Genomic Surveillance of Lassa Virus through In-Country Sequencing, Guinea
Jacob Camara
1, Giuditta Annibaldis
1
, Joon Klaps
1, Kékoura Ifono
1, Fara Raymond Koundouno, Youssouf Sidibé, Sarah Ryter, Moussa Conde, Saa Lucien Millimono, Mette Hinrichs, Julia Hinzmann, Nils Peter Petersen, Mia Le, Annick Renevey, Ehizojie Ehiremen Emua, Philippe Lemey, Simon Dellicour, Stephan Günther, N’Faly Magassouba
2, Sophie Duraffour
2, Liana Eleni Kafetzopoulou
2, and Sanaba Boumbaly
2
Author affiliation: Center for Virology Research—Laboratory for Viral Hemorrhagic Fevers in Guinea, Conakry, Guinea (J. Camara, M. Conde, N. Magassouba, S. Boumbaly); Bernhard Nocht Institute for Tropical Medicine, Hamburg, Germany (G. Annibaldis, S. Ryter, M. Hinrichs, J. Hinzmann, N.P. Petersen, M. Le, A. Renevey, S. Günther, S. Duraffour); German Center for Infection Research, partner site Hamburg–Lübeck–Borstel–Riems, Hamburg (G. Annibaldis, S. Ryter, M. Hinrichs, J. Hinzmann, N.P. Petersen, M. Le, A. Renevey, S. Günther, S. Duraffour); Rega Institute, KU Leuven, Leuven, Belgium (J. Klaps, P. Lemey, S. Dellicour, L.E. Kafetzopoulou); Laboratory for Viral Hemorrhagic Fevers of Guéckédou, Prefectural Health Department of Guéckédou, Guéckédou, Guinea (K. Ifono, F.R. Koundouno, S.L. Millimono); Laboratory for Viral Hemorrhagic Fevers at Regional Hospital of N’Zérékoré, Regional Hospital of N’Zérékoré, N’Zérékoré, Guinea (Y. Sidibé); University of Hamburg, Hamburg (M. Le); Irrua Specialist Teaching Hospital, Irrua, Nigeria (E.E. Emua); Spatial Epidemiology Lab, Université Libre de Bruxelles, Brussels, Belgium (S. Dellicour); Interuniversity Institute of Bioinformatics in Brussels, Université Libre de Bruxelles, Vrije Universiteit Brussel, Brussels (S. Dellicour); Leiden University Medical Center, Leiden, the Netherlands (L.E. Kafetzopoulou)
Main Article
Figure 1
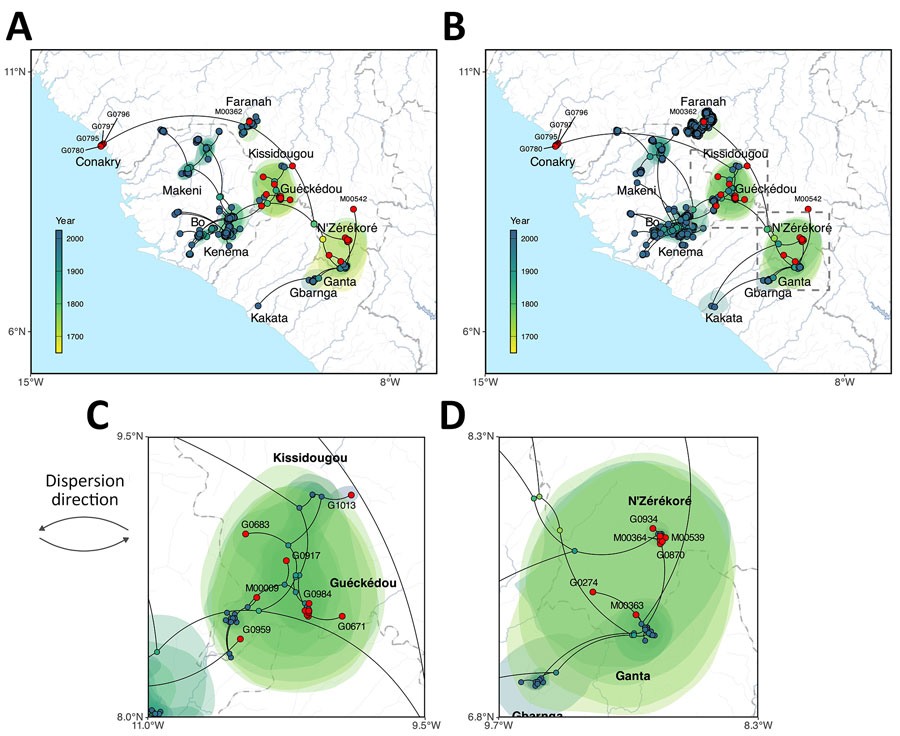
Figure 1. Phylogeographic reconstruction of the dispersal history of Lassa virus lineage IV from study of genomic surveillance of Lassa virus through in-country sequencing, Guinea. A, B) Results of the continuous phylogeographic inference based on large (A) and small (B) segment sequences. For each analysis, the corresponding maximum clade credibility tree is mapped with internal and tip nodes colored according to their estimated time of occurrence and sampling date. Tip nodes corresponding to newly sequenced cases from this study are highlighted in red. Shaded polygons represent the 80% highest posterior density regions, reflecting uncertainty in internal node location inference. The estimated root of lineage IV, dating to the 17th–18th Centuries, is indicated by dashed squares in the southeastern (forested) region of Guinea. C, D) Maps highlighting specific transmission dynamics in Guéckédou (C) and N’Zérékoré (D), illustrating the dense local clustering and the cross-border introductions from Liberia (Ganta) into the N’Zérékoré region.
Main Article
Page created: March 23, 2026
Page updated: May 08, 2026
Page reviewed: May 08, 2026
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.
