Volume 32, Number 5—May 2026
Dispatch
Clinical, Molecular, and Zoonotic Perspectives on Human Cases of Cryptosporidium sp. OTUi
Figure 2
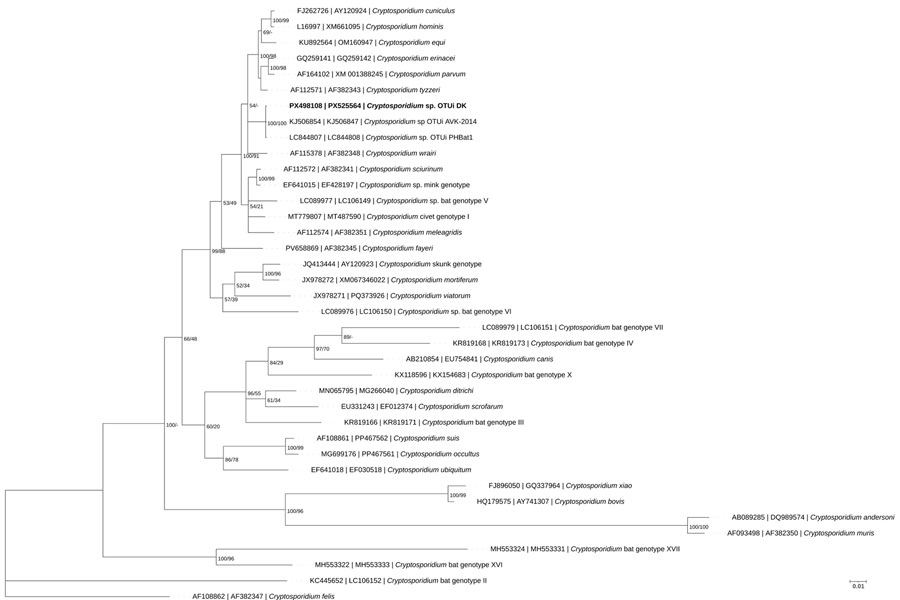
Figure 2. Phylogenetic tree inferred by partitioned Bayesian analysis on the basis of concatenated ssu and actin gene sequences of Cryptosporidium spp., including Cryptosporidium sp. OTUi identified from a woman in Denmark who had traveled to Indonesia (bold text) in a study of the clinical, molecular, and zoonotic perspectives on human cases of Cryptosporidium. GenBank accession numbers for ssu and actin sequences are provided. Alignments of ssu and actin were concatenated according to species identity and OTUi sample identity; only taxa with sequences available for both loci were included. For Bayesian analyses, substitution models were selected on the basis of the lowest Akaike information criterion score (general time reversible with invariable site plus discrete gamma model substitution for ssu and actin). Partitioned Bayesian inference was performed by using MrBayes v3.2.7 (13) with 10 million generations, 4 chains, and sampling every 1,000 generations; the first 25% of trees were discarded as burn-in. Convergence was confirmed by SD of split frequencies <0.01 and a potential scale reduction factor of 1.0. Posterior probabilities and bootstrap support values from maximum-likelihood analysis are shown to the right of the nodes. Scale bar indicates substitutions per site.
References
- Larsen TG, Ethelberg S, Nielsen HL, Hartmeyer GN, Nielsen L, Zangenberg M, et al. From rare to recognized: enhanced detection uncovers Cryptosporidium endemicity and species diversity in Denmark. Emerg Microbes Infect. 2025;14:
2529893 . DOIPubMedGoogle Scholar - Alves M, Xiao L, Sulaiman I, Lal AA, Matos O, Antunes F. Subgenotype analysis of Cryptosporidium isolates from humans, cattle, and zoo ruminants in Portugal. J Clin Microbiol. 2003;41:2744–7. DOIPubMedGoogle Scholar
- Koehler AV, Whipp M, Hogg G, Haydon SR, Stevens MA, Jex AR, et al. First genetic analysis of Cryptosporidium from humans from Tasmania, and identification of a new genotype from a traveller to Bali. Electrophoresis. 2014;35:2600–7. DOIPubMedGoogle Scholar
- Babineau M, Koehler AV, Sait ML, Mercoulia K, Dougall S, McAllister J, et al. Large-scale molecular epidemiological survey of Giardia and Cryptosporidium in Victoria, Australia (2020-2024), reveals novel subtypes and outbreak-associated lineages. J Clin Microbiol. 2026;0:
e0155825 . DOIPubMedGoogle Scholar - Stensvold CR, Larsen TG, Grüttner J, Nielsen L, Engberg J, Lebbad M. Rodent-adapted Cryptosporidium infection in humans: seven new cases and review of the literature. One Health. 2024;18:
100682 . DOIPubMedGoogle Scholar - Lebbad M, Winiecka-Krusnell J, Stensvold CR, Beser J. High diversity of Cryptosporidium species and subtypes identified in cryptosporidiosis acquired in Sweden and abroad. Pathogens. 2021;10:523. DOIPubMedGoogle Scholar
- Xu L, Fukuda Y, Murakoshi F, Alviola P, Masangkay J, Recuenco FC, et al. Molecular characterization and zoonotic risk assessment of Cryptosporidium spp. in Philippine bats. Food Waterborne Parasitol. 2024;38:
e00249 . DOIPubMedGoogle Scholar - Yang L, Wang Q, Xu L, Tu C, Huang X, He B. Detection and characterization of a novel norovirus in bats, China. Virol Sin. 2018;33:100–3. DOIPubMedGoogle Scholar
- Nowak K, Fahr J, Weber N, Lübke-Becker A, Semmler T, Weiss S, et al. Highly diverse and antimicrobial susceptible Escherichia coli display a naïve bacterial population in fruit bats from the Republic of Congo. PLoS One. 2017;12:
e0178146 . DOIPubMedGoogle Scholar - Luby SP, Rahman M, Hossain MJ, Blum LS, Husain MM, Gurley E, et al. Foodborne transmission of Nipah virus, Bangladesh. Emerg Infect Dis. 2006;12:1888–94. DOIPubMedGoogle Scholar
- Khan MS, Hossain J, Gurley ES, Nahar N, Sultana R, Luby SP. Use of infrared camera to understand bats’ access to date palm sap: implications for preventing Nipah virus transmission. EcoHealth. 2010;7:517–25. DOIPubMedGoogle Scholar
- Barbosa AD, Egan S, Feng Y, Xiao L, Ryan U. How significant are bats as potential carriers of zoonotic Cryptosporidium and Giardia? Curr Res Parasitol Vector Borne Dis. 2023;4:
100155 . DOIPubMedGoogle Scholar - Huelsenbeck JP, Ronquist F. MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics. 2001;17:754–5. DOIPubMedGoogle Scholar