Volume 32, Number 5—May 2026
Research Letter
Genomic Surveillance of Lassa Virus through In-Country Sequencing, Guinea
Cite This Article
Citation for Media
Abstract
Strengthened in-country sequencing generated 28 Lassa virus genomes from human clinical cases in Guinea, expanding knowledge of Lassa fever in the country. Phylogeographic analysis revealed cross-border exchange between Liberia and the N’Zérékoré region and a Sierra Leone introduction into Guéckédou. Enhanced genomic surveillance is crucial to guide public health.
Lassa fever (LF) is a life-threatening viral hemorrhagic disease endemic to West Africa; early clinical symptoms are indistinguishable from other febrile illnesses, complicating diagnosis and surveillance (1). The causative agent, Lassa virus (LASV), is a Mammarenavirus (Arenaviridae family) with a bisegmented (small [S] and large [L] segments) ambisense RNA genome that exhibits distinct phylogenetic structure across its endemic regions. Lineages I–III and VI circulate in Nigeria, lineage IV predominates in the Mano River Union countries (Guinea, Sierra Leone, and Liberia), and additional distinct lineages circulate in Mali/Côte d’Ivoire (lineage V) and Togo (lineage VII). Although LF cases are only sporadically reported in Guinea, serologic evidence from the southeastern (forested) and central regions indicates broad population exposure (2,3). Sequencing efforts thus far have generated partial LASV genomes from rodent reservoirs in Upper Guinea (4), whereas genomes from human infections remain limited (5,6). Sparse genomic data limit our understanding of geogenomic variation, outbreak dynamics, and clinical correlations, highlighting the need for enhanced genomic surveillance to inform diagnostics, epidemiology, and patient management.
To expand viral surveillance and diagnostic capabilities locally in Guinea, genomic capacity strengthening was initiated in 2021 at the Centre de recherche en Virologie–Laboratoire des Fièvres Hémorragiques Virales de Guinée (CRV-LFHVG; Conakry, Guinea). Sequencing infrastructure was initially established in response to the COVID-19 pandemic using targeted Nanopore sequencing (Oxford Nanopore Technologies, https://nanoporetech.com) (7). In 2022–2023, laboratory capacity was expanded to include metagenomic sequencing (8,9), integrated within the diagnostic network of 3 surveillance laboratories for viral hemorrhagic fevers across Guinea: CRV-LFHVG, the national reference laboratory in Conakry, and 2 satellite laboratories in the forest region, in Guéckédou (Laboratoire des Fièvres Hémorragiques Virales de Guéckédou; LFHV-GKD) and N’Zérékoré (Laboratoire des Fièvres Hémorragiques Virales de Hôpital Régional de N’Zérékoré; LFHV-HRNZE). During 2020–2024, this laboratory network confirmed a total of 36 LF cases (F.R. Koundouno et al., unpub. data, https://www.medrxiv.org/content/10.64898/2026.02.24.26346968v1), including a nosocomial outbreak in Conakry in 2022 (10). In-country metagenomic nanopore sequencing was performed at CRV-LFHVG to investigate the viral diversity of the confirmed LF cases. A total of 28 LF cases were successfully sequenced, all of which yielded sufficient genomic coverage for downstream phylogenetic analysis (Appendix Table). Most sequencing yielded near-complete genomes for both segments (Appendix Table), and all sequences were phylogenetically classified as lineage IV.
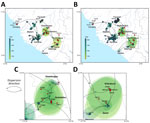
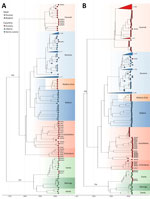
Phylogenetic analysis revealed that many of the newly sequenced genomes from Guinea are substantially divergent from previously characterized cases; branch lengths suggest years to decades of virus circulation in the natural reservoir before sampling in human cases (Figures 1, 2). Bayesian phylogeographic reconstruction estimated substitution mean rates of 8.5 × 10−4 (S segment) and 8.2 × 10−4 (L segment) substitutions/site/year, placing the root of lineage IV in the 17th–18th Centuries, likely originating in southeastern (forested) Guinea (Figures 1, 2).
