Volume 13, Number 10—October 2007
Dispatch
Foot-and-Mouth Disease Virus Serotype A in Egypt
Cite This Article
Citation for Media
Abstract
We describe the characterization of a foot-and-mouth disease (FMD) serotype A virus responsible for recent outbreaks of disease in Egypt. Phylogenetic analysis of VP1 nucleotide sequences demonstrated a close relationship to recent FMD virus isolates from East Africa, rather than to viruses currently circulating in the Middle East.
Foot-and-mouth disease (FMD) is caused by 7 immunologically distinct serotypes, O, A, C, Asia 1, South African Territories (SAT) 1, SAT 2, and SAT 3, which belong to the species Foot-and-mouth disease virus (genus Aphthovirus, family Picornaviridae). Several of these serotypes circulate currently or periodically in the Middle East and North Africa (1). In Egypt, routine prophylactic vaccination has been conducted with a locally produced serotype O vaccine. The last outbreak of serotype O was in June 2000, and other serotypes have not been reported since 1972 when serotype A occurred (2). This report describes an FMD serotype A virus responsible for recent outbreaks of disease in Egypt.
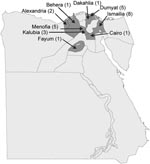
Clinical cases of FMD were first recognized on January 22, 2006, on a cattle farm at El Etehad in Ismailia, northeastern Egypt (Figure 1). Samples were submitted for laboratory investigation and serotype determination by using virus isolation, antigen ELISA, and reverse transcription–PCR (RT-PCR). Initial testing with antigen ELISA and RT-PCR assays suggested that multiple FMD virus (FMDV) serotypes may have been involved in the outbreak (data not shown), although only type A was later confirmed. On February 15, 2006, the Agriculture Ministry in Egypt notified international public health authorities (by reporting to the World Organization for Animal Health [OIE]) of 6 outbreaks of FMDV caused by serotype A in Ismailia and 12 additional outbreaks in 7 other Egyptian governorates: Alexandria (2 outbreaks), Behera (1 outbreak), Cairo (1 outbreak), Dakahlia (1 outbreak), Dumyat (5 outbreaks), Fayum (1 outbreak), and Menofia (1 outbreak). By April 6, 2006, 34 outbreaks of disease had been reported that affected >7,500 animals and involved an additional governorate (Kalubia). Most (96.7%) clinical FMD cases involved cattle; 411 cattle (mainly calves) reportedly died. Attempts to control the outbreaks were hampered by lack of an appropriate vaccine and concurrent outbreaks of highly pathogenic avian influenza. FMD became widespread in Egypt, with the following numbers of animals affected per month: 6,189 (January), 1,858 (February), 3,035 (March), 401 (April), and 297 (May). A locally produced bivalent FMDV vaccine, containing both O1 and A/Egypt/2006 isolates, was released in mid-May 2006 for the first time in Egypt. No new cases have been reported since July 2006.
Clinical material from 5 cases (collected from 3 separate locations in Egypt; Table 1) was sent to the Food and Agricultural Organization of the United Nations (FAO) World Reference Laboratory for FMD (WRLFMD) at the Institute for Animal Health, Pirbright, United Kingdom, for confirmatory diagnosis and characterization of the causative FMDV strain(s). The possibility that these samples contained multiple FMDV serotypes was also investigated. FMDV isolates causing cytopathic effects in primary bovine thyroid (BTy) cell cultures were generated from all samples. The cell culture–grown virus isolates and original clinical submissions were identified as FMDV serotype A by antigen-detection ELISA (3).
