CVA9 but also challenges its reputation as a predominantly benign childhood pathogen. This case highlights that other species B enteroviruses besides CVB3 can cause cardiomyopathy in otherwise healthy pediatric patients.
Ms. Lebreil is a molecular biology specialist developing genotyping technologies for RNA viruses. Her research interests include viral immune evasion mechanisms, host-pathogen interactions in cardiac tissue, and molecular determinants of enterovirus persistence in chronic diseases.
Acknowledgment
We sincerely thank all members of the pediatric ward at CHU de Reims, France.
References
- Machado RS, Tavares FN, Sousa IP Jr. Global landscape of coxsackieviruses in human health. Virus Res. 2024;344:
199367 . DOIPubMedGoogle Scholar - de Schrijver S, Vanhulle E, Ingenbleek A, Alexakis L, Johannesen CK, Broberg EK, et al.; ENPEN Study Collaborators. Epidemiological and clinical insights into enterovirus circulation in Europe, 2018–2023: a multicenter retrospective surveillance study. J Infect Dis. 2025;232:e104–15. DOIPubMedGoogle Scholar
- Chen BS, Lee HC, Lee KM, Gong YN, Shih SR. Enterovirus and encephalitis. Front Microbiol. 2020 Feb 20 [cited 2025 Oct 3]. https://www.frontiersin.org/journals/microbiology/articles/10.3389/fmicb.2020.00261/full
- Eisenhut M, Algawi B, Wreghitt T, Foweraker J, McKee T, Miles R, et al. Fatal coxsackie A9 virus infection during an outbreak in a neonatal unit. J Infect. 2000;40:297–8. DOIPubMedGoogle Scholar
- Hietanen E, Susi P. Recombination events and conserved nature of receptor binding motifs in coxsackievirus A9 isolates. Viruses. 2020;12:68. DOIPubMedGoogle Scholar
- Zhao H, Wang J, Chen J, Huang R, Zhang Y, Xiao J, et al. Molecular epidemiology and evolution of coxsackievirus A9. Viruses. 2022;14:822. DOIPubMedGoogle Scholar
- Glenet M, N’Guyen Y, Mirand A, Henquell C, Lebreil AL, Berri F, et al.; French Enterovirus Myocarditis Study Group. Major 5′ terminally deleted enterovirus populations modulate type I IFN response in acute myocarditis patients and in human cultured cardiomyocytes. Sci Rep. 2020;10:11947. DOIPubMedGoogle Scholar
- Weizman O, Gandjbakhch E, Magnin-Poull I, Proukhnitzky J, Bordet C, Palmyre A, et al. Molecular genetic screening after non-ischaemic sudden cardiac arrest and no overt cardiomyopathy in real life: a major tool for the aetiological diagnostic work-up. Arch Cardiovasc Dis. 2024;117:382–91. DOIPubMedGoogle Scholar
- Drazner MH, Bozkurt B, Cooper LT, Aggarwal NR, Basso C, Bhave NM, et al. Writing Committee. 2024 ACC expert consensus decision pathway on strategies and criteria for the diagnosis and management of myocarditis: a report of the American college of cardiology solution set oversight committee. J Am Coll Cardiol. 2025;85:391–431. DOIPubMedGoogle Scholar
- Mirabel M, Callon D, Bruneval P, Lebreil AL, Mousseaux E, Oudard S, et al. Late-onset giant cell myocarditis due to enterovirus during treatment with immune checkpoint inhibitors. JACC Cardiooncol. 2020;2:511–4. DOIPubMedGoogle Scholar
- Callon D, Glenet M, Lebreil AL, Heng L, Bouland N, Fichel C, et al. Major group-B enterovirus populations deleted in the noncoding 5′ region of genomic RNA modulate activation of the type I interferon pathway in cardiomyocytes and induce myocarditis. PLoS Pathog. 2024;20:
e1012125 . DOIPubMedGoogle Scholar - Grapin M, Mirand A, Pinquier D, Basset A, Bendavid M, Bisseux M, et al. Severe and fatal neonatal infections linked to a new variant of echovirus 11, France, July 2022 to April 2023. Euro Surveill. 2023;28:
2300253 . DOIPubMedGoogle Scholar - Bouin A, Gretteau PA, Wehbe M, Renois F, N’Guyen Y, Lévêque N, et al. Enterovirus persistence in cardiac cells of patients with idiopathic dilated cardiomyopathy is linked to 5′ terminal genomic RNA-deleted viral populations with viral-encoded proteinase activities. Circulation. 2019;139:2326–38. DOIPubMedGoogle Scholar
- Callon D, N’Guyen Y, Fornes P, Andreoletti L. Fatal heart arrhythmia associated with enterovirus cardiac infection and SARS-CoV-2-induced cytokine storm. Pathology. 2024;56:434–7. DOIPubMedGoogle Scholar
- Andréoletti L, Lévêque N, Boulagnon C, Brasselet C, Fornes P. Viral causes of human myocarditis. Arch Cardiovasc Dis. 2009;102:559–68. DOIPubMedGoogle Scholar
Figures
Table
Cite This ArticleOriginal Publication Date: April 10, 2026
1These authors contributed equally to this article.
Table of Contents – Volume 32, Number 4—April 2026
| EID Search Options |
|---|
|
|
|
|
|
|


Please use the form below to submit correspondence to the authors or contact them at the following address:
Laurent Andreoletti, Laboratoire de Virologie Médicale et Moléculaire, CHU & UMR-S INSERM 1320, Faculté de Médecine, 51 rue Cognacq-Jay, 51100 Reims CEDEX, France
Top