Spatial viral diversity in Guinea is organized into 3 predominant geographic clusters, mainly associated with the areas of Guéckédou, N’Zérékoré, and Faranah. That factor should be interpreted cautiously, because Guéckédou and N’Zérékoré both have surveillance laboratories, and sequences from the Faranah region originate from rodent reservoirs sampled during previous studies. Consequently, the inferred geographic clustering might be driven, at least in part, by uneven sampling. Two cocirculating sublineages of LASV were identified in the N’Zérékoré region: an older IVb lineage, which has been long-established locally (M00539, M00541, G0405, G0870, G0934); and the IVa lineage, which traces back to multiple independent introductions (M00363, M00542, M00364, G0274) from Ganta, northeastern Liberia, with common ancestors dating to the 1950s–1980s (Figure 1, panel D; Figure 2). In Guéckédou, we detected an imported LASV case originating from the Kenema area of Sierra Leone (G0959) (Figure 1, panel C; Figure 2), illustrating that sporadic introductions from neighboring LASV-endemic regions to Guinea can occur and contribute to Guinean LASV diversity.
We included 4 LF cases from Conakry, in the western part of Guinea, all of which were associated to a previously reported nosocomial transmission chain linked to a travel case from a LASV-endemic area (10) (Appendix). The LASV sequences (G0780, G0795, G0796, G0797) show minimal between-sequence variation (L segment, 0–4 mutations; S segment, 0–1 mutations), consistent with a single transmission event. We identified all 4 genomes as a reassortant LASV variant with their 2 segments clustering significantly differently (Appendix). Their L segments clustered with sequences previously identified in Faranah (lineage IVb in Figure 2, panel A), and their S segments clustered with sequences previously identified in Guéckédou (lineage IVb in Figure 2, panel B).
This study increases the available LASV sequences derived from human clinical cases and provides new genomic insights into LASV circulation in Guinea. Our findings were made possible through strengthened laboratory diagnostics in LF-endemic areas (Guéckédou and N’Zérékoré) and the establishment of new sequencing capacity for viral hemorrhagic fevers at CRV-LFHVG, Conakry. Ongoing genomic surveillance remains crucial for guiding public health interventions, as well as for the development of appropriate medical countermeasures.
Mr. Camara leads the sequencing team at Centre de recherche en Virologie–Laboratoire des Fièvres Hémorragiques Virales de Guinée (CRV-LFHVG) in Conakry, Guinea, where he oversees genomic sequencing activities and data generation. Dr. Annibaldis is based at the Bernhard Nocht Institute for Tropical Medicine in Hamburg, Germany, where she coordinates strengthening projects for viral hemorrhagic fevers surveillance. Her work focuses on sequencing capacity and laboratory system strengthening in Guinea and Nigeria.
Acknowledgments
We thank the Agence Nationale de Sécurité Sanitaire, the Ministry of Health of the Republic of Guinea, the “Délégations Régionale et Préfectorale de la Santé,” and the healthcare workers involved in the response.
This descriptive research, using anonymized diagnostic surveillance data, has been approved by the National Ethics Committee of Guinea (CNERS) under the number 009/CNERS/25. This work is part of the Nagoya permit number 006/2023/PN.
The work was supported by the German Federal Ministry of Health through support of the World Health Organization Collaborating Centre for Arboviruses and Hemorrhagic Fever Viruses at the Bernhard-Nocht-Institute for Tropical Medicine (agreement ZMV I1-2517WHO005), the Global Health Protection Program (GHPP, agreements ZMV I1-2517GHP-704, ZMVI1-2519GHP704, and ZMI1-2521GHP921 until end of 2022 and, from 2023, agreements ZMI5-2523GHP006 and ZMI5-2523GHP008), the COVID-19 surge fund (BMG ZMVI1-2520COR001), the Research and Innovation Programme of the European Union under H2020 grant agreement n°871029-EVA-GLOBAL, and the Research Foundation–Flanders (Fonds voor Wetenschappelijk Onderzoek–Vlaanderen, G005323N and G051322N, 1SH2V24N, 12X9222N). The BNITM is a member of the German Center for Infection Research (DZIF, partner site Hamburg–Lübeck–Borstel–Riems, Hamburg, Germany), and all works performed in this study have been supported by DZIF. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
G.A., F.R.K., Y.S., S.G., N.F.M., S.Du., L.E.K., and S.B. conceived and designed the study. J.C., G.A., K.I., F.R.K., Y.S., S.R., M.C., S.L.M., M.H., and J.H. collected data or performed laboratory diagnostics. J.C., G.A., K.I., S.R., M.C., S.L.M., M.H., J.H., N.P.P., M.L., and A.R. performed sequencing or sequence validation. J.C., G.A., J.K., K.I., S.R., M.C., S.L.M., and S.De. conducted formal phylogenetic analysis. J.K. and S.De. performed phylogeography analysis. J.C., G.A., J.K., K.I., F.R.K., Y.S., S.R., M.H., J.H., A.R., E.E.E., P.L., S.G., N.F.M., S.Du., L.E.K., and S.B. implemented the project. P.L., S.G., N.F.M., S.Du., and S.B. acquired funding. J.C., G.A., J.K., K.I., S.G., N.F.M., S.Du., L.E.K., and S.B. wrote the manuscript. All authors edited the manuscript. All authors read and approved the contents of the manuscript.