Total RNA was extracted from the first virus passage on BTy cells by using RNeasy kits (QIAGEN, Crawley, UK) for all 5 samples (EGY/1/2006–EGY/5/2006) (4). The complete VP1 region of the genome was amplified by RT-PCR by using 2 primer sets (A-1C562F/EUR-2B52R and A-1C612F/EUR-2B52R; Table 2) and the following thermal profile: 42°C for 30 min; 94°C for 5 min; 35 cycles of 94°C for 60 s, 55°C for 60 s, and 72°C for 90 s, followed by a final extension of 72°C for 5 min. The sequence of each amplicon was determined by cycle sequencing as previously described (4) but with the primers NK72, A-1C612F, and A-1D523R (Table 2). An unrooted neighbor-joining tree was constructed by using MEGA version 3.1 (5). The robustness of the tree topology was assessed with 1,000 bootstrap replicates as implemented in the program. Additionally, maximum parsimony (MEGA 3.1), minimum evolution (MEGA 3.1), and maximum likelihood (TREE-PUZZLE 5.2; [6]) trees were constructed; all 4 methods gave similar tree topologies (data not shown). Egyptian sequences shared a closer phylogenetic relationship with recent and historical isolates from East Africa rather than with contemporary serotype A viruses emerging from Iran, currently circulating in the Middle East and European Turkey (Figure 2).
Other conventional “typing” PCRs were performed to investigate whether additional FMDV serotypes were present in these samples. A multiplex agarose gel–based RT-PCR that targeted VP1 of O, A, C, and Asia 1 (primers P33, P38, P87–92, P40, P74–77) (7) generated a single band corresponding to the size expected (702 bp) for serotype A for all 5 samples (data not shown). In addition, a cocktail of primers (P1, P126, P150–153, P130, P159–161, P168–170) (7) recognizing VP1 of SAT1–3 serotypes did not show any bands after RT-PCR with these samples. However, amplicons of correct size (715 bp) were obtained after RT-PCR with samples EGY/1/2006, EGY/3/2006, and EGY/5/2006 when an additional primer set for SAT 1–3 VP1 (1D209F/2B208R) was used (8). Subsequent analysis of these SAT amplicons generated sequences that corresponded to serotype A, identical to the complete VP1 sequences of A/EGY/1/2006 and A/EGY/2/2006. Together, these findings support the conclusion that FMDV corresponding to a single serotype A was present in this material.
For vaccine selection, serologic tests were conducted to evaluate the extent of in vitro cross-neutralization of A/EGY/1/2006 and A/EGY/2/2006 by antisera produced against available FMDV vaccine strains (9). The match (r1 value) against the vaccine strains A22/Iraq/64 and A/Iran/96 that are regularly used elsewhere in the Middle East was less than the cut-off value of 0.3 (r1 = 0.23 and 0.24, respectively), whereas an acceptable match (r1 = 0.42) was found against the A/Eritrea/98 vaccine strain that is of East African origin. However, A/Eritrea/98 vaccine is not in routine production nor held in vaccine reserves and was therefore not available for immediate supply. A recent in vivo study demonstrated that a high potency A22/Iraq/64 vaccine could provide clinical protection against challenge with the new A/EGY/2006 virus (B. Haas, pers. comm., 2006). High-potency vaccines are known to protect even when relationship values are lower than the normal cut-off values (10).
Local interpretation of agarose-based RT-PCR assays and sequence data led the Egyptian authorities to initially suspect the involvement of at least 2 serotypes, A and SAT 2. However, tests performed at the WRLFMD conclusively showed the presence of a single serotype, A, in the samples received from Egypt. Unofficial reports suggest that the disease was introduced by animals imported from Ethiopia for slaughter (11). This hypothesis is consistent with the results of the molecular typing, which suggested a relation between strains of Egyptian and East African origin. The molecular typing confirms only that through the trade in live cattle, an East African type A strain was introduced, which was not contained at the quarantine station. The origin of the infection is unclear, since the animals in quarantine may have acquired infection at various points during shipment, including possible contaminated pens or other animals on board the ship, at the port before loading, or in transit from Ethiopia to the port of loading. Veterinary inspection of the quarantined animals also detected cases of lumpy skin disease (LSD), and possibly the origin of the LSD epidemic in Egypt in 2006 may relate to the Ethiopian animal trade, which is supported by the reports of LSD epidemics in Ethiopia in 2005. Undoubtedly, the lack of reporting of disease preimportation or at the quarantine stations did not assist the authorities in controlling the disease. Because imported animals may acquire infection at any point up until their arrival, they must be vaccinated and tested for the absence of FMDV nonstructural proteins.
