Volume 32, Number 5—May 2026
Research
Updated Genomic Epidemiologic Description of Candida (Candidozyma) auris, United States
Figure 3
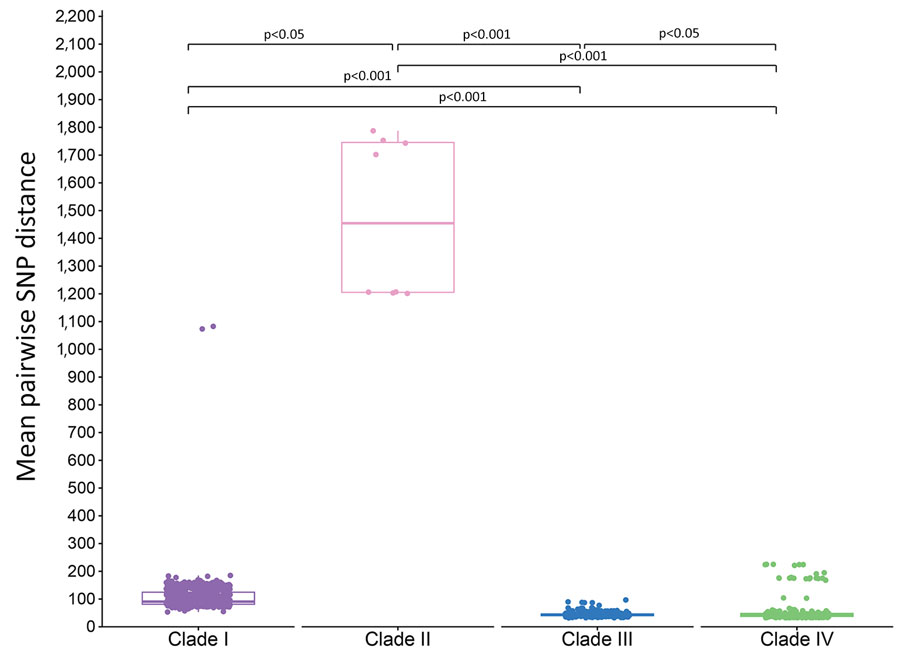
Figure 3. Pairwise SNP distances from study of genomic epidemiologic description of Candida (Candidozyma) auris isolates, by clade, United States. Tukey boxplots summarize pairwise SNP distances within each clade; horizontal line within box indicates the median, upper and lower boundaries indicate interquartile range); whiskers indicate largest and smallest distance value within 1.5 times the interquartile range. Dots correspond to each isolate’s mean pairwise SNP distance within its clade. The Kruskal-Wallis test with posthoc Dunn test indicated significant differences in pairwise SNP distances between all clade pairs. Reference strains used were as follows: clade I, B11205 (GenBank accession no. GCA_016772135.1); clade II, B11220 (GenBank accession no. GCA_003013715.2); clade III, B11221 (GenBank accession no. GCF_002775015.1); clade IV, B11243 (GenBank accession no. GCA_003014415.1). SNP, single-nucleotide polymorphism.