During the preparation of this work the authors used ChatGPT/free version to edit some sentences. After using this tool/service, the authors reviewed and edited the content as needed and take full responsibility for the content of the publication.
This article was preprinted at https://www.medrxiv.org/content/10.64898/2026.03.04.26347418v1.
References
- Kernéis S, Koivogui L, Magassouba N, Koulemou K, Lewis R, Aplogan A, et al. Prevalence and risk factors of Lassa seropositivity in inhabitants of the forest region of Guinea: a cross-sectional study. PLoS Negl Trop Dis. 2009;3:
e548 . DOIPubMedGoogle Scholar - Mariën J, Nuismer SL, Magassouba N, Soropogui B, Günther S, Becker-Ziaja B, et al. Serosurveillance identifies an endemic hotspot of Lassa fever in Faranah, Upper Guinea. J Infect Dis. 2025;232:e830–8. DOIPubMedGoogle Scholar
- Fichet-Calvet E, Ölschläger S, Strecker T, Koivogui L, Becker-Ziaja B, Camara AB, et al. Spatial and temporal evolution of Lassa virus in the natural host population in Upper Guinea. Sci Rep. 2016;6:21977. DOIPubMedGoogle Scholar
- Magassouba N, Koivogui E, Conde S, Kone M, Koropogui M, Soropogui B, et al. A sporadic and lethal Lassa fever case in Forest Guinea, 2019. Viruses. 2020;12:1062. DOIPubMedGoogle Scholar
- Wiley MR, Fakoli L, Letizia AG, Welch SR, Ladner JT, Prieto K, et al. Lassa virus circulating in Liberia: a retrospective genomic characterisation. Lancet Infect Dis. 2019;19:1371–8. DOIPubMedGoogle Scholar
- Magassouba N, Gustani-Buss E, Ifono K, Nelson EV, Camara J, Annibaldis G, et al. Two years of SARS-CoV-2 genomic surveillance capacity development in Guinea. Sci Rep. 2026;16:11225. DOIPubMedGoogle Scholar
- Camara J, Sidibé Y, Annibaldis G, Soropogui B, Ryter S, Condé M, et al. Dengue diagnosis in Guinea in two returning travelers from Côte d’Ivoire: a case report. IJID Reg. 2025;17:
100777 . DOIPubMedGoogle Scholar - Koundouno FR, Kafetzopoulou LE, Faye M, Renevey A, Soropogui B, Ifono K, et al. Detection of Marburg virus disease in Guinea. N Engl J Med. 2022;386:2528–30. DOIPubMedGoogle Scholar
- Annibaldis G, Soropogui B, Ifono K, Camara J, Kaba ML, Berete F, et al. Nosocomial outbreak of Lassa fever in Conakry, Guinea, 2022. J Infect Dis. 2026;•••:
jiag229 ; Epub ahead of print. DOIPubMedGoogle Scholar
Figures
Cite This ArticleOriginal Publication Date: April 28, 2026
1These first authors contributed equally to this article.
2These senior authors contributed equally to this article.
Table of Contents – Volume 32, Number 5—May 2026
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Giuditta Annibaldis, Bernhard Nocht Institute for Tropical Medicine (BNITM), Bernhard Nocht Straße 74, 20359, Hamburg, Germany
Top