Mr Knowles is a molecular virologist at the Institute for Animal Health. His research interest focuses on the molecular epidemiology and evolution of picornaviruses of animals, particularly FMDV.
Acknowledgments
We thank Keith Sumption for assistance in the preparation of this article.
This work was supported by Defra, UK (Reference Laboratory Contract and Research Grant nos. SE2921 and SE2935). The submission and serotyping of samples were supported by Defra and a grant from the FAO European Commission for the Control of FMD (MTF/INT/003/EEC). The latter also supported an emergency mission of the World Reference Laboratory to Egypt to provide diagnostic support in March 2006.
References
- Knowles NJ, Samuel AR. Molecular epidemiology of foot-and-mouth disease virus. Virus Res. 2003;91:65–80. DOIPubMedGoogle Scholar
- Aidaros HA. Regional status and approaches to control of foot-and-mouth disease in the Middle East and North Africa. Rev Sci Tech. 2002;21:451–8.PubMedGoogle Scholar
- Ferris NP, Dawson M. Routine application of enzyme-linked immunosorbent assay in comparison with complement fixation for the diagnosis of foot-and-mouth and swine vesicular diseases. Vet Microbiol. 1988;16:201–9. DOIPubMedGoogle Scholar
- Knowles NJ, Samuel AR, Davies PR, Midgley RJ, Valarcher J-F. Evolution and spread of a pandemic strain of foot-and-mouth disease virus serotype O. Emerg Infect Dis. 2005;11:1887–93.PubMedGoogle Scholar
- Kumar S, Tamura K, Nei M. MEGA3: Integrated software for Molecular Evolutionary Genetics Analysis and sequence alignment. Brief Bioinform. 2004;5:150–63. DOIPubMedGoogle Scholar
- Schmidt HA, Strimmer K, Vingron M, von Haeseler A. TREE-PUZZLE: maximum likelihood phylogenetic analysis using quartets and parallel computing. Bioinformatics. 2002;18:502–4. DOIPubMedGoogle Scholar
- Reid SM, Hutchings GH, Ferris NP, De Clercq K. Diagnosis of foot-and-mouth disease by RT-PCR: evaluation of primers for serotypic characterisation of viral RNA in clinical samples. J Virol Methods. 1999;83:113–23. DOIPubMedGoogle Scholar
- Reid SM, Ferris NP, Hutchings GH, Samuel AR, Knowles NJ. Primary diagnosis of foot-and-mouth disease by reverse transcription polymerase chain reaction. J Virol Methods. 2000;89:167–76. DOIPubMedGoogle Scholar
- Rweyemamu MM. Antigenic variation in foot-and-mouth disease: studies based on the virus neutralization reaction. J Biol Stand. 1984;12:323–37. DOIPubMedGoogle Scholar
- Eble PL, de Bruin MG, Bouma A, van Hemert-Kluitenberg F, Dekker A. Comparison of immune responses after intra-typic heterologous and homologous vaccination against foot-and-mouth disease virus infection in pigs. Vaccine. 2006;24:1274–81. DOIPubMedGoogle Scholar
- Sumption K, Pinto J, Lubroth J, Morzaria S, Murray T, De La Rocque S, Foot-and-mouth disease: situation worldwide and major epidemiological events in 2005–2006. EMPRES Focus On Bulletin. 2007;1:1–11. [cited 2007 Aug 21]. Available from http://www.fao.org/docs/eims/upload//225050/Focus_ON_1_07_en.pdf
Figures
Tables
Cite This ArticleTable of Contents – Volume 13, Number 10—October 2007
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Nick J. Knowles, Institute for Animal Health, Pirbright Laboratory, Ash Rd, Pirbright, Woking GU24 0NF, UK;
Top